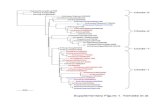
Farmer et al Supplementary Figure 1
description
Transcript of Farmer et al Supplementary Figure 1

Farmer et al Supplementary Figure 1
N
N
O
H
N
N
N
FN
O
O
O
N
N
N
O
FO
H
HKU0058684
PARP-1 IC50 = 3.2nM
KU0058948PARP-1 IC50 = 3.4nM
KU0051529PARP-1 IC50 = 730nM
a
b
0 0.00
10.
010.
1
1 10 100
0.00
01
0.00
001
[KU0058684] (M)
0 0.00
1
0.01
0.1
1 10 100
[KU0058948] (M)
0
20
40
60
80
100
120
IC50 = 1nM
% a
cti
vit
y
10-6 10-4 10-3 10-2 10-1 100 10110-5
[KU0058684] (M)
IC50 = 6nM
0
20
40
60
80
100
% a
cti
vit
y
[KU0058948] (M)
10-4 10-2 100 10110-3 10-1

1h
-4
-3
-2
-1
0
10-9
10-8
10-7
10-6
10-5
10-40
conc (M)
log
su
rviv
ing
fra
ctio
n
24h
-4
-3
-2
-1
0
10-9
10-8
10-7
10-6
10-5
10-4
0
conc (M)
log
su
rviv
ing
fra
ctio
n
4h
10-9
10-8
10-7
10-6
10-5
10-40
conc (M)
log
su
rviv
ing
fra
ctio
n
-4
-3
-2
-1
0
VC8 BAC
VC8
-4
-3
-2
-1
0
log
su
rviv
ing
fra
ctio
n
10-7
10-6
10-5
10-4
0
conc (M)
-4
-3
-2
-1
0
log
su
rviv
ing
fra
ctio
n
10-7
10-6
10-5
10-4
0
conc (M)
-4
-3
-2
-1
0lo
g s
urv
ivin
g f
ract
ion
10-7
10-6
10-5
10-4
0
conc (M)
D3Cre15Cre24
10-4
log
su
rviv
ing
fra
ctio
n
10-9
10-8
10-7
10-6
10-50
conc (M)
log
su
rviv
ing
fra
ctio
n
10-9
10-8
10-7
10-6
10-5
10-4
0
conc (M)
log
su
rviv
ing
fra
ctio
n
10-9
10-8
10-7
10-6
10-5
10-4
0
conc (M)
11COCre6Cre10
1h 4h 24h
-4
-3
-2
-1
0
-4
-3
-2
-1
0
-4
-3
-2
-1
0
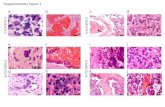
Farmer et al Supplementary Figure 2
a
b

a
10-1
1
10-1
0
10-9
10-8
10-7
10-6
10-5
10-4
0
conc (M)
log
su
rviv
ing
fra
ctio
n
-4
-3
-2
-1
0
10-1
1
10-1
0
10-9
10-8
10-7
10-6
10-5
10-4
0
conc (M)
log
su
rviv
ing
fra
ctio
n
-4
-3
-2
-1
0
-4
-3
-2
-1
0
10-1
1
10-1
0
10-9
10-8
10-7
10-6
10-5
10-4
0
conc (M)lo
g s
urv
ivin
g f
ract
ion
KU0058684 KU0058948 KU0051529
VC8 BAC
VC8
b
KU0051529
conc (M)
log
su
rviv
ing
fra
ctio
n
0
-1
10-8
10-7
10-6
10-40 10
-9
10-5
10-8
10-7
10-6
10-40 10
-9
10-5
conc (M)
KU0058948
log
su
rviv
ing
fra
ctio
n
0
-1
MCF7-3.23
MCF7-scrambled
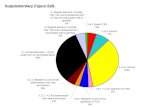
Farmer et al Supplementary Figure 3

0
1
2
3
1 2 3 4 5 6
0 10nM 1M 0 10nM 1M
Brca1 wild-type Brca1 deficient
KU0058684 concentration
Number per cell
Mean number of chromatid breaks
Mean number of complex chromatid rearrangements
D3 untreated D3 10nM Cre24 untreated Cre24 10nM0
1
2
3
0 10nM 0 10nM
Brca2 wild-type
Brca2 deficient
a
Farmer et al Supplementary Figure 4

Farmer et al Supplementary Figure 4
b
untreated10-9M 10-8M 10-7M 10-6M 10-5M
100
75
50
25
00 10-9 10-8 10-7 10-6 10-5
KU0058684 concentration (M)
10 9 8 7 6 5
100
75
50
25
00 10-9 10-8 10-7 10-6 10-5
KU0058684 concentration (M)
% of cells with >5
H2AX foci
% of cells with >5
H2AX foci
11CO
Cre10
D3
Cre24

Wild-type Brca1 deficient Brca2 deficient
0 10 0 10 0 100
10
20
30
40
50
60
70
80
90
100
M KU0058684
% of cells with >5
H2AX foci
c
Farmer et al Supplementary Figure 4
pSUPER-eCFP-control
pSUPER-eCFP-Parp-1
d
% of cells with >5 H2AX
foci
pSUPER-
eCFP-c
ontrol
pSUPER-
eCFP-P
arp-1
0
10
20
30
40
50
60

0 1 2 5 10 0 10 0 10 10
0
20
40
60
80
100
M KU0058684 M KU0051529
Wild-type Brca1 deficient Brca2 deficient Wild-type
% of cells with > 5 Rad51 foci
e
Farmer et al Supplementary Figure 4