EFFECTS OF LIVER EXTRACELLULAR MATRIX GEL … 2 Figure 1: Rat Liver Decellularization Figure 2: PEG...
Transcript of EFFECTS OF LIVER EXTRACELLULAR MATRIX GEL … 2 Figure 1: Rat Liver Decellularization Figure 2: PEG...
1
EFFECTS OF LIVER EXTRACELLULAR MATRIX GEL STIFFNESS ON
PRIMARY HEPATOCYTE FUNCTION
BY
DANIEL B. DEEGAN
A Dissertation Submitted to the Graduate Faculty of
WAKE FOREST UNIVESITY GRADUATE SCHOOL OF ARTS AND SCIENCES
in Partial Fulfillment of the Requirements
for the Degree of
DOCTOR OF PHILOSOPHY
Molecular Medicine and Translational Sciences
December 2015
Winston-Salem, North Carolina
Copyright Daniel B. Deegan 2015 (If Applicable)
Approved By:
Thomas D. Shupe, Ph.D., Advisor
Emmanuel C. Opara, Ph.D., Chair
Graça D. Almeida-Porada, M.D., Ph.D.
Michael C. Seeds, Ph.D.
Shay Soker, Ph.D.
2
DEDICATION AND ACKNOWLEDGEMENTS
A special thanks to my PI and advisor Dr. Tom Shupe for his guidance, patience,
and selflessness in mentoring me and helping me to complete the graduate school
process. Thanks to my undergraduate mentor at Virginia Tech, Dr. John McDowell, for
sparking my interest in laboratory research. Thanks to my previous mentors Dr. Tamer
Aboushwareb and Dr. Bryon Petersen for their assistance throughout graduate school.
Thanks to my labmate Cindy Zimmerman for her help and support on technical work of
my project. Thanks to my committee members, Dr. Emmanuel Opara, Dr. Graça
Almeida-Porada, Dr. Michael Seeds, and Dr. Shay Soker for their guidance on my thesis
project. Thanks to Dr. Aleksander Skardal for his support and expertise in hydrogel
formation. Thanks to Dr. Bridget Brosnihan for her counsel and advisement during
difficult times. Thanks to the Molecular Medicine and Translational Sciences program
and the Wake Forest Institute for Regenerative Medicine. Finally, thank you to my family
and friends for keeping me sane throughout the struggles of graduate school.
3
TABLE OF CONTENTS
List of Figures and Tables………………………………………………………………5-6
Abbreviations………………………………………………………………………...…6-8
Abstract……………………………………………………………………………..….9-10
Chapter 1…………………………………………………………………………..….11-59
Chapter 2……………………………………………………………………………...60-88
Chapter 3………………………………………………………………………...…..89-145
Chapter 4………………………………………………………………………...…146-159
Curriculum Vitae…………………………………………………………………..160-163
4
LIST OF FIGURES AND TABLES
Chapter 1
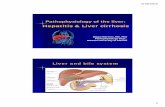
Figure 1: Hepatic Lobule Structure
Figure 2: Integrin Signaling and Regulation of Rho GTPases
Chapter 2
Figure 1: Rat Liver Decellularization
Figure 2: PEG Crosslinker Structures
Figure 3: PEGDA Crosslinking Saturation
Chapter 3
Figure 1: IHC of Decellularized Liver Tissue
Table 1: Storage Modulus Formulas
Table 2: Human Primers for RT-qPCR
Figure 2: Solubilized Liver Composition and Activity
Figure 3: Solubilized ECM Gel Increases Hepatocyte Viability
Figure 4: ECM Gels Increase Hepatocyte Attachment and Clustering
Figure 5: HA-ECM Gel Stiffnesses Formed by Varying Crosslinker Concentrations
Figure 6: Cell Viability and Filopodia Formation
Figure 7: Cytoskeletal Staining
Figure 8: Stiffness and Substrate Dependent Cell Attachment
Figure 9: Hepatocyte Phenotypic Marker Expression
Figure 10: Hepatocyte Functional Output
Figure 11: Expression of Cell Junction Molecules and Focal Adhesion Regulators
Figure 12: FAK and ILK ICC
Figure 13: Expression of the Cytoskeleton-Regulating Rho GTPases
Supplementary Figure 1: Growth Factor Binding Comparison
5
Supplementary Figure 2: Expression of HNF4α, SNAIL, and Claudin-1
Supplementary Figure 3: Integrin beta-1 Expression
Supplementary Figure 4: Hepatocyte Culture on Growth Factor Treated Gels
Supplementary Figure 5: Effects of Substrate Stiffness on Hepatic Stellate Cells
Chapter 4
No Tables or Figures
6
ABBREVIATIONS
Smooth endoplasmic reticulum SER
Cytochrome P450 CYP
Hepatic stellate cells HSC
Extracellular matrix ECM
Matrix metalloproteinases MMP
Alpha-smooth muscle actin α-SMA
Collagen-1 Col1
Glycosaminoglycans GAGs
Reactive oxygen species ROS
Blood cells RBC
Sinusoidal endothelial cells SEC
Transforming growth factor beta TGF-β
Endothelin-1 ET-1
Mitogen-activating protein kinases MAPK
Fibronectin FN
Arginine-Glycine-Aspartic acid RGD
Heparan sulfate HS
Chondroitin CS
Dermatan sulfate DS
Keratan sulfate KS
Hyaluronic acid HA
Hepatocyte growth factor HGF
Epidermal growth factor EGF
Fibroblast growth factor FGF
7
EGF receptor EGFR
Transforming growth factor alpha TGFα
Heparin binding EGF HB EGF
Basic fibroblast growth factor bFGF or FBF2
Urokinase-type plasminogen activator u-PA
N-2-acetylaminofluorene 2-AAF
Connective tissue growth factor CTGF
Mesenchymal to epithelial transition MET
Epithelial to mesenchymal transition EMT
Tissue inhibitor of metalloproteinases TIMP
Smooth muscle cells SMC
Hepatocellular carcinoma HCC
Cell adhesion molecules CAM
Focal adhesion kinase FAK
Phosphoinositide 3-kinase PI3K
Protein kinase B AKT
Crk-associated substrate CAS
c-Jun N-terminal kinases JNK
Mitogen-activated protein kinase 1 MAPK1
Mitogen-activated protein kinase 3 MAPK3
Ras homolog family member Rho
Ras-related C3 botulinum toxin substrate Rac
Growth factor receptor-bound 7 GRB7
Ras homolog family member A RhoA
Actin-related protein Arp 2/3
8
Cell division control protein 42 CDC42
Neural Wiskott-Aldrich syndrome protein N-WASP
Engulfment and Cell Motility 2 ELMO2
Integrin-linked kinase ILK
Rho-associated protein kinase ROCK
Polyethylene glycol PEG
Polyethylene glycol diacrylate PEGDA
Magnetic Resonance Elastography MRE
Loss modulus G’’
Storage modulus G’
Atomic force microscopy AFM
Superior vena cava SVC
Inferior vena cava IVC
Hepatocyte nuclear factor 4 alpha HNF4α
9
ABSTRACT
The extracellular matrix is a complex environment of mechanical and chemical
cues vital to individual cell growth, tissue formation, and ultimately complete organ
function. The field of regenerative medicine aims to recreate the unique biological
properties found in specific tissue types and to incorporate these properties into substrates
that support long term cell function for applications including tissue engineering, cell
therapies, and disease models. Because of the complexity of the extracellular matrix,
difficulties exist in characterizing and utilizing components of the matrix that contribute
to the long term function and viability of cells. Primary liver epithelial cells, or
hepatocytes, are easily isolated in large quantities. However, these cells quickly lose
viability and function once removed from the native organ. In order to support hepatocyte
phenotype in culture, decellularized liver matrix was incorporated into transplantable
hyaluronic acid hydrogels. These substrates were shown to bind growth factors, support
hepatocyte attachment, and promote cell junction formation. In liver tissue, the physical
property of matrix stiffness affects a myriad of cell behaviors including; cell attachment,
viability, function, growth factor utilization, motility, and cytoskeletal organization. Gels
were crosslinked at different stiffnesses in order to mimic and test a variety of mechanical
environments. The hepatocyte substrates were formulated within a narrow physiologic
range from slightly above to slightly below that of native liver. Primary human
hepatocyte attachment, viability, and functional output increased with stiffness. Stiffness
influenced hepatocyte morphology, cell signaling, and expression of important liver
specific markers dependent on duration in culture and presence or absence of matrix in
the gel. In conclusion, data demonstrates that inclusion of extracellular matrix in primary
10
hepatocyte cell culture substrates affects cell phenotype, and these effects are influenced
by small changes in stiffness of the substrates. This thesis work suggests that substrate
formulation and stiffness can be optimized for liver cell culture and different regenerative
medicine applications.
11
CHAPTER 1
Introduction to Liver Architecture and the Role of the
Extracellular Matrix on Liver Cellular Function
Daniel B. Deegan
Content
1. Function and Gross Anatomy……………………………………………..13
1.1 Liver Function…………………………………………………………13
1.2 Liver Vasculature……………….……………………………...……...14
1.3 Macroscopic Structures………….……………………………..….…..14
2. Cell Types of the Liver……………….……………………………………15
2.1 Hepatocytes……………………………………………………....……15
2.2 Hepatic Stellate Cells…………………………………………....…….17
2.3 Kupffer Cells……………………………………………….......……..18
2.4 Sinusoidal Endothelial Cells…………………………………...……...19
2.5 Cholangiocytes (Biliary Epithelial Cells).…………………...………..20
3. ECM Components………………………………………………….………20
12
3.1 Fibrillar Collagen…………………………………………………...….21
3.2 Non-Fibrillar Collagens………………………………………………..22
3.3 Non-Collagenous Glycoproteins…………………………….…………22
3.4 Glycosaminoglycans……………………………………………....…...24
3.5 ECM Bound Growth Factors…………………………………….……..26
4. Liver Regeneration…………………………………………………………28
4.1 Normal Liver Repair and Regeneration………………………………..28
4.2 Progenitor Mediated Repair and Regeneration………………………...30
5. Liver Disease………………………………………………………………31
5.1 Metabolic Disease………………………………………………….…..31
5.2 Cholangiopathies……………………………………………….………32
5.3 Progression of Liver Fibrosis…………………………………………..32
6. Stiffness in Tissue Microenvironments……………………………….……34
6.1 ECM Stiffness…………………………………….………….……35
6.2 Cellular Stiffness………………………………….………….……36
7. Liver ECM Stiffness during Developmental or Repair States.………….…37
8. Effects of Stiffness on Mechanotransduction and Focal Adhesion Signaling 39
8.1 Cell Adhesion Molecules……………………………………….…39
8.2 Mechanotransduction and Cytoskeletal Regulation…………….…41
13
1. Function and Gross Anatomy
1.1 Liver Function
The liver is the largest internal organ of the human body and performs over 500
unique functions (1). It is a major site of plasma protein biosynthesis, energy storage,
metabolism, and detoxification. The liver produces important compounds including
clotting factors, heparin, and albumin that give circulating blood many of its important
physical properties (1). It also produces bile, which aids the digestion of lipids into fatty
acids that are used throughout the body (2, 3). The liver is the main storage site for
glycogen and serves as a buffer for blood glucose levels through activation of
glycogenolysis, glycogenesis, or gluconeogenesis (4, 5). The organ also stores fat soluble
compounds, vitamins, and minerals for use by cells throughout the body.
The liver is not only crucial to the production and storage of many of the building
blocks used within the body, but is also vital for the removal of toxic compounds from
the system, such as cellular waste products and xenobiotics (6). The liver aids in the
filtration of damaged or dying red blood cells from the circulation. Bilirubin, formed
from breakdown of hemoglobin, causes jaundice unless processed and excreted by the
liver (7). The liver also metabolizes and excretes alcohol and toxic drug compounds like
acetaminophen. These compounds are detoxified by a two-phase process where
electrophilic intermediate metabolites are conjugated by substrates. This results in a
detoxified polar product that may be excreted in the bile or urine (8).
14
1.2 Liver Vasculature
The importance of the liver is evidenced by the fact that this organ receives ¼ of
the entire cardiac output. 25% of the blood moves directly from the heart through the
hepatic artery to the liver. The other 75% of input enters the liver though the portal vein
and is lower in oxygen content but contains nutrient rich blood from the intestines (9).
This portion of the blood supplies the liver with needed factors, while also allowing the
liver to filter and detoxify xenobiotic compounds entering the circulation through the
digestive tract.
1.3. Macroscopic Structures
The human liver is composed of four main lobes; the left, right, caudate, and
quadrate lobes. These lobes are further divided into subunits called lobules. Lobules are
hexagonal shaped arrangements that contain discontinuous fenestrated endothelium, or
sinusoids (10). These sinusoids are in intimate proximity to the liver parenchyma. The
unfettered diffusibility through the sinusoids allows for greater interactions of
parenchymal cells with compounds carried in the blood, as compared to endothelium
within other tissue types (11). Each liver lobule is fed by 6 portal triads positioned at the
periphery of the lobule. This structure is composed of a portal vein and hepatic artery, as
well as a bile duct. Blood flows out of the lobule through the central vein that is
positioned in the middle of the lobule. Bile canaliculi run parallel to the liver sinusoids
but flow in the opposite direction of the sinusoidal circulation (12). The canaliculi merge
with the biliary tree at structures called the Canals of Hering, which flow to the common
bile duct that terminates in the gall bladder (13).
15
(14)
2. Cell Types of the Liver
The liver contains least 15 unique cell types (10). These cells types can be
narrowed down to 5 main cell types found within and around the parenchyma, sinusoids,
and bile ducts of the liver. Within these liver regions, cells communicate through
autocrine and paracrine signaling to support various cell phenotypes. Together, these
cells function to affect liver homeostasis and maintain a healthy organism.
2.1 Hepatocytes
Hepatocytes are the main cell type of the liver parenchyma and can carry out all
major functions of the liver. Chief products include bile, cholesterol and albumin (15).
Hepatocytes constitute 60% of the cell number and 80% of the actual volume of the liver
(10, 16). They are polarized cells arranged in two-dimensional cords with one side of the
cell communicating with the hepatic sinusoid, while the opposing side communicates
Fig.1) Hepatic Lobule Structure, adapted from Mescher et al (14)
16
with the bile canaliculus (17, 18). The surface of the hepatocyte is covered with small
projections known as microvilli which extend into the perisinusoidal area between the
hepatocyte and the endothelium, called the space of Disse. These structures greatly
expand the surface area of the cell, facilitating interactions with blood plasma that
diffuses through the sinusoidal wall (19). Hepatocytes attach to each other by tight and
adherens junctions on their lateral surfaces (20). These junctions isolate the apical
surface, forming the canaliculus through which bile flows. These bile canaliculi provide
complex linkages surrounding the cells that allow bile to drain into the bile ducts of the
liver (17).
The functional roles of hepatocytes vary according to their location within a
lobule (21). The lobules can be divided into three zones according to differences in
oxygen, nutrient, and metabolite concentrations (22). Zone 1 is an oxygen and nutrient
rich area closest to the portal triads at the periphery of the liver lobules. In this zone,
hepatocytes have greater numbers of mitochondria which provide higher rates of
oxidative phosphorylation that drive processes requiring great amounts of energy such as
gluconeogenesis. Cells within this zone are also more frequently affected by viral
infections like hepatitis because of their proximity to the portal area (23). Zone 2 is a
transition region of lower oxygen content. Zone 3 surrounds the central vein and has
lowest oxygen and nutrient concentration. Hepatocytes in zone 3 contain less
mitochondria and a greater amount of smooth endoplasmic reticulum (SER) (24). They
carry out processes such as glycolysis, lipid production, and xenobiotic detoxification
that remove harmful compounds before they reenter the circulation (25, 26). These cells
are also vital for the detoxification of bilirubin. Because of their roles in filtration and
17
excretion, these cells highly express cytochrome P450 (CYP) enzymes and are the cells
most negatively impacted by toxin intake (27). The low oxygen content in zone 3 also
makes hepatocytes of this area most susceptible to ischemic damage resulting from
cirrhosis (26).
Upon liver injury, hepatocytes rapidly proliferate to regenerate liver tissue. The
proliferative capacity of hepatocytes allows the liver to repair itself, despite ongoing
contact with toxins or loss of up to 75% of tissue mass (28, 29). This unique ability is
most likely the result of evolutionary changes and adaptation to the liver’s role in the
body as a site of detoxification and waste processing.
2.2 Hepatic Stellate Cells (HSC)
Hepatic stellate cells are the architects of the liver and comprise 5-8% of the total
liver cell number (30). These cells exist in two states in the space of Disse, a quiescent fat
storing cell or an activated myofibroblast (31). In their normal quiescent state, stellate
cells have a spindle-like morphology with elongated processes that extend between the
sinusoids and hepatic cords and enable them to sense changes in the microenvironment
(32). These cells store fat droplets and fat soluble compounds such as vitamin A. Even if
not fully activated, cells in the less studied quiescent state have been reported to influence
the extracellular matrix (ECM) by balancing the production of ECM degrading matrix
metalloproteinases (MMPs) with low levels of matrix protein biosynthesis molecules (33,
34). Quiescent cells have also been shown to secrete soluble growth factors such as
hepatocyte growth factor (HGF) (31).
18
Hepatic stellate cells become activated when they sense physical cues in the
matrix or chemical signals released from immune cells and dead or dying hepatocytes
resulting from toxicity, metastatic tumor cells, or viral infections. Upon activation, these
cells express genes for alpha-smooth muscle actin (α-SMA) and collagen-1 (Col1) (32,
33). They adopt a myofibroblastic morphology, proliferate, and secrete various pro-
inflammatory and mitogenic cytokines. HSCs travel to sites of injury and remodel the
damaged tissue matrix by the production of MMPs, hepatocyte-influencing growth
factors, and ECM molecules (35). The majority of ECM molecules generated are
collagens. However, HSCs also produce several other ECM components including
fibronectin, laminin, and glycosaminoglycans (GAGs) (36). Although vital to repair and
acute injury response, HSCs can become overactive in a chronic liver injury state. This
imbalance causes excess deposition and accumulation of ECM in the liver, eventually
leading to fibrosis and cirrhosis (37, 38).
Like hepatocytes, HSC morphology and composition change between three zones
of the lobules. In the periportal region, stellate cells have a smaller cytoplasmic volume
and project numerous extensions into the space of Disse. In the middle zone, cells are
larger, longer, and contain greater reserves of lipids and desmin. Moving into the
pericentral zone, desmin levels in stellate cells decrease, while vitamin A amounts
increase (32).
2.3 Kupffer Cells
Kupffer cells are star shaped immune cells located within the sinusoidal barrier.
Although they only represent a small percentage of all cells within the liver, they
19
represent 80-90% of all macrophages in the body (39). Upon activation, Kupffer cells
secrete inflammatory cytokines, growth factors, and reactive oxygen species (ROS) in
response to various stimuli. These macrophages phagocytose large particles and eliminate
dead or damaged red blood cells (RBCs), cancer cells, and hepatocytes, as well as
bacteria and other foreign entities (40). Kupffer cells process millions of RBCs per
minute and breakdown their main component, hemoglobin, into reusable polypeptides,
iron, and bilirubin. Toxic bilirubin attaches to albumin within the blood and eventually
diffuses through the sinusoidal lining. In the space of Disse, hepatocytes conjugate
bilirubin for excretion out of the body (41).
2.4 Sinusoidal Endothelial Cells
Sinusoidal endothelial cells (SECs) constitute 20% of the total liver cell number.
Unlike normal endothelial cells, hepatic sinusoidal cells form a fenestrated endothelium
that lies on a discontinuous basal lamina in the liver lobules. These sinusoids control
passive diffusion of substances into the parenchyma through sieve-like action and filter
entering plasma as it passes from the blood into the space of Disse. Porosity of the
endothelium increases from the portal triad to the central vein (42). This property allows
for greater diffusion and interaction of waste products and metabolites with hepatocytes
and HSCs in zone 3. Besides passive diffusion, SECs also scavenge waste products and
cell remnants through phagocytosis or actively transport compounds from the sinusoidal
lumen to perisinusoidal space through transcytosis (16).
SECs secrete a wide variety of cytokines, growth factors, and other compounds
into the circulation. SECs contribute to the generation of fibronectin, collagen IV, and
20
participate in the activation of transforming growth factor beta (TGF-β) to the active form
(26, 43). These cells also produce autocrine vasoactive compounds, such as endothelin-1
(ET-1), to affect blood flow and uptake from the sinusoidal lumen (44). Presence and
long term exposure to certain drugs or toxins such as nicotine, endotoxin, and ethanol
decrease fenestration size and reduce the ability of molecules like cholesterol to pass
through the sinusoids for hepatocyte processing (10). This build-up in cholesterol can
lead to atherosclerosis. Defenestration also blocks retinol metabolism and activates HSCs
within the space of Disse (45). This activation exacerbates fibrosis of the perisinusoidal
space, eventually resulting in cirrhosis.
2.5 Cholangiocytes (Biliary Epithelial Cells)
Cholangiocytes are epithelial cells that represent an alternate differentiation
lineage of liver progenitor cells. Cholangiocytes vary in size and morphology and line
extrahepatic and intrahepatic bile ductules (46). Unlike hepatocytes which produce bile-
acids that must be actively transported to the canaliculus, cholangiocytes secrete
components of bile such as electrolytes that enter the bile ducts through passive ionic
exchange processes (39). Cholangiocytes modify bile by secretion and adsorption of ions
and other molecules as it passes through the biliary tree. Cholangiocytes ultimately traffic
bile through the liver, which exits to both the common bile duct and the gall bladder (47).
3. ECM components
As has been shown in multiple cell types, availability of nutrients and oxygen
controls cell morphology and functional capacity. Another factor which greatly
determines cell activity and viability is the underlying ECM. Liver cells are organized
21
and maintained by a complex extracellular microarchitecture. Unlike other organs, the
ECM composes less than 10% of the liver volume (48). Despite this small percentage,
ECM molecules play a tremendous role in liver function regulation, and direct repair and
regeneration processes within tissue (49). When out of balance, excess ECM deposition
causes dysfunction and disease states of the liver. Fibrillar collagens constitute the
majority of the liver, but non-fibrillar collagens, glycoproteins, and glycosaminoglycans
also play important roles in cell physiology.
3.1 Fibrillar Collagen
Fibrillar collagens, including collagens I, III, and V, comprise the largest
percentage of ECM proteins in the liver. These collagens contain rigid triple helix amino
acid structures and form heterogenous fiber bundles that give the liver its strength and
mechanical properties (49). Collagen I is the thickest and most prevalent type of fibrillar
collagen. It is a highly ubiquitous protein but can be found in the highest concentrations
in the periportal and pericentral regions (50). It contains several binding sites per peptide
that attach α1β1 and α1β2 integrins. Collagen I also possesses several non-specific low
affinity binding sites for cell attachment (51). Many other ECM components can complex
with collagen I to interact with cells including other collagens, fibronectin, and
proteoglycans.
Collagen III is structurally similar to collagen I and is found in all areas of the
liver. Greater amounts are present in the periportal regions (22). This collagen type also
comprises a large percentage of developing tissue. In adult liver, collagen III associates
strongly with collagen I bundles. Collagen V is the last major fibrillar collagen found
22
throughout liver. Their thin fragile fibers link multiple collagen types to each other to
form cohesive units at both ends of the sinusoids (50).
3.2 Non-Fibrillar Collagens
Non-fibrillar collagens are a category of basement membrane molecules with
fragmented triple-helical structures. Their flexible structures create networks between
other ECM components (49). Collagen IV is the main constituent of the basement
membrane and secures other basement membrane components like perlecan and laminin
(52) to the underlying structure. This collagen is chiefly secreted by endothelial cells. It
forms the discontinuous sinusoids of liver lobules that allow for blood plasma flow into
the space of Disse (49).
Other collagens of the basement membrane include collagen VI and VIII.
Collagen VI forms networks with different collagen types and assembles collagen
bundles, including collagens I and III fibers. Collagen VI affixes smaller basement
membrane molecules as well as larger structures, such as blood vessels, by attachment
and connection to collagen IV subunits (50). This collagen also functions in tissue repair
through the binding and release of soluble factors that regulate mitogen-activating protein
kinases (MAPKs). Collagen VIII also binds cytokines that may be liberated to initiate
wound healing, especially during angiogenesis (49). This collagen type connects elastic
fibers of the vasculature and contributes to the physical properties of vessels.
3.3 Non-Collagenous Glycoproteins
Fibronectin and laminin are the most widespread glycoproteins of the liver. Both
glycoproteins have numerous roles in development, injury repair, and normal cellular
23
activity. Fibronectin (FN) is a high molecular weight protein dimer that encompasses
numerous variants with two principal forms; plasma and cellular fibronectin. Plasma
fibronectin is the most abundant glycoprotein found in the liver and is synthesized
primarily by hepatocytes (53). Cellular fibronectin is an insoluble form that exists at low
levels in normal liver tissue. The highest concentrations of this form are found in the
pericentral area (22). It is secreted by a variety of cell types, but the main producer is the
stellate cell. Cellular FN binds several other ECM components including collagen I,
perlecan, and fibrin (54). It concentrates in bundles attached to hepatocyte microvilli and
collagens of the space of Disse (55). It also occupies the pericellular spaces and basal
membranes in the liver’s portal regions (56, 57)
During tissue repair, plasma FN enters the site of injured tissue and initiates ECM
degradation and remodeling by activation of Kupffer cells and hepatic stellate cells. In
this process, activated stellate cells produce MMPs that degrade plasma FN. Hepatocytes
replace plasma FN with insoluble cellular FN for incorporation into the ECM (55).
Fibronectin contains the Arginine-Glycine-Aspartic acid (RGD) cell binding sequence
that interacts with several integrin types. With these high affinity binding sites, cellular
FN acts as a chemoattractant, cell adhesion molecule, and growth factor for migrating
cells.
Laminin is another RGD-containing ligand found chiefly in the basement
membrane of the portal region and the sinusoidal wall. It is also present in certain liver
growth and developmental states in the perisinusoidal space (58). Laminin attaches cells
of the sinusoids through integrin interactions and can induce cell migration and motility.
This glycoprotein secreted by hepatic stellate cells is vital in vascular structural
24
maintenance, but is also important in the regulation of development and differentiation in
the liver (49, 59).
3.4 Glycosaminoglycans
GAGs are another important group of molecules that promote cell adhesion and
viability within the ECM. GAGs are long polysaccharide chains of various sizes and
sulfation states (60). There are six total GAG types that are classified into four
structurally distinct families; heparan sulfate (HS)/heparin, chondroitin (CS)/dermatan
sulfate (DS), keratan sulfate (KS), and hyaluronan (61). Most of these polysaccharides
covalently attach to distinct protein cores to form proteoglycans and integrate with the
ECM. The exception is HA, which does not directly covalently bond to proteins to form
proteoglycans, but directly and independently attaches to the ECM (62).
GAGs bind and affect enzymes, enzyme inhibitors, cell attachment molecules,
ECM proteins, and growth factors of the liver and other organ systems. These proteins
contain positively charged amino acids that form ionic and hydrogen bonds to negatively
charged sulfates and carboxylates of GAGs (63, 64). Specificity and strength of these
interactions are determined by protein confirmation and how certain amino acid residues
align with the arrangement of sulfated sites (62, 63). Through these interactions, GAGs
influence the mechanical properties of the ECM, enzyme activation, and activity of a vast
array of tissue specific growth factors.
GAGs greatly increase the half-life of growth factors compared to soluble growth
factors that remain free in plasma. GAG binding protects these proteins from proteolytic
degradation (65). This interaction also enhances presentation and utilization of growth
25
factors by clustering bound growth factors close to cell receptors of attached cells (66).
With the capacity to control periodic release, cells easily exploit housed growth factors
when needed for growth and functional maintenance (19). The ability to preserve and
display growth factors reduces the need of tissues to generate large quantities of growth
factors in a short period of time, which might produce adverse systemic effects.
The most common proteoglycan in the liver is perlecan, which is mainly produced
by hepatocytes and is comprised of the GAG, heparan sulfate (67). HS proteoglycans
compose 60% of all proteoglycans in the liver and are normally found in the ECM of the
sinusoids, perisinusoidal space, and sinusoidal walls (49). HS has a similar structure to
heparin but is generally longer in length and more varied in disaccharide and sulfated
configurations (63, 68). This complexity and flexibility in arrangement allows HS to
interact with a wide variety of biologically active proteins, growth factors, and cellular
receptors (69). This GAG binds to collagen, fibronectin, and laminin in the ECM to
support tissue structure and cell adhesion. HS binding to collagen organizes collagen
bundles. Interactions with fibronectin and laminin control their confirmation and
biological activity (70). HS is also a receptor for several different growth factors that
affect cell growth and function, which includes a high specificity for HGF in the liver
(62, 71, 72).
Hyaluronic acid (HA) comprises only a minor fraction of the liver ECM and total
proteoglycan volume but still has significant roles in the physical properties of liver
tissue. HA sequesters water to maintain hydration of the ECM and also provides
lubrication to enable cell motility and migration (73). HA is important during new liver
tissue formation. Concentrations of HA increase during development and tissue
26
regeneration to promote activity of proliferating cells (74). These properties and its
ability to avoid immune reactions upon transplantation make HA a suitable substrate for
cell culture and eventual use for in vivo treatments (75).
3.5 ECM Bound Growth Factors
Growth factors have a wide range of functions that vary from cell growth and
proliferation to phenotypic determination and maintenance. Glycoproteins of the ECM
bind these growth factors non-covalently through hydrogen binding and electrostatic
interactions to protect them from degradation, concentrate them in specific areas of the
tissue, and regulate interactions with cell receptors (76-78). Following attachment to
nearby ECM molecules, cells utilize proteolytic cleavage to free bound growth factors,
making them available to interact with membrane bound receptors (79). Numerous
growth factors are important to the regulation of liver function. Three of these proteins
with the most significant impact in the liver are hepatocyte growth factor (HGF),
epidermal growth factor (EGF), and fibroblast growth factor (FGF).
HGF is a unique heterodimer chiefly secreted by mesenchymal cells, especially
HSCs in the liver. It is also produced in lower levels by SECs in the liver (80, 81). After
activation of this growth factor by serine protease, HGF acts as a powerful mitogen and
stimulates cell proliferation by binding to the c-Met receptor of epithelial and endothelial
cells to activate tyrosine kinase signaling (82, 83). HGF strongly affects both liver
progenitor cells and differentiated hepatocytes (84-88). Studies with mesenchymal stem
cells showed that the presence of HGF on surrounding ECM drives differentiation toward
adult hepatocytes (89). HGF is vital to both progenitor cell and hepatocyte-mediated liver
27
regeneration. Spikes in HGF release, up to 20-fold normal plasma levels, correlate with
increased liver mass production (29, 90, 91). This protein also controls morphology,
migration, and tissue organization during reparative processes in the liver (84, 92, 93). In
animal models, HGF treatment accelerated recovery from cirrhosis and enhanced
regeneration (94). Because of its mitogenic properties, increased HGF levels have also
been implicated in aiding development of certain types of cancer (95).
HGF binds with a high affinity to the liver matrix through the GAGs, heparin and
heparan sulfate. HS mediates HGF use by cells and increases its half-life from 4 minutes
in blood plasma to several hours to match usage rates of surrounding cells (96-98). HGF
also has the ability to bind to areas of the matrix without HS through low affinity
interactions with several collagen types (99). These properties of HGF show its
importance in maintaining hepatocyte viability and function and demonstrate how
substrates could be designed to maximize HGF utilization in vitro or during cell
therapies.
While the importance of HGF in the maintenance of hepatocytes is well
established, other factors greatly affect and support liver cells. EGF is an important
growth factor that affects growth, viability, and differentiation of cells. EGF is primarily
produced in the Brunner’s gland of the duodenum (100). In the liver, it enters through the
portal vein of the portal triad and stimulates G1 to S phase transition and activates cell
division of hepatocytes. Unlike HGF, EGF acts through the tyrosine kinase EGF receptor
(EGFR) (101). It is vital for normal liver regeneration and induces Cyclin D1 (102, 103).
Other ligands of the EGFR also have similar mitogenic effects to EGF, like transforming
growth factor alpha (TGFα) and heparin binding EGF (HB EGF). They are manufactured
28
by endothelial cells and Kupffer cells in the liver and have both autocrine and paracrine
roles in proliferation (29, 104).
Another highly mitogenic growth factor family is the fibroblast growth factor
family. Like with HGF, heparin and heparan sulfate have a strong affinity for FGF. There
are 22 members of FGF family, but the most studied and recognized member is basic
fibroblast growth factor (bFGF or FBF2) (91, 105). FBF2 is secreted by HSCs in the
perisinusoidal region and, similar to HGF and EGF, is vital to wound healing and
angiogenesis in the liver (106).
4. Liver Regeneration
The roles of all of these tissue components show the importance of interplay
between cells, ECM structural molecules, and growth factors in the liver. These
relationships are not only important for normal function but also for development and
regeneration. Liver regeneration is a well-studied growth process which gives insights
into ontogeny of all organs (107). During liver regeneration, there is a complex step-wise
process that creates coordination between all liver components in order to grow and
organize new tissue. The exact mechanisms that mediate liver regeneration and the cell
phenotypes involved in the process are still hotly debated.
4.1 Normal Liver Repair and Regeneration
Loss of liver tissue, either by physical removal of tissue mass or acute chemical
toxicity, initiates the liver regenerative process. In the case of acute liver failure, dying
cells stimulate an immune response by Kupffer cells to clear dead cellular material (108).
Kupffer cells then activate HSCs and hepatocytes to begin regeneration of liver tissue
29
(29, 109). According to one model, liver regeneration is primarily accomplished by
proliferation of adult hepatocytes of the parenchyma (110). An alternate theory states that
regeneration is a delicate balance between epithelial to mesenchymal transitions (EMT)
and mesenchymal to epithelial transitions (MET), similar to the lineage transitions that
occur during development. This theory claims that following injury, epithelial cells
within the liver undergo EMT and migrate to the periportal zone. These fibroblasts
initiate repair and replicate before reverting back to hepatocytes or cholangiocytes in
order to repopulate the liver parenchyma (111).
During liver restoration, one of the first molecules secreted to regulate cells is a
proteinase called urokinase-type plasminogen activator (u-PA). It is released as rapidly as
1 minute following partial hepatectomy and correlates with an increase in its receptor u-
PAR (58). Besides releasing plasminogen, u-PA frees and activates ECM bound HGF for
immediate use and stimulation of hepatocytes. A peak in the release of HGF occurs just
one hour post-partial hepatectomy. During this time, HSCs, Kupffer cells, and SECs are
activated to release HGF, EGF, and other growth factors to promote further hepatocyte
proliferation, as well as autocrine mediated proliferation. Peaks in growth factor
production occur at 24 hours and 48 hours post-injury (90, 91, 112, 113).
MMPs are also produced to degrade old or damaged matrix and allow for
unrestricted hepatocyte replication. As a result, fibronectin and other cellular attachment
molecules decrease in the periportal regions until after the cellular mass is restored(29,
59). Hepatocytes proliferate starting in the periportal area. Proliferation proceeds into the
mid-lobular area, before finally initiating in the pericentral region. Full cellular mass is
replaced by 5-7 days in assays with rat models and around 3 months in humans (28, 114).
30
In normal intact ECM, HSCs simultaneously remodel matrix as new cells
proliferate. HSCs fully transdifferentiate into myofibroblasts and migrate to damaged
areas to lay down ECM, starting with collagen I and FN in the perisinusoidal space (18).
However, in an injury where whole tissue is lost, HSCs delay ECM production until new
hepatocyte clusters are created. HSCs infiltrate newly proliferated masses and secrete
laminin, collagen, and fibronectin to align hepatocytes into cords (59). This action
attracts endothelial cells to form new bile ducts and fenestrated sinusoids. Hepatocyte
division ceases when newly synthesized glycoproteins sequester HGF (115). Feedback
mechanisms between HSCs and hepatocytes also increase production of TGF-β1, which
halts new HGF secretion and returns proliferating hepatocytes to their resting states (35).
In the days that follow, further ECM is produced to organize cells into proper lobule
structures and restore full function in the liver.
4.2 Progenitor Cell Mediated Repair and Regeneration
In specific cases of liver repair, massive necrosis or chronic injury inhibits
proliferation of the resident hepatocyte population. This scenario is modeled in rats by
treatment of the chemical N-2-acetylaminofluorene (2-AAF) followed by partial
hepatectomy (116). In these situations, the liver activates bipotential hepatic progenitor
cells, to repopulate the tissue. In progenitor cell mediated regeneration, HSCs work in
close proximity with progenitor cells to direct migration, proliferation, and differentiation
(115, 117). HA deposits surround these two cell types during OC regeneration and
provide lubrication to allow cells to more easily migrate (38, 73). HSCs create a
provisional fibronectin rich matrix, which guides progenitor cells to damaged tissue with
the aid of bound connective tissue growth factor (CTGF) (116). CTGF and HS-bound
31
HGF stimulate progenitor cells to attach and replicate (118). These bipotential cells either
become differentiated hepatocytes or cholangiocytes to replenish lost tissue. Evidence
also exists that stellate cells themselves could undergo mesenchymal to epithelial
transition in order to bolster the compensatory hyperplasia (119). Following attachment
and differentiation, fibronectin and HA concentrations diminish, and a normal matrix
composition replaces the provisional matrix of the liver. This alternative method of
regeneration proves that nature has more than one method of repair in the body to
overcome different scenarios.
5. Liver Disease
Repair or regeneration is a delicate balance between matrix production, matrix
degradation, and cell proliferation. Despite the extraordinary abilities of the liver to repair
itself from acute trauma, problems arise when imbalances exist. Persistent stresses of
toxins, alcohol, or viral hepatitis cause fibrosis and advanced cirrhosis, which lead to
health complications or mortality in humans (52). Autoimmune responses, metabolic
disorders, and simple genetics can also initiate liver dysfunction in both infants and adults
(120). Many hallmarks of liver disease can be attributed to either the overactivation of
fibroblasts and/or dedifferentiation of hepatocytes. In certain disease states, loss of
function in cholangiocytes is also evident.
5.1 Metabolic Disease
Inheritance of faulty genes can cause specific enzyme deficiencies which result in
disorders of the liver. These deficiencies can occur in several different cell organelles and
impair various metabolic functions. Lysosomal and peroxisomal disorders can lead to
32
buildup of excess toxins and wastes that damage liver tissue (121). Specific diseases
related to these organelles include Tay-Sachs disease, Gaucher disease, and Zellweger
syndrome (122). Hemochromatosis is a deficiency in metabolism of metals in the body
which creates an accumulation of toxic levels of iron in the liver. Problems with
metabolism of galactose (galactosemia) or glycogen storage disease can lead to
impairment in regulation blood glucose levels. These imbalances can cause energy loss
and swelling of the liver (123). All of these metabolic diseases are currently incurable but
can be managed with proper nutrition or enzyme replacement therapies.
5.2 Cholangiopathies
Chronic liver disease is the progressive loss of liver function which can result in
end-stage liver disease and death. One of the cell types potentially targeted during the
development of chronic liver disease is the cholangiocyte. Cholangiopathies were
responsible for 16% of liver transplants from the years 1988 to 2014 (124). These
diseases include primary sclerosing cholangitis, biliary atresia, cystic fibrosis, and biliary
cirrhosis. During progression of these disorders, apoptosis of cholangiocytes,
inflammation, and excess collagen deposition results in obstruction or destruction of bile
ducts. This biliary degeneration can lead to increased levels of lipids and cholesterol,
portal hypertension, and liver scarring ultimately resulting in high mortality rates (124,
125).
5.3 Progression of Liver Fibrosis
One of the chief orchestrators of chronic liver disease and patient morbidity is
HSCs. Constant trauma and damage triggers release of inflammatory mediators and pro-
33
fibrotic cytokines that activate HSCs and portal fibroblasts (126). These activated cells
deposit excess quantities of ECM leading to hallmarks of fibrosis. Hepatocytes, SECs,
and cells that have undergone EMT also contribute to increased matrix and cytokine
production (43).
Collagen I, the most prevalent ECM component, increases throughout the liver
tissue and initiates further HSC activation. Cells in the space of Disse of the periportal
region deposit abnormal amounts of laminin, type IV collagen, and perlecan. Collagen IV
can increase up to 10-fold, while perlecan increases 8-times the normal liver
concentration (49). This build-up of matrix increases endothelial cell attachment and
eventually closes off the fenestrated sinusoids (42). Creating a continuous basement
membrane seals off the sinusoids and has several detrimental effects on the liver. Most
importantly wastes, toxins, and xenobiotics no longer easily interact with hepatocytes and
instead, move through the liver into the peripheral circulation. If left unprocessed, excess
bilirubin, the byproduct of hemoglobin catabolism, causes jaundice (127). Closed
sinusoids also prevent lipids from being metabolized in the parenchyma and leads to
atherosclerosis in patients (42). With increased fibrosis, sinusoids and liver vessels
constrict and block, which restricts blood flow and causes portal hypertension in patients
(128).
Changes during fibrosis also occur in the ECM of the centrilobular region. This
area accumulates irregular levels of fibronectin, type III collagen, and dermatan sulfate.
Elevated amounts of insoluble cFN activate HSCs and increase SEC attachment.
Increased plasma FN causes aberrant Kupffer cell activation and increased inflammation
34
in the liver. Without corrections in matrix deposition, cirrhosis leads to complications and
total loss of liver function.
Fibrosis is not only the result of increased matrix synthesis but additionally from
reduced matrix degradation. During normal liver repair, HSCs maintain an equilibrium
between production of MMPs and tissue inhibitor of metalloproteinases (TIMPs) (57).
However, during fibrosis, TIMPs, especially TIMP1 and TIMP2, are secreted at a greater
rate and block any action of MMPs (49). This inhibition, along with increases in matrix
production, worsens liver health and function. To treat liver disease, therapies need to
both revert HSCs and other ECM secreting cells to quiescent states and block the release
of TIMPs. Through these actions, balance and normal liver function can be restored.
6. Stiffness in Tissue Microenvironments
Increased collagen deposition and crosslinking during fibrosis cause changes in
the mechanical properties of the ECM. Mechanical properties of tissue
microenvironments contribute to the overall function and health of the organ systems
they comprise. These forces include shear stress induced by laminar flow in the
vasculature, tension from attachment to adjacent cells, and compression or bending of
tissue in response to external stimuli (129). One of the physical factors with the greatest
impact on cell behavior is stiffness. Stiffness is the mechanical property that represents
rigidity or the resistance to deformity following an applied force (130, 131). Variations in
stiffness are found in both the cells and extracellular matrix (ECM) that constitute tissue.
35
6.1 ECM Stiffness
ECM stiffness varies with tissue or organ type. Brain and fat are two of the softest
tissue types, while bone has the most rigid matrix composition and is several-fold stiffer
(132). The mechanical properties of ECM directly correlate to tissue function. Physical
characteristics of the matrix are determined by its composition of structural molecules,
the tension generated by cell attachment and migration, and exogenous physiological
forces such as blood flow.(133, 134) Stiffness also depends on location within the tissue
or organ. Gradients of bone and skeletal muscle stiffness can be found in the body. ECM
composition in the liver changes from the periportal to the pericentral regions of lobules
resulting in changes in cell phenotype, even among cells of the same lineage (22, 135).
Because of differences in the molecular arrangement throughout these liver
microstructures, one can infer that variations in stiffness also exist. A clinical study
supported this concept by measuring stiffness of liver sections in separate locations.
Results showed differing measurements between lobes of same liver and among smaller
areas within each lobe (136). Organ and tissue physiology show that the slightest
differences in mechanical properties can alter function of adult cells.
Studies have additionally shown the abilities of stiffness to direct stem cells
towards specific cell lineages. In a previous study, mesenchymal stem cells (MSCs) were
differentiated on varying substrate stiffnesses into three different cell lineages;
neurogenic, myogenic, and osteogenic cells (137). A separate group’s analysis revealed
that effects of growth factors on MSCs also changed with substrate stiffness. TGF-β
differentiated MSCs into smooth muscle cells (SMCs) at a low stiffness while
upregulating expression of chondrogenic markers at a higher stiffness level (138). By
36
simply changing this mechanical property, cell development and growth was directed to
mimic cells of a certain tissue type.
Cells not only change function but also migrate according to stiffness gradients.
In previous assays, MSCs under cell culture conditions traveled to areas of higher
substrate stiffness. The hypothesis stated that the cells migrate toward an injury-related
stiffness or a more scar-like site in order to aid in repair (139, 140). This phenomenon
called durotaxis has been observed in several cell types and is thought to be related to
mechanisms of development and wound healing (141). These studies show that physical
cues are just as important as chemical stimuli in determining cell arrangement, activation,
and phenotype.
6.2 Cellular Stiffness
Cell stiffness is directly correlated to the mechanical properties of the substrate to which
it is attached. This material could be the underlying ECM, a tissue culture biomaterial, or
a neighboring cell. Cell rigidity also depends on the cell lineage type, the cell’s maturity,
and the functional state or health of the cell (142). Cell stiffness ranges from softer
epithelial cells to stiffer smooth muscle cells and rigid osteocytes. Stem cells and
progenitor cells adapt their physical structures as they mature and differentiate (143,
144).
Certain disease and repair conditions can direct mechanical properties. Cells adapt
to dynamic physical environments and can change their activation states accordingly.
During the development of atherosclerosis, endothelial cells increase in stiffness which,
along with plaque build-up, limits flow of oxygen-rich blood (145, 146). Valvular
37
interstitial cells activate and become fibroblastic in diseased valves. They return to
quiescent states following repair and remodeling (147). In cancer, cells with decreased
stiffness have the highest metastatic potential (148). This concept holds true in ovarian
cancer cells where stiffness can be used as a biomarker for invasiveness (142).
Mechanical properties regulate cell-cell and cell-matrix interactions. Stiffness of
cells and the ECM can have independent or cumulative effects that cause systemic
imbalances and disease. Cells and matrix can also interact to balance physical conditions
and maintain normal tissue homeostasis and function. Determining methods to achieve a
healthy stiffness state in tissue will lead to effective treatments to correct disorders or
diseases in patients.
7. Liver ECM Stiffness during Developmental or Repair States
Stiffness is not a static property in liver tissue but fluctuates with organ
development, disease, and repair. In early liver development, a low stiffness environment
maintains progenitor cells in their undifferentiated form in the endoderm. As ECM
structures are created, cytokine and environmental cues eventually initiate cell migration
and differentiation of these cells into mature parenchymal liver cells (149, 150). These
observations are supported by in vitro studies which have shown the ability of low
stiffness substrates to maintain expression of stem cell markers in hepatic progenitor cells
(151).
Normal liver repair or regeneration is also initiated by elements of stiffness.
Matrix-producing hepatic stellate cells and endothelial cells are activated by damage or
increases in stiffness and migrate to these areas for repair (59). In these situations, MMPs
38
are required to degrade proteolytic-resistant collagens. These MMPs break-up or remove
damaged or excess ECM to allow formation of new matrix structures (37, 152). Physical
properties of the ECM as well as cell signaling enable new cells to repopulate these
repaired sites to restore normal tissue function (58).
In the instance of tissue resection such as a partial hepatectomy, the lost ECM
mass is fully regenerated de novo with the help of the activated matrix-producing cells.
The provisional matrix produced during early stages of repair or regeneration contains
uncrosslinked collagen I (58). In progenitor cell-mediated regeneration, this softer
composition initially maintains the hepatoblast phenotype of cells until ECM structures
are fully formed with the addition of collagen IV and the crosslinking of collagen I (153).
Regeneration initiated by proliferating hepatocytes occurs simultaneously to matrix
production by HSCs. However, physical cues provided by the ECM contribute to control
of the initiation and cessation of cell propagation (31, 35, 154).
Imbalances in these reparative processes can cause increases in stiffness and liver
dysfunction. Liver stiffness is an important parameter in the prognosis of liver diseases
including cirrhosis, hepatitis, and hepatocellular carcinoma (HCC) (155, 156). During a
fibrotic state, HSCs undergo uncontrolled activation and deposit excess amounts of
laminin and collagen, especially collagen IV (43, 59). Fibrosis also occurs because of a
loss of MMP production and increase in collagen crosslinking by lysyl oxidases (LOX)
(157). All these factors contribute to an increase in stiffness and lead to a loss of
hepatocyte phenotype and cell death (151). The liver is typically able to repair itself if the
causative stresses are eliminated. However, prolonged fibrosis can cause worsening
levels of stiffness and cellular impairment which eventually lead to irreversible cirrhosis
39
and liver failure (43, 130, 151). Elevated liver stiffness is also a precursor for the
development of HCC (158). It can predict both the appearance and metastatic potential of
tumors. Increased stiffness correlates with increased proliferation of cancer cells and
greater resistance to chemotherapeutic agents (151).
Novel research and treatments for liver disease attempt to address the mechanisms
that cause changes in tissue stiffness. Therapeutic strategies include inhibition of
activated stellate cells to limit ECM accumulation and developing agents to block LOX
to block collagen crosslinking. Groups are also investigating methods to better control the
release and action of a variety of MMPs to enable better manipulation of matrix
degradation to soften fibrotic tissue. Studies have shown that once cells are returned to a
normal stiffness environment, full function and health can be restored. Obtaining better
control over ECM stiffness levels will lead to greater management of liver disease.
8. Effects of Stiffness on Mechanotransduction and Focal Adhesion Signaling
8.1 Cell Adhesion Molecules
Cells sense mechanical properties of their microenvironment through cell
attachment initiated by integrins, syndecans, or other glycoproteins (159). Integrins are
the most common cell adhesion molecules (CAMs) and are composed of both an alpha
and beta subunit (160). These structural components determine the integrin’s specificity
to certain ECM molecules including collagens, fibronectin, laminin, or vitronectin (161).
Integrin engagement is vital for survival and normal phenotype of all epithelial cell types
(162). These transmembrane structures initiate signal transduction by formation of focal
adhesions that connect the ECM with the cytoskeleton. This mechanical force-induced
40
signaling allows cells to sense and interact with the surrounding substrate and affects cell
proliferation, motility, and function (163, 164).
Other adhesion molecules also have roles in regulating cell behavior. Syndecans
are transmembrane proteins that are linked to GAG chains of heparan sulfate and
chondroitin sulfate (165). Syndecans, with the aid of these GAGs, bind and concentrate
important growth factors, such as the liver HGF and EGF (166, 167). Although not as
prevalent as integrins, these proteoglycans also form anchoring junctions and can bind to
collagens and fibronectin of the ECM to support cell to matrix attachment. In addition,
syndecans can act alongside certain integrins to promote cell-cell adhesion (159).
Cadherins are a large family of glycoproteins that are the main effectors of cell-
cell binding. They provide connections between actin filaments of various cells to help
determine morphology and motility (168). Several classes of these calcium dependent
structures exist. Expression of E-cadherin (epithelial-cadherin) is specifically vital to
epithelial cell viability and polarity. E-cadherin forms homophilic cell-cell junctions
between hepatocytes within the liver and is critical to tissue formation (168, 169). Loss of
E-cadherin in vivo has been correlated with loss of cell phenotype and the development of
metastases in patients (170).
Tight junctions are another type of intercellular complex important to epithelial
cell adhesion, cell polarity, and function (171, 172). These structures prevent leaking of
material between cells and allow for direct diffusion and active transport of molecules
and ions between cells (173). Claudins, especially claudin-1, are required for structural
maintenance of tight junctions (174). Although not an integral structural component, the
41
tight junction protein occludin is a central protein involved in cell polarity and directional
migration. Occludin enables organization of actin filaments in response to stimuli that
lead to formation of cell protrusions and cell movement (175).
8.2 Mechanotransduction and Cytoskeletal Regulation
Cell adhesion molecules are fundamental for sensing the physical characteristics
of adjacent cells and the ECM or underlying cell culture material. Although all CAMs are
important to normal cellular functions, integrins are the molecules that are essential for
detecting substrate stiffness (176). Bound integrins develop small focal adhesion
complexes and stimulate assembly of more mature focal adhesions through recruitment
of kinases and adaptor proteins to enable cellular mechanotransduction (177, 178).
Cytoplasmic proteins serve as mediators to amplify or modify the signals generated from
integrin attachment. Signals are passed bidirectionally through either inside-out or
outside-in activation to affect cell phenotype and create changes in the surrounding ECM
microenvironment (179, 180).
During cell attachment, several different focal adhesion molecules are recruited to
interact at the cytoplasmic domains of the integrins. One of the first proteins to bind to
the integrin tail regions is talin. Talin is a key component of focal adhesions that
functionally activates integrins and determines their affinity for specific ligands (181).
Talin recruits the adaptor proteins paxillin and vinculin to focal adhesion complexes to
regulate the actin cytoskeleton (182). These linkages generate strain between integrins
and cytoskeletal structures to allow cells to sense substrate mechanical properties, which
in turn direct morphology and motility (183). Vinculin localizes to focal adhesion sites as
42
well as cadherin junctions. It directly connects talin to actin filaments within the cell.
Presence of vinculin is required for cell attachment, cell spreading, and filopodia
formation (184-186). Paxillin is another scaffold component that joins with talin to
control focal adhesion stability and turnover. It also interacts with different intracellular
kinases to regulate assembly of focal adhesion complexes and organization of the
cytoskeleton (183).
Focal adhesion kinase (FAK) is one of the main kinases that co-localizes with
integrins and affects their activation (179, 183). FAK has an integral role in
mechanotransduction. In previous studies utilizing FAK negative fibroblasts, mechanical
stiffness did not initiate normal cellular responses, and cell migration tendencies were
blocked (187). To initiate responses, FAK interacts with Src tyrosine kinase as well as
adaptor proteins and dozens of other focal adhesion signaling molecules (164). FAK
contains C-terminal and N-terminal domains that specify activity and transduction of
signals. The C-terminal domain allows FAK to migrate, attach, and disrupt focal
adhesion sites, while the N-terminus binds and regulates localization of ligands such as
paxillin (188, 189). Integrin attachment activates FAK by inducing kinase clustering and
autophosphorylation. This action subsequently allows Src to bind, activate, and further
phosphorylate FAK at various amino acid residues (164, 190).
Depending on the site targeted, phosphorylation of FAK affects a wide variety of
proteins and intracellular pathways to execute different and sometimes opposing
functions. FAK is important in cell viability and stimulates phosphoinositide 3-kinase
(PI3K) mediated upregulation of protein kinase B (AKT) (191). FAK also supports
survival by serving as a scaffold for phosphorylation of Crk-associated substrate (CAS)
43
which activates c-Jun N-terminal kinases (JNK), mitogen-activated protein kinase 1
(MAPK1), and mitogen-activated protein kinase 3 (MAPK3) (192). Previous studies have
shown the ability to prevent anoikis, or detachment related apoptosis, in epithelial cells
through FAK expression (193, 194).
In addition to influencing viability and proliferation, activated FAK in
combination with Src can phosphorylate talin and lead to the disassembly of focal
adhesion complexes (195, 196). Under unique conditions, FAK has also been observed to
strengthen integrin attachment and enhance focal adhesion formation (197, 198).
Differences in results of these studies could be explained by the ability of FAK to
stimulate at least four separate pathways of actin assembly and cell motility (199). PI3K
and CAS pathways are not only FAK targets implicated in cell survival, but also
stimulate cell spreading and migration (164, 200). Both PI3K and CAS serve as
regulators of the downstream Ras homolog family member (Rho) GTPase, Ras-related
C3 botulinum toxin substrate (Rac) (192, 201). FAK also regulates function of the
adaptor protein growth factor receptor-bound 7 (GRB7) as well as neural Wiskott-
Aldrich syndrome protein (N-WASP) (202). N-WASP controls actin polymerization and
remodeling through the actin-related protein (Arp) 2/3 complex and another type of Rho
GTPase, cell division control protein 42 (CDC42) (164, 199). Studies have also shown
that FAK can actually suppress a third type of Rho GTPase, Ras homolog family member
A (RhoA), to promote focal adhesion turnover (203).
FAK regulates Rho GTPases that are vital to focal adhesion assembly and
disassembly, actin organization, and ultimately cell motility. The three main Rho
GTPases (Rac1, CDC42, and RhoA) work in concert with each other to initiate cell
44
migration (204). However, cells seeded on varying ECM molecules or stiffnesses can
produce activation profiles of the Rac1 and CDC42 that differ from RhoA in timing or
expression levels. In research on fibroblasts seeded on fibronectin, Rac1 and CDC42
were stimulated at early time points post-seeding, while RhoA was activated much later
in vitro (199). These previous studies show that FAK activation occurs at various
phosphorylation sites or affects different pathways to control the two sets of Rho
GTPases (205).
Rac1 and CDC42 stimulate actin polymerization and extension of cellular
processes for movement. Rac1 is directly involved in formation of lamellipodia at the
leading edge of polarized cells (206, 207). Besides activation by FAK, paxillin can bind
and stimulate Rac1 activity. CDC42 binds and activates n-WASP to create thin
projections called filopodia past the frontal cellular boundary (183). Both GTPases and
FAK are required for durotaxis and other forms of directional migration controlled by
feedback signaling mechanisms (208). Lower substrate stiffness levels have been shown
to support turnover of focal adhesion sites, easier alteration of cell morphology, and
simpler cell detachment for motility (199).
RhoA can act in opposition to cell movement by increasing the assembly and
formation of mature focal adhesions and contractile actin stress fibers through stimulation
of Rho-associated protein kinase (ROCK). Increases in RhoA correlate with increases in
stiffness and create greater cytoskeletal organization, actin polarization, and stabilization
and anchorage of cell structures (209). However, overexpression of RhoA can actually
lead to build-up of stress fibers and EMT which promotes the development of cancer
(210, 211). In contrast to formation of stable adhesions, RhoA is also essential to
45
generate traction force for migration on substrates. Cells can initiate stronger traction
forces on stiffer or inflexible surfaces (209). The tradeoff is that greater stiffness in
substrates creates more stable focal adhesions that make cell detachment more difficult
(199). Therefore, an intermediate, substrate stiffness is optimal for traction and speed for
cell motility.
In addition to FAK’s effects on Rho GTPase expression, integrin-linked kinase
(ILK) can act in parallel to influence cell morphology and motility (212). ILK recruits
PINCH1 to focal adhesions to regulate assembly and disassembly during cell migration.
ILK also complexes with Engulfment and Cell Motility 2 (ELMO2) and RhoG for cell
polarization (213). Similar to FAK, ILK can stimulate PI3K-induced activation of Rac1
to control cytoskeletal rearrangement and formation of cellular processes (214, 215).
As discussed, mechanical sensing and signaling involves a complex array of focal
adhesion sites, intracellular kinases, and Rho GTPases. These various molecules act in
conjunction to affect the actin cytoskeleton and form cellular protrusions. The outside-in
signaling and resultant inside-out adjustments applied by the cell can affect morphology,
motility, and ultimately the overall function of the cell. Understanding the mechanisms
and actions of important effectors like FAK, ILK, and Rho GTPases in specific cell types
under defined culture conditions will allow researchers to better model disease and more
precisely manipulate substrates for use in testing or therapies.
46
Works Cited
(216)
1. Martini F, Bartholomew EF. Essentials of anatomy & physiology. 4th ed. San Francisco: Pearson/Benjamin Cummings; 2007. 2. Sherwood L, Cengage Learning (Firm). Human physiology : from cells to systems. 7th ed. Australia ; United States: Brooks/Cole, Cengage Learning; 2010. 3. Ganong's review of medical physiology. New York: McGraw-Hill Medical; 2010. 4. Gropper SS. Advanced nutrition and human metabolism. 6th Ed. ed. Belmont, OH: Cengage Learning; 2012. 5. Chatterjea MN, Shinde R. Textbook of medical biochemistry. 8th ed. New Delhi: Jaypee Brothers Medical Publications (P) Ltd.; 2012. xvii, 876 p. p. 6. LeCluyse EL, Witek RP, Andersen ME, Powers MJ. Organotypic liver culture models: meeting current challenges in toxicity testing. Crit Rev Toxicol. 2012;42(6):501-48. doi: 10.3109/10408444.2012.682115. PubMed PMID: 22582993; PMCID: 3423873. 7. Tzanakakis ES, Hess DJ, Sielaff TD, Hu WS. Extracorporeal tissue engineered liver-assist devices. Annu Rev Biomed Eng. 2000;2:607-32. doi: 10.1146/annurev.bioeng.2.1.607. PubMed PMID: 11701525. 8. Xu C, Li CY, Kong AN. Induction of phase I, II and III drug metabolism/transport by xenobiotics. Arch Pharm Res. 2005;28(3):249-68. PubMed PMID: 15832810. 9. Lautt WW. Hepatic Circulation: Physiology and Pathophysiology. San Rafael (CA)2009. 10. Malarkey DE, Johnson K, Ryan L, Boorman G, Maronpot RR. New insights into functional aspects of liver morphology. Toxicol Pathol. 2005;33(1):27-34. doi: 10.1080/01926230590881826. PubMed PMID: 15805053. 11. Reichen J. The Role of the Sinusoidal Endothelium in Liver Function. News Physiol Sci. 1999;14:117-21. PubMed PMID: 11390834. 12. Wingerd BD. The human body : concepts of anatomy and physiology. Third edition. ed. Philadelphia: Wolters Kluwer/Lippincott Williams & Wilkins; 2014. xvi, 528 pages p. 13. Lanza RP, Atala A. Essentials of stem cell biology. Second edition. ed. Amsterdam: Elsevier/Academic Press; 2014. xxviii, 680 pages p.
Fig.2) Integrin Signaling and Regulation of Rho GTPases, partially adapted from Mayor et al (209)
47
14. Mescher AL. Junqueira's Basic Pathology: Text & Atlas. 12 ed: McGraw-Hill Education 2010. 15. Berry MN, Edwards AM. The hepatocyte review. Dordrecht ; Boston: Kluwer Academic Publishers; 2000. xvi, 605 p. p. 16. Jungermann K, Stumpel F. Role of hepatic, intrahepatic and hepatoenteral nerves in the regulation of carbohydrate metabolism and hemodynamics of the liver and intestine. Hepatogastroenterology. 1999;46 Suppl 2:1414-7. PubMed PMID: 10431702. 17. Treyer A, Musch A. Hepatocyte polarity. Compr Physiol. 2013;3(1):243-87. doi: 10.1002/cphy.c120009. PubMed PMID: 23720287; PMCID: 3697931. 18. Arias IM. The liver : biology and pathobiology. 5th ed. Chichester, West Sussex, UK ; Hoboken, NJ: John Wiley & Sons; 2009. p. p. 19. Warren A, Le Couteur DG, Fraser R, Bowen DG, McCaughan GW, Bertolino P. T lymphocytes interact with hepatocytes through fenestrations in murine liver sinusoidal endothelial cells. Hepatology. 2006;44(5):1182-90. doi: 10.1002/hep.21378. PubMed PMID: 17058232. 20. Fu D, Wakabayashi Y, Ido Y, Lippincott-Schwartz J, Arias IM. Regulation of bile canalicular network formation and maintenance by AMP-activated protein kinase and LKB1. J Cell Sci. 2010;123(Pt 19):3294-302. doi: 10.1242/jcs.068098. PubMed PMID: 20826460; PMCID: 2939801. 21. Gumucio JJ. Hepatocyte heterogeneity: the coming of age from the description of a biological curiosity to a partial understanding of its physiological meaning and regulation. Hepatology. 1989;9(1):154-60. PubMed PMID: 2642292. 22. McClelland R, Wauthier E, Uronis J, Reid L. Gradients in the liver's extracellular matrix chemistry from periportal to pericentral zones: influence on human hepatic progenitors. Tissue Eng Part A. 2008;14(1):59-70. doi: 10.1089/ten.a.2007.0058. PubMed PMID: 18333805. 23. Bacon BR, Di Bisceglie AM. Hepatitis C virus infection. N Engl J Med. 2001;345(19):1425-6; author reply 7-8. PubMed PMID: 11794183. 24. Dufour J-Fo, Clavien P-A. Signaling pathways in liver diseases. 2nd ed. Heidelberg ; New York: Springer; 2010. xv, 526 p. p. 25. Katz NR. Metabolic heterogeneity of hepatocytes across the liver acinus. J Nutr. 1992;122(3 Suppl):843-9. PubMed PMID: 1542056. 26. Schiff ER, Maddrey WC, Sorrell MF, Schiff L. Schiff's diseases of the liver. 11th ed. Chichester, West Sussex, UK: John Wiley & Sons; 2011. p. p. 27. Hailfinger S, Jaworski M, Braeuning A, Buchmann A, Schwarz M. Zonal gene expression in murine liver: lessons from tumors. Hepatology. 2006;43(3):407-14. doi: 10.1002/hep.21082. PubMed PMID: 16496347. 28. Helling TS. Liver failure following partial hepatectomy. HPB (Oxford). 2006;8(3):165-74. doi: 10.1080/13651820510035712. PubMed PMID: 18333270; PMCID: 2131683. 29. Michalopoulos GK. Liver regeneration. J Cell Physiol. 2007;213(2):286-300. doi: 10.1002/jcp.21172. PubMed PMID: 17559071; PMCID: 2701258. 30. Geerts A. History, heterogeneity, developmental biology, and functions of quiescent hepatic stellate cells. Semin Liver Dis. 2001;21(3):311-35. doi: 10.1055/s-2001-17550. PubMed PMID: 11586463. 31. Haussinger D. Liver regeneration. Berlin: de Gruyter; 2011. xvii, 210 p. p. 32. Friedman SL. Hepatic stellate cells: protean, multifunctional, and enigmatic cells of the liver. Physiol Rev. 2008;88(1):125-72. doi: 10.1152/physrev.00013.2007. PubMed PMID: 18195085; PMCID: 2888531.
48
33. Jiang F, Parsons CJ, Stefanovic B. Gene expression profile of quiescent and activated rat hepatic stellate cells implicates Wnt signaling pathway in activation. J Hepatol. 2006;45(3):401-9. doi: 10.1016/j.jhep.2006.03.016. PubMed PMID: 16780995. 34. Coll M, Taghdouini AE, Perea L, Mannaerts I, Vila-Casadesus M, Blaya D, Rodrigo-Torres D, Affo S, Morales-Ibanez O, Graupera I, Lozano JJ, Najimi M, Sokal E, Lambrecht J, Gines P, van Grunsven LA, Sancho-Bru P. Integrative miRNA and Gene Expression Profiling Analysis of Human Quiescent Hepatic Stellate Cells. Sci Rep. 2015;5:11549. doi: 10.1038/srep11549. PubMed PMID: 26096707; PMCID: 4476106. 35. Yin C, Evason KJ, Asahina K, Stainier DY. Hepatic stellate cells in liver development, regeneration, and cancer. The Journal of clinical investigation. 2013;123(5):1902-10. doi: 10.1172/JCI66369. PubMed PMID: 23635788; PMCID: 3635734. 36. Wang P, Liu T, Cong M, Wu X, Bai Y, Yin C, An W, Wang B, Jia J, You H. Expression of extracellular matrix genes in cultured hepatic oval cells: an origin of hepatic stellate cells through transforming growth factor beta? Liver Int. 2009;29(4):575-84. doi: 10.1111/j.1478-3231.2009.01992.x. PubMed PMID: 19323784. 37. Iredale JP, Thompson A, Henderson NC. Extracellular matrix degradation in liver fibrosis: Biochemistry and regulation. Biochim Biophys Acta. 2013;1832(7):876-83. doi: 10.1016/j.bbadis.2012.11.002. PubMed PMID: 23149387. 38. Kikuchi S, Griffin CT, Wang SS, Bissell DM. Role of CD44 in epithelial wound repair: migration of rat hepatic stellate cells utilizes hyaluronic acid and CD44v6. J Biol Chem. 2005;280(15):15398-404. doi: 10.1074/jbc.M414048200. PubMed PMID: 15691832. 39. Ishibashi H, Nakamura M, Komori A, Migita K, Shimoda S. Liver architecture, cell function, and disease. Semin Immunopathol. 2009;31(3):399-409. doi: 10.1007/s00281-009-0155-6. PubMed PMID: 19468732. 40. Johnson LR. Physiology of the gastrointestinal tract. 4th ed. Burlington, MA: Elsevier Academic Press; 2006. p. p. 41. Willekens FL, Werre JM, Kruijt JK, Roerdinkholder-Stoelwinder B, Groenen-Dopp YA, van den Bos AG, Bosman GJ, van Berkel TJ. Liver Kupffer cells rapidly remove red blood cell-derived vesicles from the circulation by scavenger receptors. Blood. 2005;105(5):2141-5. doi: 10.1182/blood-2004-04-1578. PubMed PMID: 15550489. 42. Braet F, Wisse E. Structural and functional aspects of liver sinusoidal endothelial cell fenestrae: a review. Comp Hepatol. 2002;1(1):1. PubMed PMID: 12437787; PMCID: 131011. 43. Wells RG. Cellular sources of extracellular matrix in hepatic fibrosis. Clin Liver Dis. 2008;12(4):759-68, viii. doi: 10.1016/j.cld.2008.07.008. PubMed PMID: 18984465; PMCID: 2617723. 44. Kuntz E. Hepatology, textbook and atlas. 3rd ed. New York: Springer; 2008. 45. Cogger VC, Roessner U, Warren A, Fraser R, Le Couteur DG. A Sieve-Raft Hypothesis for the regulation of endothelial fenestrations. Comput Struct Biotechnol J. 2013;8:e201308003. doi: 10.5936/csbj.201308003. PubMed PMID: 24688743; PMCID: 3962122. 46. Tabibian JH, Masyuk AI, Masyuk TV, O'Hara SP, LaRusso NF. Physiology of cholangiocytes. Compr Physiol. 2013;3(1):541-65. doi: 10.1002/cphy.c120019. PubMed PMID: 23720296; PMCID: 3831353. 47. Strazzabosco M, Fabris L. Functional anatomy of normal bile ducts. Anat Rec (Hoboken). 2008;291(6):653-60. doi: 10.1002/ar.20664. PubMed PMID: 18484611; PMCID: 3743051. 48. Meadows GG. Integration/interaction of oncologic growth. Dordrecht: Springer; 2005. xix, 454 p. p. 49. Rodes J. Textbook of hepatology : from basic science to clinical practice. 3rd ed. Malden, Mass.: Blackwell; 2007.
49
50. Zern MA, Reid LM. Extracellular matrix : chemistry, biology, and pathobiology with emphasis on the liver. New York: Dekker; 1993. xiv, 591 p. p. 51. Rubin K, Borg TK, Holmdahl R, Klareskog L, Obrink B. Interactions of mammalian cells with collagen. Ciba Found Symp. 1984;108:93-116. PubMed PMID: 6569833. 52. Bedossa P, Paradis V. Liver extracellular matrix in health and disease. J Pathol. 2003;200(4):504-15. doi: DOI 10.1002/path.1397. PubMed PMID: WOS:000183925700010. 53. Pankov R, Yamada KM. Fibronectin at a glance. J Cell Sci. 2002;115(Pt 20):3861-3. PubMed PMID: 12244123. 54. Clark K, Pankov R, Travis MA, Askari JA, Mould AP, Craig SE, Newham P, Yamada KM, Humphries MJ. A specific alpha5beta1-integrin conformation promotes directional integrin translocation and fibronectin matrix formation. J Cell Sci. 2005;118(Pt 2):291-300. doi: 10.1242/jcs.01623. PubMed PMID: 15615773; PMCID: 3329624. 55. Wierzbicka-Patynowski I, Schwarzbauer JE. The ins and outs of fibronectin matrix assembly. J Cell Sci. 2003;116(Pt 16):3269-76. doi: 10.1242/jcs.00670. PubMed PMID: 12857786. 56. Moriya K, Bae E, Honda K, Sakai K, Sakaguchi T, Tsujimoto I, Kamisoyama H, Keene DR, Sasaki T, Sakai T. A fibronectin-independent mechanism of collagen fibrillogenesis in adult liver remodeling. Gastroenterology. 2011;140(5):1653-63. doi: 10.1053/j.gastro.2011.02.005. PubMed PMID: 21320502; PMCID: 3081910. 57. Boyer JL. Liver cirrhosis and its development : proceedings of the Falk Symposium 115 held in Basel, Switzerland, 22-24 October, 1999 (part II of the Basel Liver Week 1999; XI International Congress of Liver Diseases). Dordrecht ; Boston: Kluwer Academic; 2001. xii, 354 p. p. 58. Kim TH, Mars WM, Stolz DB, Petersen BE, Michalopoulos GK. Extracellular matrix remodeling at the early stages of liver regeneration in the rat. Hepatology. 1997;26(4):896-904. doi: 10.1002/hep.510260415. PubMed PMID: 9328311. 59. Martinez-Hernandez A, Amenta PS. The extracellular matrix in hepatic regeneration. Faseb J. 1995;9(14):1401-10. PubMed PMID: 7589981. 60. Taniguchi N. Glycoscience : biology and medicine. New York: Springer; 2014. pages cm p. 61. Taylor KR, Gallo RL. Glycosaminoglycans and their proteoglycans: host-associated molecular patterns for initiation and modulation of inflammation. Faseb J. 2006;20(1):9-22. doi: 10.1096/fj.05-4682rev. PubMed PMID: 16394262. 62. Esko JD, Kimata K, Lindahl U. Proteoglycans and Sulfated Glycosaminoglycans. In: Varki A, Cummings RD, Esko JD, Freeze HH, Stanley P, Bertozzi CR, Hart GW, Etzler ME, editors. Essentials of Glycobiology. 2nd ed. Cold Spring Harbor (NY)2009. 63. Hileman RE, Smith AE, Toida T, Linhardt RJ. Preparation and structure of heparin lyase-derived heparan sulfate oligosaccharides. Glycobiology. 1997;7(2):231-9. PubMed PMID: 9134430. 64. Esko JD, Linhardt RJ. Proteins that Bind Sulfated Glycosaminoglycans. In: Varki A, Cummings RD, Esko JD, Freeze HH, Stanley P, Bertozzi CR, Hart GW, Etzler ME, editors. Essentials of Glycobiology. 2nd ed. Cold Spring Harbor (NY)2009. 65. Zern BJ, Chu H, Wang Y. Control growth factor release using a self-assembled [polycation:heparin] complex. PLoS One. 2010;5(6):e11017. doi: 10.1371/journal.pone.0011017. PubMed PMID: 20543985; PMCID: 2882380. 66. Shute J. Glycosaminoglycan and chemokine/growth factor interactions. Handb Exp Pharmacol. 2012(207):307-24. doi: 10.1007/978-3-642-23056-1_13. PubMed PMID: 22566230. 67. Berry MN, Grivell AR, Grivell MB, Phillips JW. Isolated hepatocytes--past, present and future. Cell Biol Toxicol. 1997;13(4-5):223-33. PubMed PMID: 9298243.
50
68. Fromm JR, Hileman RE, Caldwell EE, Weiler JM, Linhardt RJ. Differences in the interaction of heparin with arginine and lysine and the importance of these basic amino acids in the binding of heparin to acidic fibroblast growth factor. Arch Biochem Biophys. 1995;323(2):279-87. doi: 10.1006/abbi.1995.9963. PubMed PMID: 7487089. 69. Munoz EM, Linhardt RJ. Heparin-binding domains in vascular biology. Arterioscler Thromb Vasc Biol. 2004;24(9):1549-57. doi: 10.1161/01.ATV.0000137189.22999.3f. PubMed PMID: 15231514; PMCID: 4114236. 70. Dumitriu S, Popa VI. Polymeric biomaterials. Boca Raton: CRC Press, Taylor & Francis Group; 2013. v. <1-2> p. 71. Lyon M, Deakin JA, Rahmoune H, Fernig DG, Nakamura T, Gallagher JT. Hepatocyte growth factor/scatter factor binds with high affinity to dermatan sulfate. J Biol Chem. 1998;273(1):271-8. PubMed PMID: 9417075. 72. Sarrazin S, Lamanna WC, Esko JD. Heparan sulfate proteoglycans. Cold Spring Harb Perspect Biol. 2011;3(7). doi: 10.1101/cshperspect.a004952. PubMed PMID: 21690215; PMCID: 3119907. 73. McCourt PA. How does the hyaluronan scrap-yard operate? Matrix Biol. 1999;18(5):427-32. PubMed PMID: 10601730. 74. Steinhoff G. Regenerative medicine : from protocol to patient. New York: Springer; 2013. 75. Murphy SV, Skardal A, Atala A. Evaluation of hydrogels for bio-printing applications. J Biomed Mater Res A. 2013;101(1):272-84. doi: 10.1002/jbm.a.34326. PubMed PMID: 22941807. 76. Jones JI, Gockerman A, Busby WH, Jr., Camacho-Hubner C, Clemmons DR. Extracellular matrix contains insulin-like growth factor binding protein-5: potentiation of the effects of IGF-I. J Cell Biol. 1993;121(3):679-87. PubMed PMID: 7683690; PMCID: 2119570. 77. Hudalla GA, Murphy WL. Biomaterials that regulate growth factor activity via bioinspired interactions. Adv Funct Mater. 2011;21(10):1754-68. doi: 10.1002/adfm.201002468. PubMed PMID: 21921999; PMCID: 3171147. 78. King WJ, Krebsbach PH. Growth factor delivery: how surface interactions modulate release in vitro and in vivo. Advanced drug delivery reviews. 2012;64(12):1239-56. doi: 10.1016/j.addr.2012.03.004. PubMed PMID: 22433783; PMCID: 3586795. 79. Bonnans C, Chou J, Werb Z. Remodelling the extracellular matrix in development and disease. Nat Rev Mol Cell Biol. 2014;15(12):786-801. doi: 10.1038/nrm3904. PubMed PMID: 25415508; PMCID: 4316204. 80. LeCouter J, Moritz DR, Li B, Phillips GL, Liang XH, Gerber HP, Hillan KJ, Ferrara N. Angiogenesis-independent endothelial protection of liver: role of VEGFR-1. Science. 2003;299(5608):890-3. doi: 10.1126/science.1079562. PubMed PMID: 12574630. 81. DeLeve LD. Liver sinusoidal endothelial cells and liver regeneration. The Journal of clinical investigation. 2013;123(5):1861-6. doi: 10.1172/JCI66025. PubMed PMID: 23635783; PMCID: 3635729. 82. Pediaditakis P, Monga SPS, Mars WM, Michalopoulos GK. Differential mitogenic effects of single chain hepatocyte growth factor (HGF)/scatter factor and HGF/NK1 following cleavage by factor Xa. Journal of Biological Chemistry. 2002;277(16):14109-15. doi: DOI 10.1074/jbc.M112196200. PubMed PMID: WOS:000175096000099. 83. Naldini L, Vigna E, Narsimhan RP, Gaudino G, Zarnegar R, Michalopoulos GK, Comoglio PM. Hepatocyte growth factor (HGF) stimulates the tyrosine kinase activity of the receptor encoded by the proto-oncogene c-MET. Oncogene. 1991;6(4):501-4. PubMed PMID: 1827664.
51
84. Matsumoto K, Nakamura T. Hepatocyte growth factor (HGF) as a tissue organizer for organogenesis and regeneration. Biochem Bioph Res Co. 1997;239(3):639-44. PubMed PMID: WOS:A1997YG93100001. 85. Kim SH, Akaike T. Epidermal growth factor signaling for matrix-dependent cell proliferation and differentiation in primary cultured hepatocytes. Tissue Eng. 2007;13(3):601-9. doi: 10.1089/ten.2006.0104. PubMed PMID: 17518606. 86. Steiger-Luther NC, Darwiche H, Oh SH, Williams JM, Petersen BE. Insulin-like growth factor binding protein-3 is required for the regulation of rat oval cell proliferation and differentiation in the 2AAF/PHX model. Hepatic medicine : evidence and research. 2010;2010(2):13-32. doi: 10.2147/HMER.S7660. PubMed PMID: 21852899; PMCID: 3156464. 87. Kim M, Lee JY, Jones CN, Revzin A, Tae G. Heparin-based hydrogel as a matrix for encapsulation and cultivation of primary hepatocytes. Biomaterials. 2010;31(13):3596-603. doi: 10.1016/j.biomaterials.2010.01.068. PubMed PMID: 20153045; PMCID: 2837121. 88. Sellaro TL, Ranade A, Faulk DM, McCabe GP, Dorko K, Badylak SF, Strom SC. Maintenance of human hepatocyte function in vitro by liver-derived extracellular matrix gels. Tissue Eng Part A. 2010;16(3):1075-82. doi: 10.1089/ten.TEA.2008.0587. PubMed PMID: 19845461; PMCID: 2863084. 89. Ghaedi M, Tuleuova N, Zern MA, Wu J, Revzin A. Bottom-up signaling from HGF-containing surfaces promotes hepatic differentiation of mesenchymal stem cells. Biochem Biophys Res Commun. 2011;407(2):295-300. doi: 10.1016/j.bbrc.2011.03.005. PubMed PMID: 21382341; PMCID: 3075384. 90. Stolz DB, Mars WM, Petersen BE, Kim TH, Michalopoulos GK. Growth factor signal transduction immediately after two-thirds partial hepatectomy in the rat. Cancer Res. 1999;59(16):3954-60. PubMed PMID: 10463591. 91. Bohm F, Kohler UA, Speicher T, Werner S. Regulation of liver regeneration by growth factors and cytokines. EMBO Mol Med. 2010;2(8):294-305. doi: 10.1002/emmm.201000085. PubMed PMID: 20652897; PMCID: 3377328. 92. Ishikawa H, Jo JI, Tabata Y. Liver Anti-Fibrosis Therapy with Mesenchymal Stem Cells Secreting Hepatocyte Growth Factor. Journal of biomaterials science Polymer edition. 2011. doi: 10.1163/156856211X614761. PubMed PMID: 22182291. 93. Ichihara A. BCA, HGF, and proteasomes. Biochem Biophys Res Commun. 1999;266(3):647-51. doi: 10.1006/bbrc.1999.1882. PubMed PMID: 10603302. 94. Xue F, Takahara T, Yata Y, Kuwabara Y, Shinno E, Nonome K, Minemura M, Takahara S, Li X, Yamato E, Watanabe A. Hepatocyte growth factor gene therapy accelerates regeneration in cirrhotic mouse livers after hepatectomy. Gut. 2003;52(5):694-700. PubMed PMID: 12692055; PMCID: 1773642. 95. Bottaro DP, Rubin JS, Faletto DL, Chan AM, Kmiecik TE, Vande Woude GF, Aaronson SA. Identification of the hepatocyte growth factor receptor as the c-met proto-oncogene product. Science. 1991;251(4995):802-4. PubMed PMID: 1846706. 96. Rubin JS, Day RM, Breckenridge D, Atabey N, Taylor WG, Stahl SJ, Wingfield PT, Kaufman JD, Schwall R, Bottaro DP. Dissociation of heparan sulfate and receptor binding domains of hepatocyte growth factor reveals that heparan sulfate-c-met interaction facilitates signaling. J Biol Chem. 2001;276(35):32977-83. doi: 10.1074/jbc.M105486200. PubMed PMID: 11435444. 97. Liu KX, Kato Y, Kato M, Kaku TI, Nakamura T, Sugiyama Y. Existence of two nonlinear elimination mechanisms for hepatocyte growth factor in rats. Am J Physiol. 1997;273(5 Pt 1):E891-7. PubMed PMID: 9374673. 98. Sakata H, Stahl SJ, Taylor WG, Rosenberg JM, Sakaguchi K, Wingfield PT, Rubin JS. Heparin binding and oligomerization of hepatocyte growth factor/scatter factor isoforms.
52
Heparan sulfate glycosaminoglycan requirement for Met binding and signaling. J Biol Chem. 1997;272(14):9457-63. PubMed PMID: 9083085. 99. Schuppan D, Schmid M, Somasundaram R, Ackermann R, Ruehl M, Nakamura T, Riecken EO. Collagens in the liver extracellular matrix bind hepatocyte growth factor. Gastroenterology. 1998;114(1):139-52. PubMed PMID: 9428228. 100. Orlando RC. Pathophysiology of gastroesophageal reflux disease. J Clin Gastroenterol. 2008;42(5):584-8. doi: 10.1097/MCG.0b013e31815d0628. PubMed PMID: 18364588. 101. Francavilla A, Ove P, Polimeno L, Sciascia C, Coetzee ML, Starzl TE. Epidermal growth factor and proliferation in rat hepatocytes in primary culture isolated at different times after partial hepatectomy. Cancer Res. 1986;46(3):1318-23. PubMed PMID: 3002614; PMCID: 2963441. 102. Mullhaupt B, Feren A, Fodor E, Jones A. Liver expression of epidermal growth factor RNA. Rapid increases in immediate-early phase of liver regeneration. J Biol Chem. 1994;269(31):19667-70. PubMed PMID: 7519599. 103. Mullhaupt B, Feren A, Jones A, Fodor E. DNA sequence and functional characterization of the human and rat epidermal growth factor promoter: regulation by cell growth. Gene. 2000;250(1-2):191-200. PubMed PMID: 10854792. 104. Monga SP. Role of Wnt/beta-catenin signaling in liver metabolism and cancer. Int J Biochem Cell Biol. 2011;43(7):1021-9. doi: 10.1016/j.biocel.2009.09.001. PubMed PMID: 19747566; PMCID: 2891959. 105. Ornitz DM, Itoh N. Fibroblast growth factors. Genome Biol. 2001;2(3):REVIEWS3005. PubMed PMID: 11276432; PMCID: 138918. 106. Cao R, Brakenhielm E, Pawliuk R, Wariaro D, Post MJ, Wahlberg E, Leboulch P, Cao Y. Angiogenic synergism, vascular stability and improvement of hind-limb ischemia by a combination of PDGF-BB and FGF-2. Nat Med. 2003;9(5):604-13. doi: 10.1038/nm848. PubMed PMID: 12669032. 107. Papp V, Dezso K, Laszlo V, Nagy P, Paku S. Architectural changes during regenerative and ontogenic liver growth in the rat. Liver Transpl. 2009;15(2):177-83. doi: 10.1002/lt.21665. PubMed PMID: 19177433. 108. Wynn TA, Barron L. Macrophages: master regulators of inflammation and fibrosis. Semin Liver Dis. 2010;30(3):245-57. doi: 10.1055/s-0030-1255354. PubMed PMID: 20665377; PMCID: 2924662. 109. Barrett KE. Epithelial biology in the gastrointestinal system: insights into normal physiology and disease pathogenesis. J Physiol. 2012;590(Pt 3):419-20. doi: 10.1113/jphysiol.2011.227058. PubMed PMID: 22298901; PMCID: 3379689. 110. Michalopoulos GK. Liver regeneration after partial hepatectomy: critical analysis of mechanistic dilemmas. Am J Pathol. 2010;176(1):2-13. doi: 10.2353/ajpath.2010.090675. PubMed PMID: 20019184; PMCID: 2797862. 111. Choi SS, Diehl AM. Epithelial-to-mesenchymal transitions in the liver. Hepatology. 2009;50(6):2007-13. doi: 10.1002/hep.23196. PubMed PMID: 19824076; PMCID: 2787916. 112. Boulton R, Woodman A, Calnan D, Selden C, Tam F, Hodgson H. Nonparenchymal cells from regenerating rat liver generate interleukin-1alpha and -1beta: a mechanism of negative regulation of hepatocyte proliferation. Hepatology. 1997;26(1):49-58. doi: 10.1053/jhep.1997.v26.pm0009214451. PubMed PMID: 9214451. 113. Makino H, Shimizu H, Ito H, Kimura F, Ambiru S, Togawa A, Ohtsuka M, Yoshidome H, Kato A, Yoshitomi H, Sawada S, Miyazaki M. Changes in growth factor and cytokine expression in biliary obstructed rat liver and their relationship with delayed liver regeneration after partial
53
hepatectomy. World J Gastroentero. 2006;12(13):2053-9. PubMed PMID: WOS:000239996100012. 114. Thenappan A, Li Y, Kitisin K, Rashid A, Shetty K, Johnson L, Mishra L. Role of transforming growth factor beta signaling and expansion of progenitor cells in regenerating liver. Hepatology. 2010;51(4):1373-82. doi: 10.1002/hep.23449. PubMed PMID: 20131405; PMCID: 3001243. 115. Fausto N. Liver regeneration and repair: hepatocytes, progenitor cells, and stem cells. Hepatology. 2004;39(6):1477-87. doi: 10.1002/hep.20214. PubMed PMID: 15185286. 116. Pintilie DG, Shupe TD, Oh SH, Salganik SV, Darwiche H, Petersen BE. Hepatic stellate cells' involvement in progenitor-mediated liver regeneration. Laboratory investigation; a journal of technical methods and pathology. 2010;90(8):1199-208. doi: 10.1038/labinvest.2010.88. PubMed PMID: 20440274; PMCID: 2912420. 117. Dezso K, Papp V, Bugyik E, Hegyesi H, Safrany G, Bodor C, Nagy P, Paku S. Structural analysis of oval-cell-mediated liver regeneration in rats. Hepatology. 2012;56(4):1457-67. doi: 10.1002/hep.25713. PubMed PMID: 22419534. 118. Pi L, Ding X, Jorgensen M, Pan JJ, Oh SH, Pintilie D, Brown A, Song WY, Petersen BE. Connective tissue growth factor with a novel fibronectin binding site promotes cell adhesion and migration during rat oval cell activation. Hepatology. 2008;47(3):996-1004. doi: 10.1002/hep.22079. PubMed PMID: 18167060; PMCID: 3130595. 119. Sicklick JK, Choi SS, Bustamante M, McCall SJ, Perez EH, Huang J, Li YX, Rojkind M, Diehl AM. Evidence for epithelial-mesenchymal transitions in adult liver cells. Am J Physiol Gastrointest Liver Physiol. 2006;291(4):G575-83. doi: 10.1152/ajpgi.00102.2006. PubMed PMID: 16710052. 120. Washington MK. Autoimmune liver disease: overlap and outliers. Mod Pathol. 2007;20 Suppl 1:S15-30. doi: 10.1038/modpathol.3800684. PubMed PMID: 17486048. 121. Kelly DA. Diseases of the liver and biliary system in children. 3rd ed. Chichester, UK ; Hoboken, NJ: Wiley-Blackwell; 2008. x, 630 p. p. 122. Marcdante KJ, Kliegman R. Nelson essentials of pediatrics. 7th edition. ed. Philadelphia, PA: Elsevier/Saunders; 2015. xxii, 754 pages p. 123. Hoffmann GF, Zschocke J, Nyhan WL. Inherited metabolic diseases : a clinical approach. Heidelberg: Springer; 2010. xiv, 386 p. p. 124. Lazaridis KN, LaRusso NF. The Cholangiopathies. Mayo Clin Proc. 2015;90(6):791-800. doi: 10.1016/j.mayocp.2015.03.017. PubMed PMID: 25957621; PMCID: 4533104. 125. Lazaridis KN, Strazzabosco M, Larusso NF. The cholangiopathies: disorders of biliary epithelia. Gastroenterology. 2004;127(5):1565-77. PubMed PMID: 15521023. 126. Xu R, Zhang Z, Wang FS. Liver fibrosis: mechanisms of immune-mediated liver injury. Cell Mol Immunol. 2012;9(4):296-301. doi: 10.1038/cmi.2011.53. PubMed PMID: 22157623; PMCID: 4132585. 127. Benjamin IJ, Griggs RC, Wing EJ, Fitz JG. Andreoli and Carpenter's Cecil essentials of medicine. 9th edition. ed. Philadelphia, PA: Elsevier/Saunders; 2015. p. p. 128. Rockey DC. Hepatic fibrosis, stellate cells, and portal hypertension. Clin Liver Dis. 2006;10(3):459-79, vii-viii. doi: 10.1016/j.cld.2006.08.017. PubMed PMID: 17162223. 129. Salameh A, Dhein S. Effects of mechanical forces and stretch on intercellular gap junction coupling. Biochim Biophys Acta. 2013;1828(1):147-56. doi: 10.1016/j.bbamem.2011.12.030. PubMed PMID: 22245380. 130. Wells RG. The role of matrix stiffness in regulating cell behavior. Hepatology. 2008;47(4):1394-400. doi: 10.1002/hep.22193. PubMed PMID: 18307210. 131. Baumgart E. Stiffness--an unknown world of mechanical science? Injury. 2000;31 Suppl 2:S-B14-23. PubMed PMID: 10853758.
54
132. Ma PX. Biomaterials and regenerative medicine. Cambridge, United Kingdom: Cambridge University Press; 2014. xvi, 703 pages p. 133. Mason BN, Starchenko A, Williams RM, Bonassar LJ, Reinhart-King CA. Tuning three-dimensional collagen matrix stiffness independently of collagen concentration modulates endothelial cell behavior. Acta biomaterialia. 2013;9(1):4635-44. doi: 10.1016/j.actbio.2012.08.007. PubMed PMID: 22902816; PMCID: 3508162. 134. Bhatia SK. Engineering biomaterials for regenerative medicine : novel technologies for clinical applications. New York, NY: Springer; 2012. x, 352 p. p. 135. Reid LM, Fiorino AS, Sigal SH, Brill S, Holst PA. Extracellular matrix gradients in the space of Disse: relevance to liver biology. Hepatology. 1992;15(6):1198-203. PubMed PMID: 1592356. 136. Rusak G, Zawada E, Lemanowicz A, Serafin Z. Whole-organ and segmental stiffness measured with liver magnetic resonance elastography in healthy adults: significance of the region of interest. Abdom Imaging. 2015;40(4):776-82. doi: 10.1007/s00261-014-0278-7. PubMed PMID: 25331569; PMCID: 4372679. 137. Engler AJ, Sen S, Sweeney HL, Discher DE. Matrix elasticity directs stem cell lineage specification. Cell. 2006;126(4):677-89. doi: 10.1016/j.cell.2006.06.044. PubMed PMID: 16923388. 138. Park JS, Chu JS, Tsou AD, Diop R, Tang Z, Wang A, Li S. The effect of matrix stiffness on the differentiation of mesenchymal stem cells in response to TGF-beta. Biomaterials. 2011;32(16):3921-30. doi: 10.1016/j.biomaterials.2011.02.019. PubMed PMID: 21397942; PMCID: 3073995. 139. Raab M, Swift J, Dingal PC, Shah P, Shin JW, Discher DE. Crawling from soft to stiff matrix polarizes the cytoskeleton and phosphoregulates myosin-II heavy chain. J Cell Biol. 2012;199(4):669-83. doi: 10.1083/jcb.201205056. PubMed PMID: 23128239; PMCID: 3494847. 140. Tse JR, Engler AJ. Stiffness gradients mimicking in vivo tissue variation regulate mesenchymal stem cell fate. PLoS One. 2011;6(1):e15978. doi: 10.1371/journal.pone.0015978. PubMed PMID: 21246050; PMCID: 3016411. 141. Lo CM, Wang HB, Dembo M, Wang YL. Cell movement is guided by the rigidity of the substrate. Biophysical Journal. 2000;79(1):144-52. PubMed PMID: WOS:000088048500011. 142. Xu W, Mezencev R, Kim B, Wang L, McDonald J, Sulchek T. Cell stiffness is a biomarker of the metastatic potential of ovarian cancer cells. PLoS One. 2012;7(10):e46609. doi: 10.1371/journal.pone.0046609. PubMed PMID: 23056368; PMCID: 3464294. 143. Tee SY, Fu J, Chen CS, Janmey PA. Cell shape and substrate rigidity both regulate cell stiffness. Biophys J. 2011;100(5):L25-7. doi: 10.1016/j.bpj.2010.12.3744. PubMed PMID: 21354386; PMCID: 3043219. 144. Wang YK, Chen CS. Cell adhesion and mechanical stimulation in the regulation of mesenchymal stem cell differentiation. J Cell Mol Med. 2013;17(7):823-32. doi: 10.1111/jcmm.12061. PubMed PMID: 23672518; PMCID: 3741348. 145. Holzapfel GA, Ogden RW. Mechanics of biological tissue. Berlin ; New York: Springer; 2006. xii, 522 p. p. 146. Brown XQ, Bartolak-Suki E, Williams C, Walker ML, Weaver VM, Wong JY. Effect of substrate stiffness and PDGF on the behavior of vascular smooth muscle cells: implications for atherosclerosis. J Cell Physiol. 2010;225(1):115-22. doi: 10.1002/jcp.22202. PubMed PMID: 20648629; PMCID: 2920297. 147. Rabkin-Aikawa E, Farber M, Aikawa M, Schoen FJ. Dynamic and reversible changes of interstitial cell phenotype during remodeling of cardiac valves. J Heart Valve Dis. 2004;13(5):841-7. PubMed PMID: 15473488.
55
148. Swaminathan V, Mythreye K, O'Brien ET, Berchuck A, Blobe GC, Superfine R. Mechanical stiffness grades metastatic potential in patient tumor cells and in cancer cell lines. Cancer Res. 2011;71(15):5075-80. doi: 10.1158/0008-5472.CAN-11-0247. PubMed PMID: 21642375; PMCID: 3220953. 149. Si-Tayeb K, Lemaigre FP, Duncan SA. Organogenesis and development of the liver. Dev Cell. 2010;18(2):175-89. doi: 10.1016/j.devcel.2010.01.011. PubMed PMID: 20159590. 150. Si-Tayeb K, Noto FK, Nagaoka M, Li J, Battle MA, Duris C, North PE, Dalton S, Duncan SA. Highly efficient generation of human hepatocyte-like cells from induced pluripotent stem cells. Hepatology. 2010;51(1):297-305. doi: 10.1002/hep.23354. PubMed PMID: 19998274; PMCID: 2946078. 151. Schrader J, Gordon-Walker TT, Aucott RL, van Deemter M, Quaas A, Walsh S, Benten D, Forbes SJ, Wells RG, Iredale JP. Matrix stiffness modulates proliferation, chemotherapeutic response, and dormancy in hepatocellular carcinoma cells. Hepatology. 2011;53(4):1192-205. doi: 10.1002/hep.24108. PubMed PMID: 21442631; PMCID: 3076070. 152. Han YP. Matrix metalloproteinases, the pros and cons, in liver fibrosis. J Gastroenterol Hepatol. 2006;21 Suppl 3:S88-91. doi: 10.1111/j.1440-1746.2006.04586.x. PubMed PMID: 16958682; PMCID: 2646497. 153. Zhang L, Theise N, Chua M, Reid LM. The stem cell niche of human livers: symmetry between development and regeneration. Hepatology. 2008;48(5):1598-607. doi: 10.1002/hep.22516. PubMed PMID: 18972441. 154. Kordes C, Sawitza I, Gotze S, Herebian D, Haussinger D. Hepatic stellate cells contribute to progenitor cells and liver regeneration. The Journal of clinical investigation. 2014;124(12):5503-15. doi: 10.1172/JCI74119. PubMed PMID: 25401473; PMCID: 4348953. 155. Jung KS, Kim SU, Ahn SH, Park YN, Kim do Y, Park JY, Chon CY, Choi EH, Han KH. Risk assessment of hepatitis B virus-related hepatocellular carcinoma development using liver stiffness measurement (FibroScan). Hepatology. 2011;53(3):885-94. doi: 10.1002/hep.24121. PubMed PMID: 21319193. 156. Tsukuma H, Hiyama T, Tanaka S, Nakao M, Yabuuchi T, Kitamura T, Nakanishi K, Fujimoto I, Inoue A, Yamazaki H, et al. Risk factors for hepatocellular carcinoma among patients with chronic liver disease. N Engl J Med. 1993;328(25):1797-801. doi: 10.1056/NEJM199306243282501. PubMed PMID: 7684822. 157. Cox TR, Bird D, Baker AM, Barker HE, Ho MW, Lang G, Erler JT. LOX-mediated collagen crosslinking is responsible for fibrosis-enhanced metastasis. Cancer Res. 2013;73(6):1721-32. doi: 10.1158/0008-5472.CAN-12-2233. PubMed PMID: 23345161; PMCID: 3672851. 158. Cox TR, Erler JT. Remodeling and homeostasis of the extracellular matrix: implications for fibrotic diseases and cancer. Dis Model Mech. 2011;4(2):165-78. doi: 10.1242/dmm.004077. PubMed PMID: 21324931; PMCID: 3046088. 159. Morgan MR, Humphries MJ, Bass MD. Synergistic control of cell adhesion by integrins and syndecans. Nat Rev Mol Cell Biol. 2007;8(12):957-69. doi: 10.1038/nrm2289. PubMed PMID: 17971838; PMCID: 3329926. 160. Hynes RO. Integrins: bidirectional, allosteric signaling machines. Cell. 2002;110(6):673-87. PubMed PMID: 12297042. 161. Humphries JD, Byron A, Humphries MJ. Integrin ligands at a glance. J Cell Sci. 2006;119(Pt 19):3901-3. doi: 10.1242/jcs.03098. PubMed PMID: 16988024; PMCID: 3380273. 162. Taddei ML, Giannoni E, Fiaschi T, Chiarugi P. Anoikis: an emerging hallmark in health and diseases. J Pathol. 2012;226(2):380-93. doi: 10.1002/path.3000. PubMed PMID: 21953325.
56
163. Hynes RO, Lively JC, McCarty JH, Taverna D, Francis SE, Hodivala-Dilke K, Xiao Q. The diverse roles of integrins and their ligands in angiogenesis. Cold Spring Harb Symp Quant Biol. 2002;67:143-53. PubMed PMID: 12858535. 164. Kim SH, Turnbull J, Guimond S. Extracellular matrix and cell signalling: the dynamic cooperation of integrin, proteoglycan and growth factor receptor. J Endocrinol. 2011;209(2):139-51. doi: 10.1530/JOE-10-0377. PubMed PMID: 21307119. 165. Lambaerts K, Wilcox-Adelman SA, Zimmermann P. The signaling mechanisms of syndecan heparan sulfate proteoglycans. Curr Opin Cell Biol. 2009;21(5):662-9. doi: 10.1016/j.ceb.2009.05.002. PubMed PMID: 19535238; PMCID: 2758656. 166. Derksen PW, Keehnen RM, Evers LM, van Oers MH, Spaargaren M, Pals ST. Cell surface proteoglycan syndecan-1 mediates hepatocyte growth factor binding and promotes Met signaling in multiple myeloma. Blood. 2002;99(4):1405-10. PubMed PMID: 11830493. 167. Ramani VC, Yang Y, Ren Y, Nan L, Sanderson RD. Heparanase plays a dual role in driving hepatocyte growth factor (HGF) signaling by enhancing HGF expression and activity. J Biol Chem. 2011;286(8):6490-9. doi: 10.1074/jbc.M110.183277. PubMed PMID: 21131364; PMCID: 3057851. 168. Halbleib JM, Nelson WJ. Cadherins in development: cell adhesion, sorting, and tissue morphogenesis. Genes Dev. 2006;20(23):3199-214. doi: 10.1101/gad.1486806. PubMed PMID: 17158740. 169. van Roy F, Berx G. The cell-cell adhesion molecule E-cadherin. Cell Mol Life Sci. 2008;65(23):3756-88. doi: 10.1007/s00018-008-8281-1. PubMed PMID: 18726070. 170. Onder TT, Gupta PB, Mani SA, Yang J, Lander ES, Weinberg RA. Loss of E-cadherin promotes metastasis via multiple downstream transcriptional pathways. Cancer Res. 2008;68(10):3645-54. doi: 10.1158/0008-5472.CAN-07-2938. PubMed PMID: 18483246. 171. Anderson JM, Van Itallie CM. Physiology and function of the tight junction. Cold Spring Harb Perspect Biol. 2009;1(2):a002584. doi: 10.1101/cshperspect.a002584. PubMed PMID: 20066090; PMCID: 2742087. 172. Shin K, Fogg VC, Margolis B. Tight junctions and cell polarity. Annual review of cell and developmental biology. 2006;22:207-35. doi: 10.1146/annurev.cellbio.22.010305.104219. PubMed PMID: 16771626. 173. Hirase T, Kawashima S, Wong EY, Ueyama T, Rikitake Y, Tsukita S, Yokoyama M, Staddon JM. Regulation of tight junction permeability and occludin phosphorylation by Rhoa-p160ROCK-dependent and -independent mechanisms. J Biol Chem. 2001;276(13):10423-31. doi: 10.1074/jbc.M007136200. PubMed PMID: 11139571. 174. Osanai M, Murata M, Nishikiori N, Chiba H, Kojima T, Sawada N. Occludin-mediated premature senescence is a fail-safe mechanism against tumorigenesis in breast carcinoma cells. Cancer Sci. 2007;98(7):1027-34. doi: 10.1111/j.1349-7006.2007.00494.x. PubMed PMID: 17459053. 175. Du D, Xu F, Yu L, Zhang C, Lu X, Yuan H, Huang Q, Zhang F, Bao H, Jia L, Wu X, Zhu X, Zhang X, Zhang Z, Chen Z. The tight junction protein, occludin, regulates the directional migration of epithelial cells. Dev Cell. 2010;18(1):52-63. doi: 10.1016/j.devcel.2009.12.008. PubMed PMID: 20152177. 176. Burridge K, Chrzanowska-Wodnicka M. Focal adhesions, contractility, and signaling. Annual review of cell and developmental biology. 1996;12:463-518. doi: 10.1146/annurev.cellbio.12.1.463. PubMed PMID: 8970735. 177. Zaidel-Bar R, Cohen M, Addadi L, Geiger B. Hierarchical assembly of cell-matrix adhesion complexes. Biochem Soc Trans. 2004;32(Pt3):416-20. doi: 10.1042/BST0320416. PubMed PMID: 15157150.
57
178. Wu C. Focal adhesion: a focal point in current cell biology and molecular medicine. Cell Adh Migr. 2007;1(1):13-8. PubMed PMID: 19262093; PMCID: 2633675. 179. Barczyk M, Carracedo S, Gullberg D. Integrins. Cell Tissue Res. 2010;339(1):269-80. doi: 10.1007/s00441-009-0834-6. PubMed PMID: 19693543; PMCID: 2784866. 180. Bae YH, Mui KL, Hsu BY, Liu SL, Cretu A, Razinia Z, Xu T, Pure E, Assoian RK. A FAK-Cas-Rac-lamellipodin signaling module transduces extracellular matrix stiffness into mechanosensitive cell cycling. Sci Signal. 2014;7(330):ra57. doi: 10.1126/scisignal.2004838. PubMed PMID: 24939893; PMCID: 4345117. 181. Tadokoro S, Shattil SJ, Eto K, Tai V, Liddington RC, de Pereda JM, Ginsberg MH, Calderwood DA. Talin binding to integrin beta tails: a final common step in integrin activation. Science. 2003;302(5642):103-6. doi: 10.1126/science.1086652. PubMed PMID: 14526080. 182. Wang P, Ballestrem C, Streuli CH. The C terminus of talin links integrins to cell cycle progression. J Cell Biol. 2011;195(3):499-513. doi: 10.1083/jcb.201104128. PubMed PMID: 22042621; PMCID: 3206343. 183. Mitra SK, Hanson DA, Schlaepfer DD. Focal adhesion kinase: in command and control of cell motility. Nat Rev Mol Cell Biol. 2005;6(1):56-68. doi: 10.1038/nrm1549. PubMed PMID: 15688067. 184. Xu W, Baribault H, Adamson ED. Vinculin knockout results in heart and brain defects during embryonic development. Development. 1998;125(2):327-37. PubMed PMID: 9486805. 185. Carisey A, Tsang R, Greiner AM, Nijenhuis N, Heath N, Nazgiewicz A, Kemkemer R, Derby B, Spatz J, Ballestrem C. Vinculin regulates the recruitment and release of core focal adhesion proteins in a force-dependent manner. Current biology : CB. 2013;23(4):271-81. doi: 10.1016/j.cub.2013.01.009. PubMed PMID: 23375895; PMCID: 3580286. 186. Humphries JD, Wang P, Streuli C, Geiger B, Humphries MJ, Ballestrem C. Vinculin controls focal adhesion formation by direct interactions with talin and actin. J Cell Biol. 2007;179(5):1043-57. doi: 10.1083/jcb.200703036. PubMed PMID: 18056416; PMCID: 2099183. 187. Wang HB, Dembo M, Hanks SK, Wang Y. Focal adhesion kinase is involved in mechanosensing during fibroblast migration. Proc Natl Acad Sci U S A. 2001;98(20):11295-300. doi: 10.1073/pnas.201201198. PubMed PMID: 11572981; PMCID: 58723. 188. Bertolucci CM, Guibao CD, Zheng J. Structural features of the focal adhesion kinase-paxillin complex give insight into the dynamics of focal adhesion assembly. Protein Sci. 2005;14(3):644-52. doi: 10.1110/ps.041107205. PubMed PMID: 15689512; PMCID: 2279287. 189. Scheswohl DM, Harrell JR, Rajfur Z, Gao G, Campbell SL, Schaller MD. Multiple paxillin binding sites regulate FAK function. J Mol Signal. 2008;3:1. doi: 10.1186/1750-2187-3-1. PubMed PMID: 18171471; PMCID: 2246129. 190. Westhoff MA, Serrels B, Fincham VJ, Frame MC, Carragher NO. SRC-mediated phosphorylation of focal adhesion kinase couples actin and adhesion dynamics to survival signaling. Mol Cell Biol. 2004;24(18):8113-33. doi: 10.1128/MCB.24.18.8113-8133.2004. PubMed PMID: 15340073; PMCID: 515031. 191. Hanks SK, Ryzhova L, Shin NY, Brabek J. Focal adhesion kinase signaling activities and their implications in the control of cell survival and motility. Front Biosci. 2003;8:d982-96. PubMed PMID: 12700132. 192. Cho SY, Klemke RL. Extracellular-regulated kinase activation and CAS/Crk coupling regulate cell migration and suppress apoptosis during invasion of the extracellular matrix. J Cell Biol. 2000;149(1):223-36. PubMed PMID: 10747099; PMCID: 2175095. 193. Sonoda Y, Matsumoto Y, Funakoshi M, Yamamoto D, Hanks SK, Kasahara T. Anti-apoptotic role of focal adhesion kinase (FAK). Induction of inhibitor-of-apoptosis proteins and
58
apoptosis suppression by the overexpression of FAK in a human leukemic cell line, HL-60. J Biol Chem. 2000;275(21):16309-15. PubMed PMID: 10821872. 194. Kurenova E, Xu LH, Yang X, Baldwin AS, Jr., Craven RJ, Hanks SK, Liu ZG, Cance WG. Focal adhesion kinase suppresses apoptosis by binding to the death domain of receptor-interacting protein. Mol Cell Biol. 2004;24(10):4361-71. PubMed PMID: 15121855; PMCID: 400455. 195. Hamadi A, Bouali M, Dontenwill M, Stoeckel H, Takeda K, Ronde P. Regulation of focal adhesion dynamics and disassembly by phosphorylation of FAK at tyrosine 397. J Cell Sci. 2005;118(Pt 19):4415-25. doi: 10.1242/jcs.02565. PubMed PMID: 16159962. 196. Nagano M, Hoshino D, Koshikawa N, Akizawa T, Seiki M. Turnover of focal adhesions and cancer cell migration. Int J Cell Biol. 2012;2012:310616. doi: 10.1155/2012/310616. PubMed PMID: 22319531; PMCID: 3272802. 197. Kwong L, Wozniak MA, Collins AS, Wilson SD, Keely PJ. R-Ras promotes focal adhesion formation through focal adhesion kinase and p130(Cas) by a novel mechanism that differs from integrins. Mol Cell Biol. 2003;23(3):933-49. PubMed PMID: 12529399; PMCID: 140691. 198. Wozniak MA, Modzelewska K, Kwong L, Keely PJ. Focal adhesion regulation of cell behavior. Biochim Biophys Acta. 2004;1692(2-3):103-19. doi: 10.1016/j.bbamcr.2004.04.007. PubMed PMID: 15246682. 199. Li S, Guan JL, Chien S. Biochemistry and biomechanics of cell motility. Annu Rev Biomed Eng. 2005;7:105-50. doi: 10.1146/annurev.bioeng.7.060804.100340. PubMed PMID: 16004568. 200. Kim SH, Kim SH. Antagonistic effect of EGF on FAK phosphorylation/dephosphorylation in a cell. Cell Biochem Funct. 2008;26(5):539-47. doi: 10.1002/cbf.1457. PubMed PMID: 18543351. 201. Welch HC, Coadwell WJ, Stephens LR, Hawkins PT. Phosphoinositide 3-kinase-dependent activation of Rac. FEBS Lett. 2003;546(1):93-7. PubMed PMID: 12829242. 202. Rhoads DS, Guan JL. Analysis of directional cell migration on defined FN gradients: role of intracellular signaling molecules. Exp Cell Res. 2007;313(18):3859-67. doi: 10.1016/j.yexcr.2007.06.005. PubMed PMID: 17640633; PMCID: 2083117. 203. Ren XD, Kiosses WB, Sieg DJ, Otey CA, Schlaepfer DD, Schwartz MA. Focal adhesion kinase suppresses Rho activity to promote focal adhesion turnover. J Cell Sci. 2000;113 ( Pt 20):3673-8. PubMed PMID: 11017882. 204. Sakumura Y, Tsukada Y, Yamamoto N, Ishii S. A molecular model for axon guidance based on cross talk between rho GTPases. Biophys J. 2005;89(2):812-22. doi: 10.1529/biophysj.104.055624. PubMed PMID: 15923236; PMCID: 1366631. 205. Chen R, Kim O, Li M, Xiong X, Guan JL, Kung HJ, Chen H, Shimizu Y, Qiu Y. Regulation of the PH-domain-containing tyrosine kinase Etk by focal adhesion kinase through the FERM domain. Nat Cell Biol. 2001;3(5):439-44. doi: 10.1038/35074500. PubMed PMID: 11331870. 206. Kurokawa K, Itoh RE, Yoshizaki H, Nakamura YO, Matsuda M. Coactivation of Rac1 and Cdc42 at lamellipodia and membrane ruffles induced by epidermal growth factor. Mol Biol Cell. 2004;15(3):1003-10. doi: 10.1091/mbc.E03-08-0609. PubMed PMID: 14699061; PMCID: 363057. 207. Ballestrem C, Wehrle-Haller B, Hinz B, Imhof BA. Actin-dependent lamellipodia formation and microtubule-dependent tail retraction control-directed cell migration. Mol Biol Cell. 2000;11(9):2999-3012. PubMed PMID: 10982396; PMCID: 14971. 208. Hsia DA, Mitra SK, Hauck CR, Streblow DN, Nelson JA, Ilic D, Huang S, Li E, Nemerow GR, Leng J, Spencer KS, Cheresh DA, Schlaepfer DD. Differential regulation of cell motility and invasion by FAK. J Cell Biol. 2003;160(5):753-67. doi: 10.1083/jcb.200212114. PubMed PMID: 12615911; PMCID: 2173366.
59
209. Tojkander S, Gateva G, Lappalainen P. Actin stress fibers--assembly, dynamics and biological roles. J Cell Sci. 2012;125(Pt 8):1855-64. doi: 10.1242/jcs.098087. PubMed PMID: 22544950. 210. Bhowmick NA, Ghiassi M, Bakin A, Aakre M, Lundquist CA, Engel ME, Arteaga CL, Moses HL. Transforming growth factor-beta1 mediates epithelial to mesenchymal transdifferentiation through a RhoA-dependent mechanism. Mol Biol Cell. 2001;12(1):27-36. PubMed PMID: 11160820; PMCID: 30565. 211. Reffay M, Parrini MC, Cochet-Escartin O, Ladoux B, Buguin A, Coscoy S, Amblard F, Camonis J, Silberzan P. Interplay of RhoA and mechanical forces in collective cell migration driven by leader cells. Nat Cell Biol. 2014;16(3):217-23. doi: 10.1038/ncb2917. PubMed PMID: 24561621. 212. Khyrul WA, LaLonde DP, Brown MC, Levinson H, Turner CE. The integrin-linked kinase regulates cell morphology and motility in a rho-associated kinase-dependent manner. J Biol Chem. 2004;279(52):54131-9. doi: 10.1074/jbc.M410051200. PubMed PMID: 15485819. 213. Ho E, Irvine T, Vilk GJ, Lajoie G, Ravichandran KS, D'Souza SJ, Dagnino L. Integrin-linked kinase interactions with ELMO2 modulate cell polarity. Mol Biol Cell. 2009;20(13):3033-43. doi: 10.1091/mbc.E09-01-0050. PubMed PMID: 19439446; PMCID: 2704155. 214. Qian Y, Zhong X, Flynn DC, Zheng JZ, Qiao M, Wu C, Dedhar S, Shi X, Jiang BH. ILK mediates actin filament rearrangements and cell migration and invasion through PI3K/Akt/Rac1 signaling. Oncogene. 2005;24(19):3154-65. doi: 10.1038/sj.onc.1208525. PubMed PMID: 15735674. 215. Tu Y, Huang Y, Zhang Y, Hua Y, Wu C. A new focal adhesion protein that interacts with integrin-linked kinase and regulates cell adhesion and spreading. J Cell Biol. 2001;153(3):585-98. PubMed PMID: 11331308; PMCID: 2190577. 216. Mayor R, Carmona-Fontaine C. Keeping in touch with contact inhibition of locomotion. Trends Cell Biol. 2010;20(6):319-28. doi: 10.1016/j.tcb.2010.03.005. PubMed PMID: 20399659; PMCID: 2927909.
60
CHAPTER 2
Decellularization and the Incorporation of Liver Extracellular
Matrix in Cell Culture Substrates
Daniel B. Deegan
Content
1. ECM Isolation through Decellularization Methods .………………..……61
2. Substrate Forms………………………………………………….…….….65
2.1 Decellularized Tissue Constructs…………………………...………...65
2.2 ECM Hydrogels……………………………………………...………..66
2.3 Species Differences………………………………………………..….67
3. Results and Advantages of ECM Use in Cell Culture……..………….…67
4. Altering Stiffness in Cell Culture Substrates……………………………..69
4.1 Methods for Manipulating Hydrogel Stiffness ……….…….………..69
4.2 Measuring Stiffness……………………………………………..…….73
5. Conclusions and Potential Applications…………………………………..77
61
In order to optimize cell culture conditions, substrates must contain the proper
milieu of structural components and growth factors. These molecules also must be
present in the correct ratios to grow and maintain specific cell types. The method of
decellularization solves this problem by removing native cells from tissue while
preserving the natural ECM arrangement. The byproduct is a non-immunogenic scaffold
that can be directly seeded or indirectly incorporated into tissue culture substrates and
cell therapies. In turn, these substrates can be utilized to test the effects of certain
substrate properties on cell signaling and function. Our own assays used the versatile
nature of liver ECM hydrogels in order to study the effects of mechanical stiffness on
primary hepatocyte function.
1. ECM Isolation through Decellularization Methods
A wide variety of approaches were developed to decellularize tissue or whole
organs for ECM isolation and eventual use in cell culture. Methods are dependent upon
the tissue and animal type being processed. Variations exist in the mechanical or
chemical procedures used. Protocols utilize different physical forces, compounds, and
reagents for cellular removal. Incubations in solutions and subsequent washing steps vary
in duration. These diverse procedures for decellularization have experienced a wide range
of success and failure.
Each decellularization technique possesses a set of advantages and disadvantages.
One of the first methods developed in this field ruptured cells through utilization of
distilled water and salt solutions in combination with mechanical force (1). Use of
hypertonic and hypotonic solutions to lyse cells does minimum harm to the underlying
62
ECM structure but on the other hand, does not remove smaller cellular components from
the tissue (2-8). Addition of physical disruption and agitation of tissue promotes full
clearance, however, it can also lead to damage of the intact architecture and unintended
removal of ECM constituents (9).
Another early decellularization technique implemented the use of freezing and
thawing to eliminate cells from tissue constructs (10-14). This simple method does not
require creation of solutions or the involvement of physical force. Repeated freeze thaw
effectively kills and lyses cells but can ultimately degrade certain proteins of the ECM,
especially growth factors which exhibit limited half-lives (15). Because of the sensitivity
of specific molecules and the unknown extent of ECM damage due to this process, freeze
thaw cycles should be kept to a minimum.
Newer methods incorporated varying concentrations of enzymatic reagents and
detergents into decellularization protocols. Enzymes including trypsin and DNase are
often used to treat scaffolds for cellular removal (16-24). These compounds are highly
successful in liberating cells from the ECM and break down immunogenic cellular
components such as nucleic acids (16). During natural processes in the body, apoptotic
cells trigger caspase-activated DNases which degrade DNA into 180 base pair fragments.
These nucleic acids are further degraded by DNase II of macrophages to avoid initiation
of innate immune reactions (25). For these reasons, important criteria established for
decellularization include degrading any residual DNA fragments to below 200 base pairs
(bp) in length and reducing DNA levels to below 50 ng of DNA per mg of dried tissue
(9). These actions are vital in avoiding immune responses when these substrates are
implanted into in vivo systems. Despite abilities to reduce immunoreactivity, lengthy
63
enzymatic treatments can also lead to destruction of ECM proteins and
glycosaminoglycans (GAGs) (26). Difficulties also exist in subsequent removal of
enzymes from the scaffold. Remnant enzymes could be harmful to animal models or
human patients and induce their own immune reactions.
An alternative option to enzymes for elimination of cellular material is detergents.
Previous publications have reported advantages of detergents over enzymatic methods in
maintenance of mechanical and elastic properties of the ECM (27). Detergents vary in
structure and strength, which both dictate concentrations applied and length of time
utilized. The two main types of detergents used in decellularization are ionic and non-
ionic detergents. Ionic detergents contain a charged head group and hydrophobic tail
group. This structure allows for rapid and effective disruption of the cellular membrane
and full removal of all cellular components from tissue (16, 24, 28-31). However,
because of its strength, this compound can easily denature or remove ECM molecules
such as GAGs and basement membrane proteins if not limited in concentration or
duration of use (32, 33). In comparison, non-ionic detergents are gentle on the underlying
matrix. Their hydrophilic groups are uncharged but are still able to disrupt lipid bonds
(28, 34-38). However, their mild nature means greater detergent concentrations and
longer washes are required for full cellular removal. Non-ionic detergents are often
combined with other reagents to ensure complete and timely decellularization (7, 39, 40).
The decellularization method designed by our research group and chosen for use
in this thesis project involves use of both ionic and non-ionic detergents. The
decellularization protocol has been optimized for use with livers of rat models. The
process takes advantage of the rat’s intact vasculature in order to perfuse solutions
64
Fig.1) Rat Liver Decellularization, adapted from Shupe et al (1)
A) PBS washing removes blood from the liver. B) Triton-X 100 detergent solubilizes lipid
membranes, while C) SDS detergent removes remnant cellular material.
throughout the organ. This approach enables solutions to infiltrate all areas of the tissue
for maximal contact with cells. The impact is faster and more efficient decellularization
as compared to simply immersing tissue surfaces. These developed methods allow
preservation of structural and biologically active components vital for effective growth
and maintenance of cells. For whole organ engineering, perfusion also allows for later
use of the vasculature during reseeding of cells. Through this established protocol,
decellularized liver tissue or substrates supplemented with the isolated ECM improve
liver cell viability and long term function in the cell culture assays of this thesis work.
65
2. Substrate Forms
ECM isolated from the decellularization process can be utilized in several
different configurations for many different applications. The intended objective
determines how this material must be used.
2.1 Decellularized Tissue Constructs
One of the main uses of decellularized organs and vascular constructs is for tissue
engineering. With a large disparity between donor organs and patients requiring
transplantations, the goal of organ engineering is to narrow this gap by supplying lab
grown tissues, preferably with patients’ own cells to avoid an immune response. To
engineer complex organs, one of the primary objectives is to create a fully cellularized
and patent vasculature. This task is made easier by maintaining an intact vessel basement
membrane.
Several groups have successfully generated re-endothelialized and functional
vascular grafts through seeding of decellularized scaffolds (41-46). Similar strides have
been made at a reduced scale in the recellularization of small animal organs or small
tissue samples (39, 47-49). Decellularization not only preserves the ECM composition
but also the structural architecture of the tissue. This remnant microarchitecture allows
cells to attach, align, and self-organize into proper functional units within both
parenchymal and vascular regions.
Challenges remain in whole organ engineering. Consistency of decellularized
tissue samples and the ability to control the physical and biochemical properties of the
decellularized matrix remain a problem. The exact physical and biological toll various
66
detergents take on ECM during decellularization is not fully controllable or known.
Certain key growth factors or smaller microstructures may need to be replaced or
reconstructed in order to produce an optimally functional cell scaffold. Microscopic
structures are easily damaged by decellularization reagents, including glomeruli of the
kidney, as well as bile canaliculi and sinusoids of the liver. Future studies should focus
on modifying and improving methods for preserving these delicate but important
structures. Research is still needed to optimize exact concentrations of reagents, duration
of individual steps, and storage methods for decellularized tissue. Studies also need to
gain a better understanding of complex interactions within tissue-specific ECM,
including the binding and biological activity of growth factors. New knowledge will
build upon current advances and lead the field closer to achieving functional tissue
products.
2.2 ECM Hydrogels
Hydrogels provide a versatile platform to isolate and test individual physical and
chemical properties. The flexibility of gel substrates allows for alteration of properties
such as stiffness, growth factor content, and ECM composition. These substrates can be
applied to everything from drug testing to a variety of novel therapies and treatments.
Supplementing these gels with organ-specific ECM improves attachment, growth, and
phenotype of cells similar to levels exhibited in fully functional tissue.
One of the uses of ECM hydrogels is to reproduce conditions found in vivo in order to
model certain disease or repair states. Three-dimensional cell culture in gels effectively
model tumors and metastatic phenotypes for cancer research (50, 51). High stiffness gels
67
can simulate diseases like liver cirrhosis, while lower stiffness mimics a developmental
or regenerative environment (52, 53). An upregulation of cellular production of specific
growth factors and matrix proteins also occurs during disease or regeneration (54-57).
Exogenous growth factors can be bound to active sites in ECM supplemented gels, or
ratios of specific structural molecules can be altered. This capacity for modifications
allows accurate modeling of ECM microenvironments for many tissue types and
physiological scenarios. Allows us to explore effects of composition
2.3 Species Differences
For this thesis project, decellularized rat liver tissue was integrated into HA gels
for preliminary primary rat hepatocyte studies and subsequently, primary human
hepatocyte culture. ECM components including collagens, polysaccharides, and
fibronectin are highly ubiquitous in mammalian tissue (58, 59). Molecular structures and
cell binding sites of these proteins are highly conserved between rats and humans and
interact with common cellular factors (60-62). GAGs and associated growth factors also
contain conserved structures and functions (58, 63, 64). In our own assays, we have
observed human growth factors including HGF and EGF stimulating rat cells and vice
versa. Because of their structural similarities, species differences between rat and human
liver ECM proved to be inconsequential in our assays.
3. Results and Advantages of ECM Use in Cell Culture
Traditional cell culture methods use synthetic materials or incorporate single
protein coatings like collagen into substrates for cell attachment and growth. With the
development of decellularization, a fuller representation of organ-specific ECM
68
components could be included. Several studies have shown the advantages of culturing
different cell types on natural, tissue-specific ECM substrates compared to purely
synthetic substrates. ECM from different tissue types can direct stem cells towards
various cell lineages (65-70). In liver culture systems, specific ECM cues can promote
the differentiation of hepatic progenitor cells into fully mature hepatocytes or
cholangiocytes (44, 71-74).
To develop effective and successful treatments, long term viability and phenotype
maintenance needs to be supported. Heparin and HA hydrogels are favorable substrate
bases because these molecules are naturally found in the body, do not induce immune
reactions upon implantation, and have the ability to bind and present growth factors (75,
76). We hypothesize addition of tissue-specific ECM would allow cells to function at
higher levels for use in cell based therapies. Previous studies have shown how tissue
specific ECM improves long term cell function in vitro. Both liver progenitor cells and
adult hepatocytes seeded onto isolated liver ECM maintained viability and function over
longer periods of time than cells grown on tissue culture plastic or collagen alone (44, 77-
79). Decellularized matrix has supported growth and phenotype of cells of several
different tissue types for a variety of uses (80-82). By recreating a natural
microenvironment in culture, cells are able to function and interact as they would within
normal, healthy tissue of the body.
With the addition of diverse species of GAGs present in natural ECM, greater
control over growth factor binding and release could be achieved to aid the efficiency of
cell therapies. Improvements in phenotypic maintenance were observed in cells
encapsulated with a combination of several structural molecules and growth factors,
69
especially hepatocyte growth factor (HGF) and epidermal growth factor (EGF) with
hepatocytes (83, 84). New data on the cellular effects of various substrate compositions
and properties will enable better design of cell therapies. Combining innovative methods
and knowledge will optimize their effectiveness in combating or correcting diseases in
patients.
4. Altering Stiffness in Cell Culture Substrates
4.1 Methods for Manipulating Hydrogel Stiffness
As mentioned in the previous chapter, physical properties are as important as the
presence of specific ECM components for differentiation or maintenance of various cell
types during seeding of intact tissue. Mechanical properties are modified by a wide
variety of strategies and techniques in cell culture. Previous groups have used increased
concentrations of proteins or synthetic compounds such as acrylamide to bolster substrate
stiffness (85-87). However, these alterations can influence cells in ways beyond stiffness,
and tend to complicate assay results. To isolate the property of stiffness during
experiments, the best techniques avoid addition of biologically active molecules.
One of the simplest and most common methods for modifying stiffness is through
substrate crosslinking. Crosslinking allows precise manipulation with limited
confounding effects on attachment sites or biological activity of the substrate. Stiffness
alterations can be made by varying the crosslinker type, the structure and arm length of
the crosslinker, the crosslinker concentration, or the duration of the crosslinker treatment
(88). The choice for crosslinker type is contingent on which molecules are available in
the substrate. These reagents chemically react with several different moieties including;
70
thiols, amines, carboxyls, hydroxyls, and carbonyls (89). Crosslinkers can also be
photoreactive and are activated by exposure to ultraviolet light (90, 91).
A crosslinker is either classified as homofunctional or heterofunctional, reacting
and covalently binding to one or more classes of functional groups. Homofunctional
crosslinkers act at rates and strengths dependent on the concentration of the crosslinker,
as well as the density of reactive groups in the substrate’s constituents. Greater
concentrations of the reagent cause a faster and higher degree of crosslinking. A larger
quantity of active sites in the substrate also translates to quicker bonding and stiffer
materials (92). Multiple homofunctional crosslinker types can be used in procedures to
gradually stiffen materials during activities such as bioprinting (93). Unlike
homofunctional reagents, a single heterofunctional linker type can be utilized in multiple
step reactions to produce viable substrates. Instead of combining all substrate components
at once, the crosslinker is allowed to react with one molecule before additional
compounds are added. This process provides greater control over specificity of
interactions and spacing of bonds for more exact material construction (93, 94).
Properties of the crosslinker are not only dependent on its chemical specificity but
also vary according to the length and number of structural arms it contains. A longer
distance between crosslinks creates a looser assembly of compounds and a softer
substrate. Therefore, to increase stiffness, one must use a crosslinker that is shorter in
length (92, 95). The typical reagent is bifunctional and holds two binding sites. However,
crosslinkers have been engineered to contain as many as eight arms. Greater amount of
branching leads to more reactive ends and a greater degree of binding. The geometry of
71
multi-armed crosslinkers is also more compact which generates tighter packed and stiffer
structures (96).
For this thesis project, our group chose a polyethylene glycol (PEG) based
compound for crosslinking applications during formation of liver ECM hydrogels. PEG
based chemicals are nontoxic and do not influence cell function (97). They also do not
initiate immune reactions upon implantation in vivo (98, 99). Because PEG is water-
soluble, clumping of the compound is avoided, and crosslinking is evenly spaced within
the polymer (100). PEG creates hydrophilic substrates which have been shown to
promote better cell adhesion, proliferation, spreading, and function than hydrophobic
materials (101, 102). The combination of these characteristics makes PEG an ideal
crosslinker for use in primary hepatocyte culture and development of cell treatments.
Liver ECM gels were created by mixing solubilized liver matrix with hyaluronic
acid and denatured collagen solutions. These HA gel components were specifically
modified to contain reactive thiols for crosslinking (103). Acrylate is a type of molecule
often attached to a PEG spacer to form a crosslinker that contains reactive double bonds
which bind to sulfhydryl groups. Our assays tested several geometric variations of PEG
acrylate in order to solidify the liver hydrogels at various stiffnesses.
Polyethylene glycol diacrylate (PEGDA) is one of the most commonly used
crosslinkers. It has a two-arm linear structure that produces a softer gel type (104). This
substrate stiffness was used as a baseline for our assays. PEG acrylate variations
containing four or eight arms also exist (105). More acrylate arms allow for a greater
number of reactive ends to bind thiol groups. In our own assays, PEGDA, four-arm PEG
72
acrylate, and eight-arm PEG acrylate were added to HA-ECM gel constituents at various
concentrations. Results showed that eight-arm PEG acrylate at the highest concentration
of 6% weight by volume produced the highest stiffness.
Although most combinations of crosslinker geometry and concentration produced
distinctively different stiffness levels, a saturation point of each crosslinker was also
determined. Higher concentrations of crosslinker past this limit did not significantly
affect stiffness, regardless of additional reaction time (Figure 3). This upper limit is
reached when all thiol groups are utilized. To ensure excess crosslinker is not retained in
the gel, no concentration above the observed saturation threshold was chosen for
subsequent assays. Furthermore, gels were thoroughly washed with PBS to clear out
excess crosslinker.
Fig.2) PEG Crosslinker Structures, adapted from Zhu et al (119) Greater PEG branching or arm number creates a tighter molecular network and increases
substrate stiffness.
73
4.2 Measuring Stiffness
Several methods and instrumentation were developed to measure the property of
stiffness. For centuries, the simple practice of palpation was used to diagnose patients
(106). Clinicians now utilize a non-invasive method called elastography to image and
quantitate stiffness of intact organs. This technique employs the use of ultrasound or
magnetic resonance imaging (MRI).
Several variations of ultrasound elastography evolved over time. Early forms
involved application of a manual force to the exterior of patients which was then
measured internally by ultrasound to calculate organ stiffness (107). Accuracy improved
through use of mechanical pulse generators and more advanced sonography systems.
During transient elastography, an automated device creates shear waves that penetrate
through the skin into organs of the abdominal cavity such as the liver (108, 109). The
0
200
400
600
800
1000
1200
1400
1% 2% 4%
G' (P
a)
Crosslinker Concentration % (w/v)
Crosslinked Gel Stiffness
Fig.3) PEGDA Crosslinking Saturation- HA-ECM gels were crosslinked with 1%,
2, or 4% w/v PEGDA (3.4 kDa). Only the 1% and 2% PEGDA gels were
significantly different (*p<.05, Student’s t test, n=4)
74
velocity of the waves is measured by ultrasound as they pass through the tissue of
interest. This data allows clinicians to calculate the stiffness of a large representative area
of the organ for diagnosis (110). Instead of acting on the surface of the skin, newer
adaptations employ concentrated sonography waves to generate acoustic radiation forces
at the site of measurement within the tissue (111-113). These processes improve the
distance pulses can be directed within the body. Acoustic radiation forces also increase
the precision of deep tissue ultrasound measurements through their ability to travel across
fluid build-up in the peritoneal cavity which normally impedes transient elastography
(110, 112).
Magnetic Resonance Elastography (MRE) incorporates these same mechanisms to
propagate shear waves. However, the velocity of these waves is subsequently measured
by MRI. Unlike ultrasound technology, a three-dimensional image of the entire tissue is
produced alongside a map of various stiffness values (114, 115). This tool provides a
more complete analysis of the organ being tested. MRE has proven to be a reliable tool
for detection of fibrosis in diseased liver (116). It has also successfully evaluated
stiffnesses in organs such as the brain that were unattainable by ultrasound methods
(117). Further refinement is needed to optimize the use of MRE in other tissue or organ
types (114).
In a laboratory setting, a rheometer is one of the most popular instruments used to
measure stiffness for samples prepared ex vivo. Unlike non-invasive clinical methods,
Rheometry is used to analyze biopsies or excised tissue outside of a living organism. This
technique is also commonly used in calculating stiffness of hydrogels, biomaterials, or
synthetic substrates used in cell culture. Because samples are isolated from irrelevant
75
tissues, fluids, or other confounding factors, rheometry increases accuracy and precision
of measurements compared to ultrasound and MRE (118). Engineers developed several
variations of rheometers based upon the material being tested and the type of data
required for analyses.
Rotational or shear rheometry is often used to measure stiffness in viscoelastic
soft tissue, hydrogels, or polymers (119, 120). These methods were applied in our own
assays to determine stiffness in liver ECM hydrogels and decellularized liver tissue.
During testing with these rheometers, a sample is stabilized on a fixed platform or plate.
An oscillating geometry in the form of a cone, plate, or cylinder is subsequently lowered
until it comes into contact with the sample (118). From the various forces applied, an
elastic modulus can be measured and calculated. The elastic modulus, or resistance to
deformity, can be expressed by three types of calculations; Young’s modulus, bulk
modulus and shear modulus. Young’s modulus is the ratio of tensile stress to tensile
strain or the force required to compress an object (121). The bulk modulus is the amount
of force needed to uniformly deform a material or the ratio of volumetric stress to
volumetric strain (122). The shear modulus describes how a sample responds to opposing
forces parallel to its surface and is calculated by dividing shear stress by shear strain
(123).
In the stiffness tests we performed, the Discovery Series HR-2 Rheometer (TA
Instruments, New Castle, DE) was used with a parallel plate geometry. Oscillation stress
sweeps incrementally increased shear stress on the samples up to 10.0 Pa and were run at
a constant oscillatory frequency and axial force. Both the shear storage modulus (G’) and
shear loss modulus (G’’) were measured and recorded. The storage modulus describes the
76
elastic properties or stored energy, while the loss modulus describes the viscous
characteristics or energy lost as heat (124). The storage modulus was the main stiffness
value utilized for reporting and comparison of hydrogels in our assays.
Shear rheometry provides an overall representation of stiffness in materials by
assessing a substantial portion of the substrate. For rheology measurements of smaller
microstructures or individual cells, atomic force microscopy (AFM) is required. An
atomic force microscope contains a high resolution nano-indenter or tip and detection
probe that scans indented samples to evaluate microscopic areas for stiffness (125-128).
Instead of measuring the combined effects of cell and ECM stiffness, AFM allows
separate analysis of components. Singular structures and cells in a sample can easily be
distinguished and measured to assess conditions such as cancer (129). Multiple readings
of a larger homogenous substrate can also be obtained and averaged to calculate more
wide-ranging stiffness (130, 131).
Despite AFM’s exceptional resolution, disadvantages also exist. AFM performed
on a heterogeneous sample can give an incomplete or inaccurate representation of the
material tested if too few readings are acquired or only a partial area is covered (132).
Problems can also arise depending on the indenter type used. Improper tips cause artifacts
to develop in samples (127, 131). Indenters can also adhere to softer materials and cause
miscalculations of stiffness (133).
Every analysis of stiffness presents unique objectives and obstacles. Clinicians
require a quick and noninvasive strategy, while a laboratory setting affords greater time
and control over samples. By effectively evaluating the overall stiffness of cell culture
77
substrates, shear rheometry demonstrated the greatest ability to study altered stiffness in
our liver ECM gels.
5. Conclusions and Potential Applications
In previous studies, hepatocytes cultured on different substrate stiffnesses have
been reported to have generalized effects. Low stiffness is associated with a growth
arrested and differentiated adult phenotype. High stiffness is associated with increased
viability and cell proliferation but also dedifferentiation and possible initiation of EMT
(53). However, the exact stiffness ranges defined in these studies and substrate
compositions vary greatly. Stiffness levels are also often at non-physiologically
achievable levels.
The assays we conducted utilized liver specific ECM isolated by decellularization
to form gels so that substrate components were similar to what composes normal liver
tissue. We also designed our study to focus on a narrow physiological relevant range of
stiffness. The adjustable system was created to culture primary human hepatocytes to
determine optimal conditions for cell function and distinguish specific mechanisms for
stiffness-induced changes in cell mechanics and morphology. We hypothesized that the
substrate stiffness closest to the actual stiffness of native liver tissue promotes the highest
long term function.
This knowledge can enable advancement in the creation of substrates for liver cell
therapies to treat disease in patients. This technology was developed to easily manipulate
and test effects of mechanical properties on cells. Future studies can use these abilities to
simulate the effects of disease states in a variety of organ systems and determine disease
78
mechanisms in order to form possible treatment strategies. Increased dimensions of
complexity in architecture and cell types could also be added to the system. These ECM-
containing hydrogels could eventually be used for tissue engineering applications.
Because of its semi-fluidic properties, cells encapsulated in gels can be injected into
numerous sites in vivo (134-138). Gels can also be “printed” into various patterns for
tissue construction. Bioprinting is a new technology that rapidly deposits substrates into a
predetermined form. The process controls substrate shape, composition, and physical
properties like stiffness. After gels are printed into constructs, materials can be fully
cross-linked depending upon the desired tissue type. Bioprinted tissues currently in
development include everything from skin patches and vessels to complex organs like the
liver (80, 139-142). With the use of ECM hydrogels and new emerging technologies, the
possibilities for new therapeutic products and treatments are endless.
Works Cited
1. Wicha MS, Lowrie G, Kohn E, Bagavandoss P, Mahn T. Extracellular matrix promotes mammary epithelial growth and differentiation in vitro. Proc Natl Acad Sci U S A. 1982;79(10):3213-7. PubMed PMID: 6954472; PMCID: 346385. 2. Remlinger NT, Czajka CA, Juhas ME, Vorp DA, Stolz DB, Badylak SF, Gilbert S, Gilbert TW. Hydrated xenogeneic decellularized tracheal matrix as a scaffold for tracheal reconstruction. Biomaterials. 2010;31(13):3520-6. doi: 10.1016/j.biomaterials.2010.01.067. PubMed PMID: 20144481. 3. Courtman DW, Pereira CA, Kashef V, McComb D, Lee JM, Wilson GJ. Development of a pericardial acellular matrix biomaterial: biochemical and mechanical effects of cell extraction. Journal of biomedical materials research. 1994;28(6):655-66. doi: 10.1002/jbm.820280602. PubMed PMID: 8071376. 4. Xu CC, Chan RW, Tirunagari N. A biodegradable, acellular xenogeneic scaffold for regeneration of the vocal fold lamina propria. Tissue Eng. 2007;13(3):551-66. doi: 10.1089/ten.2006.0169. PubMed PMID: 17518602. 5. Shafiq MA, Gemeinhart RA, Yue BY, Djalilian AR. Decellularized human cornea for reconstructing the corneal epithelium and anterior stroma. Tissue engineering Part C, Methods. 2012;18(5):340-8. doi: 10.1089/ten.TEC.2011.0072. PubMed PMID: 22082039; PMCID: 3338110.
79
6. Lee W, Miyagawa Y, Long C, Cooper DK, Hara H. A comparison of three methods of decellularization of pig corneas to reduce immunogenicity. Int J Ophthalmol. 2014;7(4):587-93. doi: 10.3980/j.issn.2222-3959.2014.04.01. PubMed PMID: 25161926; PMCID: 4137190. 7. Rosario DJ, Reilly GC, Ali Salah E, Glover M, Bullock AJ, Macneil S. Decellularization and sterilization of porcine urinary bladder matrix for tissue engineering in the lower urinary tract. Regen Med. 2008;3(2):145-56. doi: 10.2217/17460751.3.2.145. PubMed PMID: 18307398. 8. Flynn LE, Prestwich GD, Semple JL, Woodhouse KA. Proliferation and differentiation of adipose-derived stem cells on naturally derived scaffolds. Biomaterials. 2008;29(12):1862-71. doi: 10.1016/j.biomaterials.2007.12.028. PubMed PMID: 18242690. 9. Crapo PM, Gilbert TW, Badylak SF. An overview of tissue and whole organ decellularization processes. Biomaterials. 2011;32(12):3233-43. doi: 10.1016/j.biomaterials.2011.01.057. PubMed PMID: 21296410; PMCID: 3084613. 10. Gulati AK. Evaluation of acellular and cellular nerve grafts in repair of rat peripheral nerve. J Neurosurg. 1988;68(1):117-23. doi: 10.3171/jns.1988.68.1.0117. PubMed PMID: 3335896. 11. Jackson DW, Grood ES, Wilcox P, Butler DL, Simon TM, Holden JP. The effects of processing techniques on the mechanical properties of bone-anterior cruciate ligament-bone allografts. An experimental study in goats. Am J Sports Med. 1988;16(2):101-5. PubMed PMID: 3377093. 12. Nonaka PN, Campillo N, Uriarte JJ, Garreta E, Melo E, de Oliveira LV, Navajas D, Farre R. Effects of freezing/thawing on the mechanical properties of decellularized lungs. J Biomed Mater Res A. 2014;102(2):413-9. doi: 10.1002/jbm.a.34708. PubMed PMID: 23533110. 13. Lu H, Hoshiba T, Kawazoe N, Chen G. Comparison of decellularization techniques for preparation of extracellular matrix scaffolds derived from three-dimensional cell culture. J Biomed Mater Res A. 2012;100(9):2507-16. doi: 10.1002/jbm.a.34150. PubMed PMID: 22623317. 14. Burk J, Erbe I, Berner D, Kacza J, Kasper C, Pfeiffer B, Winter K, Brehm W. Freeze-thaw cycles enhance decellularization of large tendons. Tissue engineering Part C, Methods. 2014;20(4):276-84. doi: 10.1089/ten.TEC.2012.0760. PubMed PMID: 23879725; PMCID: 3968887. 15. Kisand K, Kerna I, Kumm J, Jonsson H, Tamm A. Impact of cryopreservation on serum concentration of matrix metalloproteinases (MMP)-7, TIMP-1, vascular growth factors (VEGF) and VEGF-R2 in Biobank samples. Clin Chem Lab Med. 2011;49(2):229-35. doi: 10.1515/CCLM.2011.049. PubMed PMID: 21118050. 16. Sullivan DC, Mirmalek-Sani SH, Deegan DB, Baptista PM, Aboushwareb T, Atala A, Yoo JJ. Decellularization methods of porcine kidneys for whole organ engineering using a high-throughput system. Biomaterials. 2012;33(31):7756-64. doi: 10.1016/j.biomaterials.2012.07.023. PubMed PMID: 22841923. 17. Prasertsung I, Kanokpanont S, Bunaprasert T, Thanakit V, Damrongsakkul S. Development of acellular dermis from porcine skin using periodic pressurized technique. J Biomed Mater Res B Appl Biomater. 2008;85(1):210-9. doi: 10.1002/jbm.b.30938. PubMed PMID: 17853423. 18. Chen RN, Ho HO, Tsai YT, Sheu MT. Process development of an acellular dermal matrix (ADM) for biomedical applications. Biomaterials. 2004;25(13):2679-86. PubMed PMID: 14751754. 19. Meyer SR, Chiu B, Churchill TA, Zhu L, Lakey JR, Ross DB. Comparison of aortic valve allograft decellularization techniques in the rat. J Biomed Mater Res A. 2006;79(2):254-62. doi: 10.1002/jbm.a.30777. PubMed PMID: 16817222.
80
20. Schenke-Layland K, Vasilevski O, Opitz F, Konig K, Riemann I, Halbhuber KJ, Wahlers T, Stock UA. Impact of decellularization of xenogeneic tissue on extracellular matrix integrity for tissue engineering of heart valves. J Struct Biol. 2003;143(3):201-8. PubMed PMID: 14572475. 21. Yang M, Chen CZ, Wang XN, Zhu YB, Gu YJ. Favorable effects of the detergent and enzyme extraction method for preparing decellularized bovine pericardium scaffold for tissue engineered heart valves. J Biomed Mater Res B Appl Biomater. 2009;91(1):354-61. doi: 10.1002/jbm.b.31409. PubMed PMID: 19507136. 22. Yang B, Zhang Y, Zhou L, Sun Z, Zheng J, Chen Y, Dai Y. Development of a porcine bladder acellular matrix with well-preserved extracellular bioactive factors for tissue engineering. Tissue engineering Part C, Methods. 2010;16(5):1201-11. doi: 10.1089/ten.TEC.2009.0311. PubMed PMID: 20170425. 23. Fitzpatrick JC, Clark PM, Capaldi FM. Effect of decellularization protocol on the mechanical behavior of porcine descending aorta. Int J Biomater. 2010;2010. doi: 10.1155/2010/620503. PubMed PMID: 20689621; PMCID: 2910464. 24. Elder BD, Eleswarapu SV, Athanasiou KA. Extraction techniques for the decellularization of tissue engineered articular cartilage constructs. Biomaterials. 2009;30(22):3749-56. doi: 10.1016/j.biomaterials.2009.03.050. PubMed PMID: 19395023; PMCID: 2743309. 25. Nagata S, Hanayama R, Kawane K. Autoimmunity and the clearance of dead cells. Cell. 2010;140(5):619-30. doi: 10.1016/j.cell.2010.02.014. PubMed PMID: 20211132. 26. Kasimir MT, Rieder E, Seebacher G, Silberhumer G, Wolner E, Weigel G, Simon P. Comparison of different decellularization procedures of porcine heart valves. The International journal of artificial organs. 2003;26(5):421-7. PubMed PMID: 12828309. 27. Zou Y, Zhang Y. Mechanical evaluation of decellularized porcine thoracic aorta. J Surg Res. 2012;175(2):359-68. doi: 10.1016/j.jss.2011.03.070. PubMed PMID: 21571306; PMCID: 3100660. 28. Seddon AM, Curnow P, Booth PJ. Membrane proteins, lipids and detergents: not just a soap opera. Biochim Biophys Acta. 2004;1666(1-2):105-17. doi: 10.1016/j.bbamem.2004.04.011. PubMed PMID: 15519311. 29. Allaire E, Guettier C, Bruneval P, Plissonnier D, Michel JB. Cell-free arterial grafts: morphologic characteristics of aortic isografts, allografts, and xenografts in rats. J Vasc Surg. 1994;19(3):446-56. PubMed PMID: 8126857. 30. DeQuach JA, Mezzano V, Miglani A, Lange S, Keller GM, Sheikh F, Christman KL. Simple and high yielding method for preparing tissue specific extracellular matrix coatings for cell culture. PLoS One. 2010;5(9):e13039. doi: 10.1371/journal.pone.0013039. PubMed PMID: 20885963; PMCID: 2946408. 31. Bolland F, Korossis S, Wilshaw SP, Ingham E, Fisher J, Kearney JN, Southgate J. Development and characterisation of a full-thickness acellular porcine bladder matrix for tissue engineering. Biomaterials. 2007;28(6):1061-70. doi: 10.1016/j.biomaterials.2006.10.005. PubMed PMID: 17092557. 32. Faulk DM, Carruthers CA, Warner HJ, Kramer CR, Reing JE, Zhang L, D'Amore A, Badylak SF. The effect of detergents on the basement membrane complex of a biologic scaffold material. Acta biomaterialia. 2014;10(1):183-93. doi: 10.1016/j.actbio.2013.09.006. PubMed PMID: 24055455; PMCID: 3857635. 33. Gratzer PF, Harrison RD, Woods T. Matrix alteration and not residual sodium dodecyl sulfate cytotoxicity affects the cellular repopulation of a decellularized matrix. Tissue Eng. 2006;12(10):2975-83. doi: 10.1089/ten.2006.12.2975. PubMed PMID: 17518665.
81
34. Vavken P, Joshi S, Murray MM. TRITON-X is most effective among three decellularization agents for ACL tissue engineering. J Orthop Res. 2009;27(12):1612-8. doi: 10.1002/jor.20932. PubMed PMID: 19504590; PMCID: 2824567. 35. Ozeki M, Narita Y, Kagami H, Ohmiya N, Itoh A, Hirooka Y, Niwa Y, Ueda M, Goto H. Evaluation of decellularized esophagus as a scaffold for cultured esophageal epithelial cells. Journal of Biomedical Materials Research Part A. 2006;79a(4):771-8. doi: DOI 10.1002/jbm.a.30885. PubMed PMID: WOS:000242360400002. 36. Ren H, Shi X, Tao L, Xiao J, Han B, Zhang Y, Yuan X, Ding Y. Evaluation of two decellularization methods in the development of a whole-organ decellularized rat liver scaffold. Liver Int. 2013;33(3):448-58. doi: 10.1111/liv.12088. PubMed PMID: 23301992. 37. Guyette JP, Gilpin SE, Charest JM, Tapias LF, Ren X, Ott HC. Perfusion decellularization of whole organs. Nat Protoc. 2014;9(6):1451-68. doi: 10.1038/nprot.2014.097. PubMed PMID: 24874812. 38. Mendoza-Novelo B, Avila EE, Cauich-Rodriguez JV, Jorge-Herrero E, Rojo FJ, Guinea GV, Mata-Mata JL. Decellularization of pericardial tissue and its impact on tensile viscoelasticity and glycosaminoglycan content. Acta biomaterialia. 2011;7(3):1241-8. doi: 10.1016/j.actbio.2010.11.017. PubMed PMID: 21094703. 39. Shupe T, Williams M, Brown A, Willenberg B, Petersen BE. Method for the decellularization of intact rat liver. Organogenesis. 2010;6(2):134-6. PubMed PMID: 20885860; PMCID: 2901817. 40. Reing JE, Brown BN, Daly KA, Freund JM, Gilbert TW, Hsiong SX, Huber A, Kullas KE, Tottey S, Wolf MT, Badylak SF. The effects of processing methods upon mechanical and biologic properties of porcine dermal extracellular matrix scaffolds. Biomaterials. 2010;31(33):8626-33. doi: 10.1016/j.biomaterials.2010.07.083. PubMed PMID: 20728934; PMCID: 2956268. 41. Ott HC, Matthiesen TS, Goh SK, Black LD, Kren SM, Netoff TI, Taylor DA. Perfusion-decellularized matrix: using nature's platform to engineer a bioartificial heart. Nat Med. 2008;14(2):213-21. doi: 10.1038/nm1684. PubMed PMID: 18193059. 42. Ravi S, Chaikof EL. Biomaterials for vascular tissue engineering. Regen Med. 2010;5(1):107-20. doi: 10.2217/rme.09.77. PubMed PMID: 20017698; PMCID: 2822541. 43. Gui L, Muto A, Chan SA, Breuer CK, Niklason LE. Development of decellularized human umbilical arteries as small-diameter vascular grafts. Tissue Eng Part A. 2009;15(9):2665-76. doi: 10.1089/ten.TEA.2008.0526. PubMed PMID: 19207043; PMCID: 2735599. 44. Baptista PM, Siddiqui MM, Lozier G, Rodriguez SR, Atala A, Soker S. The use of whole organ decellularization for the generation of a vascularized liver organoid. Hepatology. 2011:n/a-n/a. doi: 10.1002/hep.24067. 45. Yazdani SK, Watts B, Machingal M, Jarajapu YP, Van Dyke ME, Christ GJ. Smooth muscle cell seeding of decellularized scaffolds: the importance of bioreactor preconditioning to development of a more native architecture for tissue-engineered blood vessels. Tissue Eng Part A. 2009;15(4):827-40. doi: 10.1089/ten.tea.2008.0092. PubMed PMID: 19290806. 46. Ko IK, Peng L, Peloso A, Smith CJ, Dhal A, Deegan DB, Zimmerman C, Clouse C, Zhao W, Shupe TD, Soker S, Yoo JJ, Atala A. Bioengineered transplantable porcine livers with re-endothelialized vasculature. Biomaterials. 2015;40:72-9. doi: 10.1016/j.biomaterials.2014.11.027. PubMed PMID: 25433603. 47. Uygun BE, Soto-Gutierrez A, Yagi H, Izamis ML, Guzzardi MA, Shulman C, Milwid J, Kobayashi N, Tilles A, Berthiaume F, Hertl M, Nahmias Y, Yarmush ML, Uygun K. Organ reengineering through development of a transplantable recellularized liver graft using decellularized liver matrix. Nat Med. 2010;16(7):814-20. doi: 10.1038/nm.2170. PubMed PMID: 20543851; PMCID: 2930603.
82
48. Petersen TH, Calle EA, Zhao L, Lee EJ, Gui L, Raredon MB, Gavrilov K, Yi T, Zhuang ZW, Breuer C, Herzog E, Niklason LE. Tissue-engineered lungs for in vivo implantation. Science. 2010;329(5991):538-41. doi: 10.1126/science.1189345. PubMed PMID: 20576850; PMCID: 3640463. 49. Bonvillain RW, Scarritt ME, Pashos NC, Mayeux JP, Meshberger CL, Betancourt AM, Sullivan DE, Bunnell BA. Nonhuman primate lung decellularization and recellularization using a specialized large-organ bioreactor. J Vis Exp. 2013(82):e50825. doi: 10.3791/50825. PubMed PMID: 24378384. 50. Xu X, Prestwich GD. Inhibition of tumor growth and angiogenesis by a lysophosphatidic acid antagonist in an engineered three-dimensional lung cancer xenograft model. Cancer. 2010;116(7):1739-50. doi: 10.1002/cncr.24907. PubMed PMID: 20143443; PMCID: 2847044. 51. Griffini P, Smorenburg SM, Verbeek FJ, van Noorden CJ. Three-dimensional reconstruction of colon carcinoma metastases in liver. J Microsc. 1997;187(Pt 1):12-21. PubMed PMID: 9263437. 52. Wells RG. Cellular sources of extracellular matrix in hepatic fibrosis. Clin Liver Dis. 2008;12(4):759-68, viii. doi: 10.1016/j.cld.2008.07.008. PubMed PMID: 18984465; PMCID: 2617723. 53. Wells RG. The role of matrix stiffness in regulating cell behavior. Hepatology. 2008;47(4):1394-400. doi: 10.1002/hep.22193. PubMed PMID: 18307210. 54. Bohm F, Kohler UA, Speicher T, Werner S. Regulation of liver regeneration by growth factors and cytokines. EMBO Mol Med. 2010;2(8):294-305. doi: 10.1002/emmm.201000085. PubMed PMID: 20652897; PMCID: 3377328. 55. Huh CG, Factor VM, Sanchez A, Uchida K, Conner EA, Thorgeirsson SS. Hepatocyte growth factor/c-met signaling pathway is required for efficient liver regeneration and repair. Proc Natl Acad Sci U S A. 2004;101(13):4477-82. doi: 10.1073/pnas.0306068101. PubMed PMID: 15070743; PMCID: 384772. 56. Fausto N, Laird AD, Webber EM. Liver regeneration. 2. Role of growth factors and cytokines in hepatic regeneration. Faseb J. 1995;9(15):1527-36. PubMed PMID: 8529831. 57. Pi L, Ding X, Jorgensen M, Pan JJ, Oh SH, Pintilie D, Brown A, Song WY, Petersen BE. Connective tissue growth factor with a novel fibronectin binding site promotes cell adhesion and migration during rat oval cell activation. Hepatology. 2008;47(3):996-1004. doi: 10.1002/hep.22079. PubMed PMID: 18167060; PMCID: 3130595. 58. Alberts B. Molecular biology of the cell. 4th ed. New York: Garland Science; 2002. xxxiv, 1548 p. p. 59. Ramalingam M. Tissue engineering and regenerative medicine : a nano approach. Boca Raton: CRC Press; 2013. xxii, 570 p. p. 60. Lessey BA, Albelda S, Buck CA, Castelbaum AJ, Yeh I, Kohler M, Berchuck A. Distribution of integrin cell adhesion molecules in endometrial cancer. Am J Pathol. 1995;146(3):717-26. PubMed PMID: 7887452; PMCID: 1869177. 61. Grose R, Hutter C, Bloch W, Thorey I, Watt FM, Fassler R, Brakebusch C, Werner S. A crucial role of beta 1 integrins for keratinocyte migration in vitro and during cutaneous wound repair. Development. 2002;129(9):2303-15. PubMed PMID: 11959837. 62. Williams D. Collagen: ubiquitous in nature, multifunctional in devices. Med Device Technol. 1998;9(6):10-3. PubMed PMID: 10182119. 63. Simon Davis DA, Parish CR. Heparan sulfate: a ubiquitous glycosaminoglycan with multiple roles in immunity. Front Immunol. 2013;4:470. doi: 10.3389/fimmu.2013.00470. PubMed PMID: 24391644; PMCID: 3866581.
83
64. Dreyfuss JL, Regatieri CV, Jarrouge TR, Cavalheiro RP, Sampaio LO, Nader HB. Heparan sulfate proteoglycans: structure, protein interactions and cell signaling. An Acad Bras Cienc. 2009;81(3):409-29. PubMed PMID: 19722012. 65. Watt FM, Huck WT. Role of the extracellular matrix in regulating stem cell fate. Nat Rev Mol Cell Biol. 2013;14(8):467-73. doi: 10.1038/nrm3620. PubMed PMID: 23839578. 66. Santiago JA, Pogemiller R, Ogle BM. Heterogeneous differentiation of human mesenchymal stem cells in response to extended culture in extracellular matrices. Tissue Eng Part A. 2009;15(12):3911-22. doi: 10.1089/ten.tea.2008.0603. PubMed PMID: 19911955; PMCID: 2792070. 67. Lin H, Yang G, Tan J, Tuan RS. Influence of decellularized matrix derived from human mesenchymal stem cells on their proliferation, migration and multi-lineage differentiation potential. Biomaterials. 2012;33(18):4480-9. doi: 10.1016/j.biomaterials.2012.03.012. PubMed PMID: 22459197. 68. Ross EA, Williams MJ, Hamazaki T, Terada N, Clapp WL, Adin C, Ellison GW, Jorgensen M, Batich CD. Embryonic stem cells proliferate and differentiate when seeded into kidney scaffolds. Journal of the American Society of Nephrology : JASN. 2009;20(11):2338-47. doi: 10.1681/ASN.2008111196. PubMed PMID: 19729441; PMCID: 2799178. 69. Discher DE, Mooney DJ, Zandstra PW. Growth factors, matrices, and forces combine and control stem cells. Science. 2009;324(5935):1673-7. doi: 10.1126/science.1171643. PubMed PMID: 19556500; PMCID: 2847855. 70. Sundelacruz S, Kaplan DL. Stem cell- and scaffold-based tissue engineering approaches to osteochondral regenerative medicine. Semin Cell Dev Biol. 2009;20(6):646-55. doi: 10.1016/j.semcdb.2009.03.017. PubMed PMID: 19508851; PMCID: 2737137. 71. Gattazzo F, Urciuolo A, Bonaldo P. Extracellular matrix: a dynamic microenvironment for stem cell niche. Biochim Biophys Acta. 2014;1840(8):2506-19. doi: 10.1016/j.bbagen.2014.01.010. PubMed PMID: 24418517; PMCID: 4081568. 72. Tanimizu N, Miyajima A, Mostov KE. Liver progenitor cells develop cholangiocyte-type epithelial polarity in three-dimensional culture. Mol Biol Cell. 2007;18(4):1472-9. doi: 10.1091/mbc.E06-09-0848. PubMed PMID: 17314404; PMCID: 1838984. 73. Tanimizu N, Saito H, Mostov K, Miyajima A. Long-term culture of hepatic progenitors derived from mouse Dlk+ hepatoblasts. J Cell Sci. 2004;117(Pt 26):6425-34. doi: 10.1242/jcs.01572. PubMed PMID: 15572411. 74. Malinen MM, Kanninen LK, Corlu A, Isoniemi HM, Lou YR, Yliperttula ML, Urtti AO. Differentiation of liver progenitor cell line to functional organotypic cultures in 3D nanofibrillar cellulose and hyaluronan-gelatin hydrogels. Biomaterials. 2014;35(19):5110-21. doi: 10.1016/j.biomaterials.2014.03.020. PubMed PMID: 24698520. 75. Murphy SV, Skardal A, Atala A. Evaluation of hydrogels for bio-printing applications. J Biomed Mater Res A. 2013;101(1):272-84. doi: 10.1002/jbm.a.34326. PubMed PMID: 22941807. 76. Burdick JA, Prestwich GD. Hyaluronic acid hydrogels for biomedical applications. Adv Mater. 2011;23(12):H41-56. doi: 10.1002/adma.201003963. PubMed PMID: 21394792; PMCID: 3730855. 77. Zhou P, Lessa N, Estrada DC, Severson EB, Lingala S, Zern MA, Nolta JA, Wu JA. Decellularized Liver Matrix as a Carrier for the Transplantation of Human Fetal and Primary Hepatocytes in Mice. Liver Transplant. 2011;17(4):418-27. doi: Doi 10.1002/Lt.22270. PubMed PMID: WOS:000289004000009. 78. Skardal A, Smith L, Bharadwaj S, Atala A, Soker S, Zhang Y. Tissue specific synthetic ECM hydrogels for 3-D in vitro maintenance of hepatocyte function. Biomaterials. 2012;33(18):4565-75. doi: 10.1016/j.biomaterials.2012.03.034. PubMed PMID: 22475531; PMCID: 3719050.
84
79. Nakamura S, Ijima H. Solubilized matrix derived from decellularized liver as a growth factor-immobilizable scaffold for hepatocyte culture. J Biosci Bioeng. 2013;116(6):746-53. doi: DOI 10.1016/j.jbiosc.2013.05.031. PubMed PMID: WOS:000329087800016. 80. Pati F, Jang J, Ha DH, Won Kim S, Rhie JW, Shim JH, Kim DH, Cho DW. Printing three-dimensional tissue analogues with decellularized extracellular matrix bioink. Nat Commun. 2014;5:3935. doi: 10.1038/ncomms4935. PubMed PMID: 24887553; PMCID: 4059935. 81. Lin P, Chan WC, Badylak SF, Bhatia SN. Assessing porcine liver-derived biomatrix for hepatic tissue engineering. Tissue Eng. 2004;10(7-8):1046-53. doi: 10.1089/ten.2004.10.1046. PubMed PMID: 15363162. 82. Fishman JM, Lowdell MW, Urbani L, Ansari T, Burns AJ, Turmaine M, North J, Sibbons P, Seifalian AM, Wood KJ, Birchall MA, De Coppi P. Immunomodulatory effect of a decellularized skeletal muscle scaffold in a discordant xenotransplantation model. Proc Natl Acad Sci U S A. 2013;110(35):14360-5. doi: 10.1073/pnas.1213228110. PubMed PMID: 23940349; PMCID: 3761636. 83. Zhang FT, Wan HJ, Li MH, Ye J, Yin MJ, Huang CQ, Yu J. Transplantation of microencapsulated umbilical-cord-blood-derived hepatic-like cells for treatment of hepatic failure. World journal of gastroenterology : WJG. 2011;17(7):938-45. doi: 10.3748/wjg.v17.i7.938. PubMed PMID: 21412504; PMCID: 3051145. 84. Kim M, Lee JY, Jones CN, Revzin A, Tae G. Heparin-based hydrogel as a matrix for encapsulation and cultivation of primary hepatocytes. Biomaterials. 2010;31(13):3596-603. doi: 10.1016/j.biomaterials.2010.01.068. PubMed PMID: 20153045; PMCID: 2837121. 85. Tse JR, Engler AJ. Stiffness gradients mimicking in vivo tissue variation regulate mesenchymal stem cell fate. PLoS One. 2011;6(1):e15978. doi: 10.1371/journal.pone.0015978. PubMed PMID: 21246050; PMCID: 3016411. 86. Mih JD, Sharif AS, Liu F, Marinkovic A, Symer MM, Tschumperlin DJ. A multiwell platform for studying stiffness-dependent cell biology. PLoS One. 2011;6(5):e19929. doi: 10.1371/journal.pone.0019929. PubMed PMID: 21637769; PMCID: 3103526. 87. You J, Park SA, Shin DS, Patel D, Raghunathan VK, Kim M, Murphy CJ, Tae G, Revzin A. Characterizing the effects of heparin gel stiffness on function of primary hepatocytes. Tissue Eng Part A. 2013;19(23-24):2655-63. doi: 10.1089/ten.TEA.2012.0681. PubMed PMID: 23815179; PMCID: 3856597. 88. Ratner BD, Bryant SJ. Biomaterials: where we have been and where we are going. Annu Rev Biomed Eng. 2004;6:41-75. doi: 10.1146/annurev.bioeng.6.040803.140027. PubMed PMID: 15255762. 89. Green NS, Reisler E, Houk KN. Quantitative evaluation of the lengths of homobifunctional protein cross-linking reagents used as molecular rulers. Protein Sci. 2001;10(7):1293-304. doi: 10.1110/ps.51201. PubMed PMID: 11420431; PMCID: 2374107. 90. Radosevich JA. UV radiation : properties, effects, and applications. New York: Nova Publishers; 2014. xiii, 355 pages p. 91. Mattson G, Conklin E, Desai S, Nielander G, Savage MD, Morgensen S. A practical approach to crosslinking. Mol Biol Rep. 1993;17(3):167-83. PubMed PMID: 8326953. 92. Narain R. Chemistry of bioconjugates : synthesis, characterization, and biomedical applications. Hoboken, New Jersey: Wiley; 2014. x, 464 pages p. 93. Skardal A, Zhang J, McCoard L, Xu X, Oottamasathien S, Prestwich GD. Photocrosslinkable hyaluronan-gelatin hydrogels for two-step bioprinting. Tissue Eng Part A. 2010;16(8):2675-85. doi: 10.1089/ten.TEA.2009.0798. PubMed PMID: 20387987; PMCID: 3135254.
85
94. Guvendiren M, Burdick JA. Stiffening hydrogels to probe short- and long-term cellular responses to dynamic mechanics. Nat Commun. 2012;3:792. doi: 10.1038/ncomms1792. PubMed PMID: 22531177. 95. Hermanson GT. Bioconjugate techniques. Third edition. ed. London ; Waltham, MA: Elsevier/AP; 2013. xvii, 1146 pages p. 96. Rubinstein M, Colby RH. Polymer physics. Oxford ; New York: Oxford University Press; 2003. xi, 440 p. p. 97. Ali ME, Lamprecht A. Polyethylene glycol as an alternative polymer solvent for nanoparticle preparation. Int J Pharm. 2013;456(1):135-42. doi: 10.1016/j.ijpharm.2013.07.077. PubMed PMID: 23958752. 98. Webster R, Didier E, Harris P, Siegel N, Stadler J, Tilbury L, Smith D. PEGylated proteins: evaluation of their safety in the absence of definitive metabolism studies. Drug Metab Dispos. 2007;35(1):9-16. doi: 10.1124/dmd.106.012419. PubMed PMID: 17020954. 99. Zalipsky S, Mullah N, Harding JA, Gittelman J, Guo L, DeFrees SA. Poly(ethylene glycol)-grafted liposomes with oligopeptide or oligosaccharide ligands appended to the termini of the polymer chains. Bioconjug Chem. 1997;8(2):111-8. doi: 10.1021/bc9600832. PubMed PMID: 9095350. 100. Oliveira CP, Ribeiro ME, Ricardo NM, Souza TV, Moura CL, Chaibundit C, Yeates SG, Nixon K, Attwood D. The effect of water-soluble polymers, PEG and PVP, on the solubilisation of griseofulvin in aqueous micellar solutions of Pluronic F127. Int J Pharm. 2011;421(2):252-7. doi: 10.1016/j.ijpharm.2011.10.010. PubMed PMID: 22001534. 101. Bacakova L, Filova E, Parizek M, Ruml T, Svorcik V. Modulation of cell adhesion, proliferation and differentiation on materials designed for body implants. Biotechnol Adv. 2011;29(6):739-67. doi: 10.1016/j.biotechadv.2011.06.004. PubMed PMID: 21821113. 102. Benhabbour SR, Sheardown H, Adronov A. Cell adhesion and proliferation on hydrophilic dendritically modified surfaces. Biomaterials. 2008;29(31):4177-86. doi: 10.1016/j.biomaterials.2008.07.016. PubMed PMID: 18678405. 103. Lee KY, Mooney DJ. Hydrogels for tissue engineering. Chem Rev. 2001;101(7):1869-79. PubMed PMID: 11710233. 104. Tseng H, Cuchiara ML, Durst CA, Cuchiara MP, Lin CJ, West JL, Grande-Allen KJ. Fabrication and mechanical evaluation of anatomically-inspired quasilaminate hydrogel structures with layer-specific formulations. Ann Biomed Eng. 2013;41(2):398-407. doi: 10.1007/s10439-012-0666-5. PubMed PMID: 23053300; PMCID: 3545057. 105. Zhu J. Bioactive modification of poly(ethylene glycol) hydrogels for tissue engineering. Biomaterials. 2010;31(17):4639-56. doi: 10.1016/j.biomaterials.2010.02.044. PubMed PMID: 20303169; PMCID: 2907908. 106. Wells PN, Liang HD. Medical ultrasound: imaging of soft tissue strain and elasticity. Journal of the Royal Society, Interface / the Royal Society. 2011;8(64):1521-49. doi: 10.1098/rsif.2011.0054. PubMed PMID: 21680780; PMCID: 3177611. 107. Ophir J, Cespedes I, Ponnekanti H, Yazdi Y, Li X. Elastography: a quantitative method for imaging the elasticity of biological tissues. Ultrason Imaging. 1991;13(2):111-34. PubMed PMID: 1858217. 108. Ganne-Carrie N, Ziol M, de Ledinghen V, Douvin C, Marcellin P, Castera L, Dhumeaux D, Trinchet JC, Beaugrand M. Accuracy of liver stiffness measurement for the diagnosis of cirrhosis in patients with chronic liver diseases. Hepatology. 2006;44(6):1511-7. doi: 10.1002/hep.21420. PubMed PMID: 17133503. 109. Sandrin L, Fourquet B, Hasquenoph JM, Yon S, Fournier C, Mal F, Christidis C, Ziol M, Poulet B, Kazemi F, Beaugrand M, Palau R. Transient elastography: a new noninvasive method
86
for assessment of hepatic fibrosis. Ultrasound Med Biol. 2003;29(12):1705-13. PubMed PMID: 14698338. 110. Ferraioli G, Tinelli C, Lissandrin R, Zicchetti M, Dal Bello B, Filice G, Filice C. Point shear wave elastography method for assessing liver stiffness. World journal of gastroenterology : WJG. 2014;20(16):4787-96. doi: 10.3748/wjg.v20.i16.4787. PubMed PMID: 24782633; PMCID: 4000517. 111. Parker KJ, Doyley MM, Rubens DJ. Imaging the elastic properties of tissue: the 20 year perspective. Phys Med Biol. 2011;56(1):R1-R29. doi: 10.1088/0031-9155/56/1/R01. PubMed PMID: 21119234. 112. Ferraioli G, Parekh P, Levitov AB, Filice C. Shear wave elastography for evaluation of liver fibrosis. J Ultrasound Med. 2014;33(2):197-203. doi: 10.7863/ultra.33.2.197. PubMed PMID: 24449721. 113. Nightingale K. Acoustic Radiation Force Impulse (ARFI) Imaging: a Review. Curr Med Imaging Rev. 2011;7(4):328-39. doi: 10.2174/157340511798038657. PubMed PMID: 22545033; PMCID: 3337770. 114. Mariappan YK, Glaser KJ, Ehman RL. Magnetic resonance elastography: a review. Clin Anat. 2010;23(5):497-511. doi: 10.1002/ca.21006. PubMed PMID: 20544947; PMCID: 3066083. 115. Muthupillai R, Lomas DJ, Rossman PJ, Greenleaf JF, Manduca A, Ehman RL. Magnetic resonance elastography by direct visualization of propagating acoustic strain waves. Science. 1995;269(5232):1854-7. PubMed PMID: 7569924. 116. Yin M, Talwalkar JA, Glaser KJ, Manduca A, Grimm RC, Rossman PJ, Fidler JL, Ehman RL. Assessment of hepatic fibrosis with magnetic resonance elastography. Clin Gastroenterol Hepatol. 2007;5(10):1207-13 e2. doi: 10.1016/j.cgh.2007.06.012. PubMed PMID: 17916548; PMCID: 2276978. 117. Sarvazyan A, Hall TJ, Urban MW, Fatemi M, Aglyamov SR, Garra BS. An Overview of Elastography - an Emerging Branch of Medical Imaging. Curr Med Imaging Rev. 2011;7(4):255-82. PubMed PMID: 22308105; PMCID: 3269947. 118. Klatt D, Friedrich C, Korth Y, Vogt R, Braun J, Sack I. Viscoelastic properties of liver measured by oscillatory rheometry and multifrequency magnetic resonance elastography. Biorheology. 2010;47(2):133-41. doi: 10.3233/BIR-2010-0565. PubMed PMID: 20683156. 119. Chan RW, Rodriguez ML. A simple-shear rheometer for linear viscoelastic characterization of vocal fold tissues at phonatory frequencies. J Acoust Soc Am. 2008;124(2):1207-19. doi: 10.1121/1.2946715. PubMed PMID: 18681608; PMCID: 2561277. 120. Wang M, Kornfield JA. Measuring shear strength of soft-tissue adhesives. J Biomed Mater Res B Appl Biomater. 2012;100(3):618-23. doi: 10.1002/jbm.b.31981. PubMed PMID: 22323271. 121. Askeland DR, Fulay PP, Wright WJ. The science and engineering of materials. 6th ed. Stamford, CT: Cengage Learning; 2011. xxi, 921 p. p. 122. Megson THG. Structural and stress analysis. Third edition. ed. Amsterdam ; Boston: Butterworth-Heinemann is an imprint of Elsevier; 2014. xvii, 748 pages p. 123. McNaught AD, Wilkinson A, International Union of Pure and Applied Chemistry. Compendium of chemical terminology : IUPAC recommendations. 2nd ed. Oxford England ; Malden, MA, USA: Blackwell Science; 1997. vii, 450 p. p. 124. Meyers MA, Chawla KK. Mechanical behavior of materials. Upper Saddle River, N.J.: Prentice Hall; 1999. xv, 680 p. p. 125. Yang J, Shao Z. Recent advances in biological atomic force microscopy. Micron. 1995;26(1):35-49. PubMed PMID: 7704519.
87
126. Giessibl FJ, Morita S. Non-contact AFM. J Phys Condens Matter. 2012;24(8):080301. doi: 10.1088/0953-8984/24/8/080301. PubMed PMID: 22311696. 127. Hofmann T, Pielmeier F, Giessibl FJ. Chemical and crystallographic characterization of the tip apex in scanning probe microscopy. Phys Rev Lett. 2014;112(6):066101. PubMed PMID: 24580694. 128. Roduit C, Sekatski S, Dietler G, Catsicas S, Lafont F, Kasas S. Stiffness tomography by atomic force microscopy. Biophys J. 2009;97(2):674-7. doi: 10.1016/j.bpj.2009.05.010. PubMed PMID: 19619482; PMCID: 2711326. 129. Lekka M, Gil D, Pogoda K, Dulinska-Litewka J, Jach R, Gostek J, Klymenko O, Prauzner-Bechcicki S, Stachura Z, Wiltowska-Zuber J, Okon K, Laidler P. Cancer cell detection in tissue sections using AFM. Arch Biochem Biophys. 2012;518(2):151-6. doi: 10.1016/j.abb.2011.12.013. PubMed PMID: 22209753. 130. Wiesendanger R. Scanning probe microscopy and spectroscopy : methods and applications. Cambridge England ; New York: Cambridge University Press; 1994. xxii, 637 p. p. 131. Eaton PJ, West P. Atomic force microscopy. Oxford ; New York: Oxford University Press; 2010. viii, 248 p. p. 132. Korkin A, Krstic P, Miskovic Z, Yu H, Zhitomirsky I. Nanoscale Science and Technology for Electronics, Photonics and Renewable Energy Applications: Selected Papers from NGC2009 & CSTC2009 conference (http://asdn.net/ngc2009/). Nanoscale Res Lett. 2010;5(3):453. doi: 10.1007/s11671-010-9564-7. PubMed PMID: 20672095; PMCID: 2894060. 133. Le Rouzic J, Vairac P, Cretin B, Delobelle P. Sensitivity optimization of the scanning microdeformation microscope and application to mechanical characterization of soft materials. Rev Sci Instrum. 2008;79(3):033707. doi: 10.1063/1.2894208. PubMed PMID: 18377015. 134. Heile A, Brinker T. Clinical translation of stem cell therapy in traumatic brain injury: the potential of encapsulated mesenchymal cell biodelivery of glucagon-like peptide-1. Dialogues Clin Neurosci. 2011;13(3):279-86. PubMed PMID: 22034462; PMCID: 3182013. 135. Karle P, Muller P, Renz R, Jesnowski R, Saller R, von Rombs K, Nizze H, Liebe S, Gunzburg WH, Salmons B, Lohr M. Intratumoral injection of encapsulated cells producing an oxazaphosphorine activating cytochrome P450 for targeted chemotherapy. Adv Exp Med Biol. 1998;451:97-106. PubMed PMID: 10026857. 136. Levit RD, Landazuri N, Phelps EA, Brown ME, Garcia AJ, Davis ME, Joseph G, Long R, Safley SA, Suever JD, Lyle AN, Weber CJ, Taylor WR. Cellular encapsulation enhances cardiac repair. J Am Heart Assoc. 2013;2(5):e000367. doi: 10.1161/JAHA.113.000367. PubMed PMID: 24113327; PMCID: 3835246. 137. Goren A, Dahan N, Goren E, Baruch L, Machluf M. Encapsulated human mesenchymal stem cells: a unique hypoimmunogenic platform for long-term cellular therapy. Faseb J. 2010;24(1):22-31. doi: 10.1096/fj.09-131888. PubMed PMID: 19726759. 138. Knippenberg S, Thau N, Dengler R, Brinker T, Petri S. Intracerebroventricular injection of encapsulated human mesenchymal cells producing glucagon-like peptide 1 prolongs survival in a mouse model of ALS. PLoS One. 2012;7(6):e36857. doi: 10.1371/journal.pone.0036857. PubMed PMID: 22745655; PMCID: 3380029. 139. Duan B, Hockaday LA, Kang KH, Butcher JT. 3D bioprinting of heterogeneous aortic valve conduits with alginate/gelatin hydrogels. J Biomed Mater Res A. 2013;101(5):1255-64. doi: 10.1002/jbm.a.34420. PubMed PMID: 23015540; PMCID: 3694360. 140. Khalil S, Sun W. Bioprinting endothelial cells with alginate for 3D tissue constructs. J Biomech Eng. 2009;131(11):111002. doi: 10.1115/1.3128729. PubMed PMID: 20353253. 141. Murphy SV, Atala A. 3D bioprinting of tissues and organs. Nat Biotechnol. 2014;32(8):773-85. doi: 10.1038/nbt.2958. PubMed PMID: 25093879.
88
142. Chang R, Emami K, Wu H, Sun W. Biofabrication of a three-dimensional liver micro-organ as an in vitro drug metabolism model. Biofabrication. 2010;2(4):045004. doi: 10.1088/1758-5082/2/4/045004. PubMed PMID: 21079286.
89
CHAPTER 3
Stiffness of Hyaluronic Acid Gels Containing Liver
Extracellular Matrix Supports Human Hepatocyte Function
and Alters Cell Morphology
Daniel B. Deegan
This chapter includes and expands upon the results of the published manuscript,
“Stiffness of Hyaluronic Acid Gels Containing Liver Extracellular Matrix Supports
Human Hepatocyte Function and Alters Cell Morphology.”
Manuscript Reference:
Deegan D.B., Zimmerman C., Skardal A, Atala A., Shupe T., Stiffness of Hyaluronic Acid Gels
Containing Liver Extracellular Matrix Supports Hepatocyte Function and Alters Cell Morphology. Journal
of the Mechanical Behavior of Biomedical Materials. 2015; pp. 87-103
90
Abstract
Tissue engineering and cell based liver therapies have utilized primary hepatocytes with
limited success due to the failure of hepatocytes to maintain their phenotype in vitro. In
order to overcome this challenge, hyaluronic acid (HA) cell culture substrates were
formulated to closely mimic the composition and stiffness of the normal liver cellular
microenvironment. The stiffness of the substrate was modulated by adjusting HA
hydrogel crosslinking. Additionally, the repertoire of bioactive molecules within the HA
substrate was bolstered by supplementation with normal liver extracellular matrix
(ECM). Primary human hepatocyte viability and phenotype were determined over a
narrow physiologically relevant range of substrate stiffnesses from 600 to 4600 Pa in
both the presence and absence of liver ECM. Cell attachment, viability, and organization
of the actin cytoskeleton improved with increased stiffness up to 4600 Pa. These
differences were not evident in earlier time points or substrates containing only HA.
However, gene expression for the hepatocyte markers hepatocyte nuclear factor 4 alpha
(HNF4α) and albumin significantly decreased on the 4600 Pa stiffness at day 7 indicating
that cells may not have maintained their phenotype long-term at this stiffness. Function,
as measured by albumin secretion, varied with both stiffness and time in culture, peaking
at day 7 at the 1200 Pa stiffness, slightly below the stiffness of normal liver ECM at 3000
Pa. Overall, gel stiffness affected primary human hepatocyte cell adhesion, functional
marker expression, and morphological characteristics dependent on both the presence of
liver ECM in gel substrates and time in culture.
91
1. Introduction
Rising incidence and cost of liver disease as well as a limited supply of
transplantable organs have increased demand for new liver treatments. Experimental
treatments currently being developed include liver cell transplant on biological scaffolds
(1-5). However, several limitations intrinsic to primary hepatocytes have slowed
development of these technologies. Challenges remain in the maintenance of viability and
cell function of primary hepatocytes outside of the normal liver microenvironment (5).
Therefore, developing substrates that support hepatocyte viability and function is critical
to maximize the efficacy of primary hepatocytes in tissue engineering and cell therapies
for liver disease.
Previous studies have identified several advantages of using tissue-specific ECM
instead of generic matrix formulations such as Matrigel for supporting primary cells (6).
Cells on natural liver ECM demonstrated increased ability to proliferate and maintain
function over extended time periods (7). Hepatocytes cultured on intact or solubilized
organ specific matrix also maintained normal cell morphologies as compared to cells
grown on tissue culture plastic or collagen (3, 8, 9). The underlying mechanisms for this
improved phenotypic stability are still unknown but likely relate to inclusion of several
components of the natural liver microenvironment, including cell and growth factor
binding sites.
For this study, decellularization of normal liver was used to isolate acellular ECM
for incorporation into hyaluronic acid (HA) hydrogels. The decellularization process
involved perfusion of the organ with detergents to remove native cellular material, while
92
preserving structural proteins and glycosaminoglycans (GAGs) (10). Removal of native
cells leaves a non-immunogenic ECM scaffold that can be seeded with hepatocytes that
remain viable for several weeks (11).
Many previous studies utilized ECM coated tissue culture plastic for growing
primary hepatocytes. However, this method is only suitable for two-dimensional culture
and is not easily translated into a three-dimensional form for use in cell based therapies
(12, 13). The current study uses a hydrogel that may be formed into three-dimensional
units suitable for transplantation. Preliminary findings indicated that solubilized ECM
combined with hyaluronic acid gel (HA) maintained a normal epithelial morphology and
strong tight junction formation with little evidence of transition to a fibroblast-like
morphology, commonly seen in traditional collagen cultures. Solubilization of the ECM
also prevented hepatocyte apoptosis induced by phagocytosis of cryomilled ECM
powders. Other studies published by our group demonstrated that HA based hydrogels
avoided an immunogenic response measured by human lymphocyte proliferation, while
other gel types such as rat tail collagen 1 induced a low level immune response (11, 14).
Better phenotype maintenance and biocompatibility upon transplantation signaled that
HA gel was the best substrate for incorporating decellularized liver in future assays.
In the development of a liver ECM substrate, the physical characteristic of
stiffness is important to cell physiology and mechanics. The general conclusions of
previous publications indicate that stiff substrates promote hepatocyte spreading,
proliferation, and dedifferentiation; while soft substrates promote maintenance of the
functional hepatocyte phenotype (15, 16). Substrate stiffness also regulates cell
aggregation, growth factor responsiveness, and cell motility with the highest cell
93
migration occurring on intermediate stiffness levels (17, 18). However, several of these
previous studies were done on stiffness ranges well above physiological liver stiffness,
which has been reported anywhere from 1.5 to 8.5 kPa (19-24).
For the current study, the crosslinker concentration was the sole variable for
adjusting gel stiffness. Crosslinking eliminated potential confounding effects found in
more complex systems where concentrations of structural compounds are altered. Gel
stiffnesses were also limited to a narrow, physiologically relevant range of 600-4600 Pa
that bracketed the stiffness of normal liver ECM at 3000 Pa. Assays were conducted to
measure overall hepatocyte function as well as the mechanisms by which substrate
stiffness affected cell phenotype. Specific tests performed in this study were designed to
explore the mechanisms by which substrate stiffness affected substrate/cell adhesion,
primary hepatocyte function, and morphology. Cell junction formation is critical for the
maintenance of epithelial cell phenotype; regulating a broad range of cellular properties
through cell-cell communication, adhesion, and diffusion of water and solutes (25, 26).
Tight junctions act as semi-permeable barriers that establish cell polarity, which is crucial
for the maintenance of bile canaliculi (27, 28). The tight junction protein, claudin-1, is
required for maintenance of the selective barrier properties between adjacent hepatocytes
(29). Occludin is not essential for maintaining tight junction structures, but plays a role in
regulating cytoskeletal organization, cell polarity, and cell migration (30-32).
The effects of substrate stiffness on intracellular signaling were also studied
through measurement of two cytoplasmic kinases: focal adhesion kinase (FAK) and
integrin-linked kinase (ILK). Cells adapt their cytoskeletal structure to specific
microenvironmental conditions through interactions between focal adhesion molecules
94
and intracellular kinases. Appropriate expression of these kinases in hepatocytes is
required for normal growth factor responsiveness, prevention of apoptosis, and normal
metabolic function (33-35). Silencing or overexpression of these factors can lead to
cellular dysfunction and, in the case of neoplasia, tumor progression (36). Increased
expression also mediates cytoskeletal reorganization and cell protrusion formation which
drives cell migration (37, 38). In other experimental systems, FAK and ILK expression
have been shown to correlate with cell junction remodeling, specifically through
interactions with occludin. FAK and ILK also act through downstream activation of Ras
homolog family member (Rho) GTPases; cell division control protein 42 (CDC42), Ras-
related C3 botulinum toxin substrate (Rac1), and Ras homolog family member A (RhoA)
(39, 40). CDC42 and Rac are the two GTPases involved in actin cytoskeletal
reorganization for cell motility, including formation of cells extensions called filopodia
and lamellipodia. RhoA expression increases focal adhesion and stress fiber formation
and is correlated with a more contractile morphology (41-44).
By studying the stiffness responsive factors that influence hepatocyte structure
and function, we have gained insights into the mechanisms through which substrate
stiffness affects the viability and phenotypic stability of primary hepatocytes. These
findings have informed the development of a transplantable gel containing liver specific
ECM at an optimal stiffness for therapeutic use in hepatocyte cell therapies.
95
2. Materials and Methods
2.1 Liver Decellularization
Intact rat livers were decellularized according to our group’s previously published
methods (10). Ten week old male F344 rats were euthanized by administering a lethal
dose of sodium pentobarbital (100 mg/kg). Livers were then cannulated and perfused by
methods previously diagrammed and reported in our cell isolation protocol (45). In brief,
a euthanized rat was pinned on a dissection board, and a mid-line incision was made
through the peritoneum starting from the diaphragm down to the groin. Visceral organs
were shifted to expose the inferior vena cava (IVC) and liver. The IVC was cannulated
with a 20-gauge catheter and tied in place with sutures. The portal vein was then cut, and
the superior vena cava (SVC) was clamped. Solutions were held in flasks at 37°C in a
water bath (Fisher Scientific, Pittsburgh, PA) and then perfused through the catheter at a
flow rate of 5 mL/minute using tubing and a Masterflex L/S peristaltic pump (Cole-
Parmer Instrument Company, Vernon Hill, IL).
100 mL of PBS was first perfused to clear residual blood. The liver was then
perfused with approximately 300 mL each of 1%, 2% and 3% Triton X-100 solutions in
PBS. A final solution of 0.1% SDS in PBS was used to clear remaining DNA and Triton
detergent. The liver was then fully excised, and overnight PBS washing was used to
remove residual SDS from the matrices. All animal procedures were approved by the
Institutional Animal Care and Use Committee (IACUC) at Wake Forest University.
96
2.2 Immunohistochemistry (IHC) on Decellularized Tissue
Following decellularization, we immunostained rat livers to assess cell clearance
and retention of structural components such as collagens and laminin. As reported in our
previous manuscript, decellularization completely removed cellular material from tissue
sections, while collagen and laminin were preserved and represented (Figure 1) (10).
For this study, Alcian blue staining for mucopolysaccharides that comprise GAGs
was performed on frozen decellularized liver sections. Tissue was frozen in OCT blocks
and cut into 5 μm slices that were then fixed in acetone on slides at 4ºC for two minutes.
Sections were then stained according to a previously established protocol (46). In short,
Fig.1) IHC of Decellularized Liver Tissue, adapted from Shupe et al (10)
A) Hematoxylin and eosin staining (H&E) shows full cellular removal. B)
Trichrome staining shows presence of structural collagens. C) Staining for
laminin and D) collagen IV indicate presence of a basement membrane.
97
sections were incubated in 0.5% Alcian blue 8G solution in 0.5% acetic acid for 30
minutes (Sigma-Aldrich, St. Louis, MO). Sections were then washed with PBS and
imaged.
2.3 GAG Function in Decellularized Tissue
To assess the ability of residual GAGs to bind growth factors following
decellularization, 5 mg samples of decellularized liver were incubated at room
temperature for one hour in PBS containing an array of concentrations of recombinant
epidermal growth factor (EGF) and human hepatocyte growth factor (HGF) up to 1000
ng/mL (Peprotech, Rocky Hill, NJ). Following pre-treatment, tissue was washed for 1
hour at 4ºC in four separate 50 mL volumes of PBS to remove unbound growth factor.
We confirmed that following the fourth wash, greater washing of up to ten washes
created no differences in bound growth factor quantity. Following lyophilization,
cryomilling, and solubilization of the tissue matrix in HCl-pepsin buffer solution, we
quantified EGF or HGF levels in each sample by ELISA (RayBiotech Inc., Norcross,
GA). Samples were normalized through measurement of total protein using the Pierce
BCA Protein Assay Kit (Thermo, Rockport, IL).
2.4 Liver ECM Solubilization
Gels with added ECM were created according to previously published protocols
from our group (8). In brief, decellularized liver tissue was frozen at -80ºC and
lyophilized using a SuperModulyo Freeze Dryer (Thermo, Waltham, MA). Samples were
maintained under vacuum at a pressure of 0.1 mbar and temperature of -80ºC for 24
hours. Tissue was then ground in liquid nitrogen using a mortar and pestle until a fine
98
powder was attained. Liver ECM was solubilized by mixing lyophilized decellularized
tissue with 0.1 N HCl and pepsin (Porcine gastric mucosa, 3400 units of protein, Fischer
Scientific, Fair Lawn, NJ). After an incubation of 48 hours at room temperature, the
solution was centrifuged and filtered with a 0.2 µm syringe filter. The pH was adjusted to
7.0 and the total protein concentration to 1.0 mg/mL.
2.5 Solubilized Liver ECM Quantitative Analysis
DNA from solubilized liver ECM solutions was isolated and purified using the
Qiagen DNeasy Kit. DNA amounts were quantified using a set of standards and the
Quant-iT PicoGreen reagent kit (Invitrogen Corp., Carlsbad, CA) that could detect lower
than 1 ng/mL of DNA in solution.
GAG levels were quantified using the Blyscan sGAG Assay kit (Biocolor,
Newtownabbey, UK). 40 μL of solubilized ECM was mixed with 160 μL of 0.2 M
sodium phosphate buffer containing 1 mg/mL of papain and incubated at 65ºC for 3
hours. Samples were then mixed with Blyscan dye, pelleted, and washed. After
dissociation reagent was added, samples and GAG standards were loaded into a 96 well
plate and measured in a SpectraMax M5 Multi-Mode Microplate Reader at an absorbance
of 656 nm (Molecular Devices, Sunnyvale, CA).
Collagen levels were quantified using the Sircol Soluble Collagen Assay kit
(Biocolor, Newtownabbey, UK). 20 μL of solubilized ECM was mixed with 130 μL of
0.5 M acetic acid containing 0.1 mg/mL pepsin and then incubated overnight at 4ºC.
Samples were then mixed with Sircol dye, pelleted, and washed. Dissociation reagent
99
was added, and samples and standards were read in a 96 well plate at an absorbance of
555 nm.
2.6 ECM Gel Formation
Liver ECM solution was combined at a 1:1 ratio with thiol modified HA gel
components (HyStem Hydrogels, ESI BIO, Alameda, CA). Components include 3,3′-
Dithiobis(propanoic hydrazide) modified HA (Glycosil or HA-DTPH, Mw 168 kDa, Mn
79 kDa, polydispersity index 2.13), chemically modified gelatin-DTPH (Gelin-S, Mw
∼50 kDa), and heparin-DTPH (Heprasil, Mw ∼17-19 kDa). Protocols for synthesis of
these proprietary compounds can be found in the literature (47, 48). HA alone does not
provide sites for attachment during cell seeding. By including gelatin, or denatured
collagen, and supplemented liver ECM, adequate binding sites for cells were supplied. At
a sufficient stiffness, these molecules enable timely and efficient cell adhesion that
resembles what is observed in seeding of tissue scaffolds or protein coated tissue culture
plastic. These additions allow the hydrogels to mimic the composition and physical
properties of a normal tissue microenvironment.
To test the effects of mechanical properties, three different gel stiffnesses were
created. For the two lower stiffnesses, 1% and 2% weight/volume polyethylene glycol
diacrylate (PEGDA/Extralink, 3.4 kDa) were added to other gel components (ESI BIO,
Alameda, CA). The stiffest gel was created with 6% weight/volume 4-arm PEG acrylate
(20 kDa) (Creative PEGWorks, Chapel Hill, NC). As controls, gels containing only
components of the HA gel kit were created without the addition of solubilized ECM.
100
Gel solutions were added to wells of chamber slides or tissue culture plates and
allowed to fully crosslink for two hours. Gels completely filled the growth area of the
wells and polymerized at a thickness of over 1.5 mm to prevent the tissue culture plastic
bottom from contributing to the stiffness of the substrate. For cell culture assays
involving immunocytochemistry (ICC), 125 μL of each gel type was layered per well into
8-well Permanox Plastic Nunc Lab-Tek Chamber Slides to fill an area 80 mm² (Thermo
Scientific, Waltham, MA). For stiffness quantification, 325 µL of gel was added to 24-
well plates to fill an area of 200 mm². For assays measuring function, viability, and gene
expression, 50 µL of each gel type was layered into 96-well plates to fill an area of 32
mm² for cell seeding (Tissue culture treated plastic, Thermo Scientific, Waltham, MA).
2.7 Stiffness Quantification
ECM-HA and HA only gels were created in 24-well plates (80 mm²) with each
crosslinker concentration (1% PEGDA, 2% PEGDA, and 6% 4-arm PEG w/v). The
Discovery Series HR-2 Rheometer (TA Instruments, New Castle, DE), a type of
rotational or shear rheometer, was used to test stiffness following creation of gel
substrates. Decellularized tissue was used as a reference control and comparison to
normal liver ECM in this study. During testing with this type of rheometer, the sample
was stabilized on the fixed stage of the rheometer. A geometry in the form of a 12 mm
flat parallel plate was subsequently lowered until it came into contact with the sample.
The rheometer performed oscillatory stress sweep tests where shear stress was
incrementally increased on the samples from 0.6 Pa up to 10.0 Pa with a constant
oscillatory frequency of 1 Hz and a normal axial force of 0.4 N, as has been previously
described (49-52).
101
From the various forces applied, an elastic modulus could be measured and
calculated by the instrument. The elastic modulus, or resistance to deformity, can be
expressed by three types of calculations; Young’s modulus, bulk modulus and shear
elastic modulus. We analyzed the shear elastic modulus, which describes how a sample
responds to opposing forces parallel to its surface and is calculated by dividing shear
stress by shear strain. Both the shear storage modulus (G’) and shear loss modulus (G’’)
were measured. The storage modulus describes the elastic properties or stored energy,
while the loss modulus describes the viscous characteristics or energy lost as heat (53).
The storage modulus (G’) from oscillatory shear was the main value utilized for reporting
stiffness characteristics of the hydrogels in our assays.
To calculate G’, measurements of phase lag (the phase shift between stress and
strain), stress, and strain are needed. The formulas listed in Table 1 were used to calculate
G’ and its different variables. Formulas were modified from the Discovery Series HR-2
Rheometer Reference Guide (TA Instruments New Castle, DE).
Table 1: Storage modulus formulas
Formulas Variables
G′ = 𝑠𝑡𝑜𝑟𝑎𝑔𝑒 𝑚𝑜𝑑𝑢𝑙𝑢𝑠 = cos(δ)σ
γ
δ = phase lag or shift, σ = oscillation stress, γ
= strain
σ = oscillation stress = Kσ∗M(sample)
Kσ = stress constant, M = oscillation torque
(on the sample)
Kσ = stress constant = 2
π∗R³
R = radius of parallel plate
γ = strain = Kγ∗Ѳ Kγ = strain constant, Ѳ = oscillation or
angular displacement
Kγ = strain constant = R
H
R = radius of parallel plate, H = gap
102
2.8 GAG Function in ECM gels
GAG function was analyzed in gel substrates of the same 600 Pa stiffness in 96
well plates either containing HA only or HA supplemented with solubilized liver ECM.
Gels were incubated in solutions containing 500 ng/mL of HGF in PBS overnight and
then washed for an hour in four separate 250 uL volumes of PBS. A HGF dependent
epithelial cell line, 4MBr-5, was seeded at 20,000 cell/well in Ham's F-12K Medium with
2 mM L-glutamine, 1.5 g/L sodium bicarbonate, and 30 ng/mL of epidermal growth
factor (ATCC, Manassas, VA). DNA synthesis of seeded cells was quantified after 48
hours in culture to assess the activity of GAG-bound HGF. Values were also standardized
by cells grown on non-growth factor treated gels of each substrate type.
2.9 Primary Hepatocyte Isolation and Seeding
In preliminary assays, primary rat hepatocytes were freshly isolated from rat
livers perfused with collagenase and calcium chloride solutions through the inferior vena
cava at 37ºC. To purify isolations, hepatocytes were passed through a 100µm nylon mesh
and centrifuged at 1000RPM as described in previous studies (102, 103).
For assays involving primary human hepatocytes, cryopreserved human
hepatocytes (Triangle Research Labs, Research Triangle Park, NC) were seeded onto gels
in Clonetics Hepatocyte Basal Medium with UltraGlutamine-1 and Hepatocyte Culture
Medium supplements (HCM; Lonza, Allendale, NJ). Supplements include ascorbic acid,
bovine serum albumin-fatty acid free, hydrocortisone, human epidermal growth factor,
transferrin, insulin, and gentamicin. 50,000 or 125,000 primary hepatocytes per well were
seeded on pre-formed gels in 96-well plates or 8-well chamber slides respectively. Cells
103
were allowed to attach to gel binding sites for 5 hours before PBS washing removed dead
or non-adherent cells. Hepatocytes were maintained in incubators (37ºC, 5% CO2), and
media was changed following PBS washing every 2 days until conclusion of the
experiment.
2 and 7 days post-seeding were used as time points in all of our cellular assays.
The majority of cell death occurs in the first 24 hours post-hepatocyte seeding (54). The
day 2 time point allows for initial cell attachment, establishment of cell viability, and
washing away of non-adherent or dead cells. Primary hepatocytes have also been
reported to take 3-6 days to stabilize metabolic functions, especially albumin production
(55). The day 7 time point allows cells to complete this adaptation period on the cell
culture substrates.
2.10 DNA Quantification
DNA was extracted from 4MBr-5 cells of each well by proteinase K digestion for
2 hours at 56ºC and purified using the Qiagen DNeasy Kit. DNA was quantified using the
Quant-iT PicoGreen reagent kit and a set of DNA standards (Invitrogen Corp., Carlsbad,
CA). Total DNA amounts were standardized by DNA levels from cells grown on non-
growth factor treated HA only or ECM containing gels.
Similar to the GAG-bound HGF assay, DNA was quantified in primary
hepatocytes to assess cell numbers retained on gels at various time points. DNA was
extracted and purified from cells of each well at the day 2 or day 7 post-seeding time
points. DNA was then quantified using the Quant-iT PicoGreen reagent kit (Invitrogen
104
Corp., Carlsbad, CA). Total DNA amounts were compared among various stiffness
groups.
2.11 Immunocytochemistry (ICC)
For ICC, cells were seeded on gels in Nunc™ Lab-Tek™ 8-well Chamber Slides.
After 7 days in culture, cells were fixed overnight in 10% neutral buffer formalin (NBF)
at 4ºC. Cells were then stained using the Actin Cytoskeleton and Focal Adhesion Staining
Kit (Millipore, Billerica, MA), or for FAK and ILK staining, the rabbit monoclonal
antibodies anti-FAK (EP695Y) and anti-ILK (EPR1592) (Abcam, Cambridge, MA) were
used. Cells were blocked for 30 minutes in 8% milk and then stained with a 1:500
dilution of the anti-Vinculin, 1:500 dilution of anti-FAK, or 1:250 dilution of anti-ILK
primary antibody for 1 hour. After washing, anti-Vinculin cells were then stained with
1:500 diluted TRITC-conjugated Phalloidin and 1:500 Goat anti-Mouse IgG (H+L)
Secondary Antibody, Alexa Fluor 488 conjugate (Life Technologies, Grand Island, NY).
Fluorescent-conjugated phalloidin stained F-actin in the microfilaments of the
cytoskeleton to assess cell structure. Cells incubated with Anti-FAK or Anti-ILK were
stained with 1:500 Goat anti-Rabbit IgG (H+L) Secondary Antibody, Alexa Fluor 594
conjugate. Following further washing, all cells were incubated in 1:1000 DAPI
counterstain for 5 minutes. DAPI co-staining was used to visualize individual cell nuclei
for quantification.
2.12 Cell Counting
To confirm DNA quantification methods at 7 days post-seeding, hepatocytes were
quantified by counting nuclei of stained cells per 0.25 mm² area. For each gel type and
105
stiffness, six separate 10x magnification images (1000 μm x 1000 μm) of DAPI stained
hepatocytes were captured in randomized areas of six different substrate-containing wells
of an 8-well Permanox Plastic Nunc Lab-Tek Chamber Slide (Thermo Scientific,
Waltham, MA). Images were broken down into four quarters (500 μm x 500 μm), and the
top left quarter of the image was used for counting purposes (0.25 mm² area). DAPI
stained cells were hand-counted per image and averaged to assess cell adhesion for each
substrate condition.
2.13 Calcein-AM Staining
Primary hepatocytes seeded on ECM-HA or HA only containing gels were
immunofluorescently labeled using a Live/Dead Viability Assay Kit (Life Technologies,
Grand Island, NY). Viable cells were identified by green-fluorescent Calcein-AM
positive staining.
2.14 Hepatocyte Functional Analysis
Media samples were collected at various time points from wells containing
primary hepatocytes seeded on gels of varying stiffness. The Human Serum Albumin
ELISA kit (Alpha Diagnostic International, San Antonio, TX) was used to compare
secreted albumin in experimental groups to negative controls containing medium alone.
Quantification was achieved by comparing values to a standard curve generated from
purified albumin.
106
2.15 Liver Cell Junction and Kinase Expression
Primary hepatocytes seeded on various gel types were lysed by syringe
homogenization in QIAzol solution. Solutions from triplicate wells were combined to
create a total volume of 700uL of homogenized QIAzol/cell mixture. A total of 140 µL of
chloroform was added to this solution and vortexed for 15 seconds. Solutions were then
centrifuged at 12,000 x g at 4ºC for 15 minutes. The upper aqueous layers were added to
miRNeasy spin columns, and processed RNA was used for real time PCR (Qiagen,
Germantown, MD).
Isolated RNA was quantified using a NanoDrop 2000 UV-Vis Spectrophotometer
(Thermo Scientific, Waltham, MA). A total of 200 ng RNA was used to create cDNA
using a thermocycler and the SuperScript Vilo cDNA Kit (Life Technologies, Grand
Island, NY). cDNA was mixed with the SYBR Select Master Mix (Life Technologies,
Grand Island, NY) and Eurofin primers custom created from sequences designed using
Primer-Blast (NCBI) (Table 2) or from published literature (56, 57). An Applied
Biosystems 7300 Real-Time PCR System was used to calculate samples’ Ct values for
targets, which were standardized by GAPDH. Expression was compared relative to
Human Liver Total RNA (Life Technologies, Waltham, MA).
107
Table 2: Human primers for RT-qPCR
Targets Forward Primer (5’ to 3’) Reverse Primer (5’ to 3’)
Claudin-1 CCGTTGGCATGAAGTGTATG CCAGTGAAGAGAGCCTGACC
E-Cadherin TGAGTGTCCCCCGGTATCTT AGAATCATAAGGCGGGGCTG
ILK AAGGTGCTGAAGGTTCGAGA ATACGGCATCCAGTGTGTGA
FAK (PTK2) TTATTGGCCACTGTGGATGA TACTCTTGCTGGAGGCTGGT
GAPDH (56) CCCACTCCTCCACCTTTGAC TTGCTGTAGCCAAATTCGTTGT
Integrin Beta 1
(57) GAAGGGTTGCCCTCCAGA GCTTGAGCTTCTCTGCTGTT
RhoA GTCCACGGTCTGGTCTTCAG TTTCCACAGGCTCCATCACC
CDC42 variant
1/2 GGTGGAGAAGCTGAGGTCAT CATCGCCCACAACAACACAC
Rac1/3 ATCACCTATCCGCAGGGTCT CGGATCGCTTCGTCAAACAC
HNF4α TCAACCCGAGAAAACAAACC ACCTGCTCTACCAGCCAGAA
Albumin TTGGCACAATGAAGTGGGTA AAAGGCAATCAACACCAAGG
SNAIL CCCTCAAGATGCACATCCGAA GACTCTTGGTGCTTGTGGAGCA
2.16 Statistics
Data was expressed as the mean +/−standard deviation. Statistical analysis was
performed using Student’s t test and analysis of variance (ANOVA) where noted. A p-
value of less than 0.05 was considered significant.
108
3. Results
3.1 Liver Decellularization and Solubilization Preserved ECM Components and Their
Functions
This study incorporated liver ECM into gels through the process of
decellularization and solubilization. A previous manuscript published by our group
demonstrated full cellular removal and retention of specific ECM components, including
B)
0
100
200
300
400
500
0 50 100 500 1000
pg
HG
F/g
EC
M
HGF Pretreatment Concentration (ng/mL)
Growth Factor Binding C)
0
10
20
30
40
50
60
70
HA ECM
To
tal D
NA
(n
g)
HGF Treated Gel Type
HGF-Dependent Cell Growth D)
A)
Fig.2) Solubilized Liver Composition and Activity- A) Alcian blue staining detected GAG presence in
decellularized tissue. B) Solubilized liver ECM solutions were analyzed for collagen, GAG, and DNA
content (n=3, ±SD). DNA content was non-detectable (ND) at an assay limit of <1 ng/mL. C) During
growth factor treatments to determine GAG functionality, HGF bound to isolated ECM in a dose-
dependent manner (*p<.05, Student’s t-test, n=5). D) The activity of growth factor treated gels was
tested by seeding a HGF-dependent cell line on HA gels or HA gels supplemented with solubilized ECM
after 500 ng/mL HGF pre-treatment. Cell growth was quantified by DNA (*p<.05, Student’s t-test, n=7).
109
collagen IV and laminin, in the decellularized liver tissue (Figure 1-Methods) (10).
Alcian blue staining was performed in a new IHC assay to confirm presence of GAGs in
the tissue (Figure 2A). GAG-containing decellularized matrix also demonstrated the
ability to bind HGF in a concentration dependent manner (Figure 2C). In addition, the
GAG binding growth factor HGF abound in greater quantities compared to EGF, which
only has non-specific interactions with collagens of the matrix (Supplementary Figure 1).
Following liver ECM solubilization, key molecular constituents were quantified.
DNA levels were insignificant and below detection levels (<1 ng/mL). However,
significant levels of GAGs and collagen were detected (Figure 2B). An assay was
performed to measure the ability of GAGs in solubilized liver ECM gels to bind and
present growth factor. The assay showed increased growth of a HGF-dependent epithelial
cell line on HGF pre-treated HA gels supplemented with solubilized ECM compared to
pre-treated gels only containing HA (Figure 2D).
In cell culture experiments with rat hepatocyte, substrates with solubilized ECM
improved viability compared to gels containing cryomilled tissue (Figure 3).
Incorporation of decellularized liver tissue into HA cell culture substrates also showed
benefits over single protein coatings or unmodified gel substrates. Preliminary studies
showed greater hepatocyte aggregation on HA gels containing liver ECM versus basic
collagen I gels (Figure 4). Compared to cells grown on gels containing solely HA,
primary hepatocytes seeded on HA gels supplemented with solubilized ECM attached in
greater numbers and showed greater E-cadherin expression (Figure 4E and 4F).
110
Fig.3) Primary rat hepatocytes were seeded for 7 days on gel coatings with
A+C) powdered or B+D) solubilized liver ECM. Ethidium bromide levels stained
dead cells red, while calcein-AM stained live cells green. Cell viability was A) low
on gels with powdered ECM and B) high on solubilized ECM gels. Phase contrast
images showed C) vacuolization and development of lamellar bodies in
hepatocytes grown on powdered ECM. D) Hepatocytes on solubilized ECM gels
maintained normal cell morphologies.
111
Fig.4) Primary rat hepatocytes were seeded for 7 days on either A+C) pure collagen I gels or
B+D) HA gels containing solubilized liver ECM. Phase contrast imaging and cytochrome
P450 staining with a DAPI counterstain show greater cell aggregation on liver ECM gels.
ECM gels also initiated greater E) cell attachment and F) E-cadherin expression (*p<.05,
**p<.01, Student’s t test, DNA n=10, E-cadherin n=3).
112
3.2 Crosslinker Concentration Altered ECM Gel Stiffness
1% PEGDA, 2% PEGDA, and 6% 4-arm PEG (w/v) crosslinker formed three
different ECM gel stiffness groups for stiffness testing. Oscillatory stress rheometry
quantified the storage modulus (G’) to compare each crosslinked gel’s stiffness to the
stiffness of normal liver ECM. Gels formed by 6% 4-arm PEG had a mean G’ of 4600
Pa. This value was higher than the mean stiffness of normal liver ECM, which fluctuated
around 3000 Pa, but fell within the standard deviation of the data set. The stiffnesses of
gel groups formed by 1% and 2% PEGDA at 600 and 1200 Pa respectively were
signifcantly lower than the storage moduli of both the 6% 4arm PEG crosslinked gels and
normal liver ECM. The 600 and 1200 Pa gel groups were also signicantly different from
each other (Figure 5).
Fig.5) HA-ECM Gel Stiffnesses Formed by Varying Crosslinker Concentrations-
HA-ECM gels were formed with 1% PEGDA, 2% PEGDA, or 6% 4-arm PEG (20kDa). All gel
stiffness values were significantly different from each other. Decellularized liver tissue had a
significantly higher stiffness than 1% and 2% PEGDA, however, it was not significantly
different compared to the 6% 4-arm PEG stiffness. (*p<.05, Student’s t test, n=4,)
0
1000
2000
3000
4000
5000
6000
7000
PEGDA (3.4kDa) 1% PEGDA (3.4kDa) 2% 4-arm PEG (20kDa) 6% Decellularized Liver
G' (P
a)
Crosslinker Concentration % (w/v)
HA-ECM Gel Stiffness
113
3.3 ECM Supplementation and Stiffness Affected Viable Hepatocyte Attachment and
Morphology
Calcein-AM, which is only metabolized in the cytoplasm of live cells, was used to
stain hepatocytes to assess the role of ECM gel stiffness in cell viability and morphology.
An increase in viable human hepatocytes was observed on ECM containing gels with
increased stiffness at day 7 post-seeding. Hepatocytes also survived on ECM gels in
greater numbers than cells grown on gels containing HA alone (Figure 6).
Cell morphology varied among substrate groups. No obvious differences were
observed between cells seeded on varying stiffnesses of gels containing only HA (Figure
6A, 6C, and 6E). However, cells on ECM supplemented gels of 4600 Pa developed
organized, polyhedral morphologies compared to irregularly-shaped cells on the lower
stiffnesses of 600 and 1200 Pa. Increased cytoplasmic protrusions or filopodia formation
at day 7 post-seeding was demonstrated with Calcein-AM staining on 600 and 1200 Pa
ECM gels (Figure 6B and 6D).
114
Fig.6) Cell Viability and Filopodia Formation- Green-fluorescent Calcein-AM staining was
used to visualize primary hepatocyte viability and morphology at Day 7 post-seeding at 4x
magnification. Mean gel stiffnesses were; A+B) 600 Pa, C+D) 1200 Pa, or E+F) 4600 Pa.
Gels were composed of only HA (A,C,E) or were supplemented with liver ECM (D,B,F).
Filopodia were highlighted with white arrows.
115
Fig.7) Cytoskeletal Staining- Phalloidin and DAPI staining was used to visualize F-actin
filaments and nuclei of primary hepatocytes at day 7 post-seeding at 10x and 20x
magnification. Mean gel stiffnesses were; A+B) 600 Pa, C+D) 1200 Pa, or E+F) 4600 Pa.
Gels were composed of only HA (A,C,E) or were supplemented with liver ECM (D,B,F). Actin
stress fibers were highlighted by white arrows.
116
More detailed immunocytochemistry was then used to analyze hepatocyte
attachment and cytoskeletal structure of the hepatocytes. Clear differences were observed
in phalloidin staining of the hepatocytes’ actin cytoskeletons on varying ECM gel
stiffnesses. Increased gel stiffness correlated with increased attachment and cytoskeletal
organization, evidenced by greater actin microfilament presence and stress fiber
formation (Figure 7F). ECM supplemented gels of 600 and 1200 Pa stiffnesses generated
lower cell densities and less actin staining (Figure 7B and 7D). Differences in
cytoskeletal structure were less obvious in cells seeded on HA only gels but in general,
cells on the 4600 Pa gel stiffness experienced greater stress fiber formation than cells on
the 600 and 1200 Pa stiffnesses (Figure 7E).
From the DAPI staining, cells were counted by number of nuclei per mm² of gel.
Results showed a significant increase in cell attachment with increased ECM gel
stiffness. Stiffness did not significantly influence cell number on gels composed of only
HA. The data also indicated significant differences in attachment between ECM-
containing and HA-only gels at 600 Pa and 4600 Pa. Human hepatocytes attached with
greater efficiency on HA gels at 600 Pa. At 4600 Pa, a more physiologically normal
stiffness, hepatocytes attached best to gels containing ECM (Figure 8A).
An additional assay was used to validate these findings. In this assay, DNA was
isolated and quantified from different stiffness groups at two different time points. ECM
gels at day 2 did not show significant differences. However, by day 7, 4600 Pa ECM gels
showed significantly greater DNA levels (Figure 8B).
117
3.4 ECM Gel Stiffness Altered Human Hepatocyte Marker Expression and Functionality
over Time
Expression levels of the hepatocyte markers HNF4α and albumin on gels of varying
stiffnesses were measured and normalized to native human liver expression. There were
no significant differences in HNF4α expression at the day 2 time point (Supplementary
Figure 2A). However, at day 7 post-seeding, cells expressed both HNF4α and albumin at
significantly lower levels on the 4600 Pa gel stiffness, while cells on the 1200 Pa stiffness
demonstrated increased expression of these genes (Figure 9). SNAIL expression was also
quantified to determine if any stiffness-induced epithelial-mesenchymal transition was
0
100
200
300
400
500
600
700
800
900
600 1200 4600
Ce
lls
/mm
²
Gel Stiffness (Pa)
Cell Attachment on Varying Gel Types
HA
ECM
A)
0
10
20
30
40
50
60
70
80
600 1200 4600
DN
A p
er
we
ll (
ng
)
Gel Stiffness (Pa)
DNA Quantification on ECM Gels
Day 2
Day 7
B)
Fig.8) Stiffness and Substrate Dependent Cell Attachment- A) Cell attachment was
assessed at 7 days post-seeding on HA gels of various stiffnesses, with or without ECM
supplementation. Cell number increased with stiffness in ECM gels. There were also
significant differences in cell numbers between simple HA gels and HA gels containing liver
ECM at the 600 and 4600 Pa stiffness levels. B) DNA of cells attached to ECM containing
HA gels was quantified to confirm cell counting methods and to test an earlier time point of 2
days post-seeding. A significant decrease in DNA was measured on the 1200 Pa stiffness
between day 2 and 7 of culture. However, at the 4600 Pa ECM gel stiffness, no loss in
viable cells was observed over this same period of time. (*p<.05, **p<.01, ***p<.001,
Student’s t test, n=6)
118
occurring. No significant differences were seen at either time point (Supplementary
Figure 2B and 2D).
ELISA for albumin production was used as a measure of human hepatocyte
function, and to determine if functional output matched hepatocyte marker expression. A
significant increase in albumin production was seen with increasing gel stiffness at day 2
post-seeding. This increase remained significant when albumin was normalized to total
DNA (Figure 10A). By day 7, albumin production improved in cells seeded on all gel
stiffnesses, but cells on the 4600 Pa gel stiffness no longer produced albumin at a
significantly higher rate. Cells on the 1200 Pa stiffness created albumin at a significantly
higher rate than the lowest stiffness and appeared to overtake the functional rate of cells
on the 4600 Pa stiffness (Figure 10B).
0
0.1
0.2
0.3
0.4
0.5
0.6
600 1200 4600
Fo
ld C
ha
ng
e (
Re
lati
ve
to
Liv
er
Ex
pre
ss
ion
)
Gel Stiffness (Pa)
Human ALB Expression B)
0
0.1
0.2
0.3
0.4
0.5
0.6
600 1200 4600
Fo
ld C
ha
ng
e (
Re
lati
ve
to
Liv
er
Ex
pre
ss
ion
)
Gel Stiffness (Pa)
Human HNF4α Expression A)
Fig.9) Hepatocyte Phenotypic Marker Expression- RT-qPCR measured expression of the
hepatic markers A) HNF4α and B) albumin in primary human hepatocytes on liver ECM
supplemented HA gels of varying gel stiffnesses relative to normal liver expression. Cells on the gel
stiffness of 4600 Pa had significantly lower expression of both markers compared to the 600 and
1200 Pa stiffnesses at day 7 post-seeding (**p<.01, ***p<.001, Student’s t test, n=3).
119
3.5 Stiffness Affects Cell Morphology and Structure through Cytoplasmic Kinase
Expression
To further characterize underlying mechanisms of primary hepatocyte
organization and morphology on various ECM gel stiffnesses, cell junction maintenance
was determined by RT-qPCR. Data was again normalized to expression in normal human
liver. Claudin-1 is an important protein in tight junction formation and maintenance in
epithelial cells. Primary hepatocytes grown on the 4600 Pa stiffness for 7 days expressed
the highest levels of claudin-1. Hepatocytes on all stiffnesses expressed claudin-1 at
levels greater than those measured in normal liver (Figure 11A).
Expression of occludin, another constituent of tight junctions, also proved to be
dependent on substrate stiffness, but in an inverse manner. Expression of occludin
decreased relative to stiffness, although all values were at or above levels of expression
seen in normal liver (Figure 11B). Taken together, the expression patterns of occludin
Fig.10) Hepatocyte Functional Output- ELISA measured albumin secretion of primary human
hepatocytes per well on ECM containing HA gels of varying stiffnesses at A) day 2 and B) day 7
post-seeding. Quantities were standardized for cell number by DNA quantity per well (*p<.05,
**p<.01, Student’s t test, n=12,).
0
0.5
1
1.5
2
2.5
600 1200 4600
g A
lbu
min
/g D
NA
Gel Stiffness (Pa)
Albumin Secretion- Day 2 A)
0
2
4
6
8
10
12
600 1200 4600
g A
lbu
min
/g D
NA
Gel Stiffness (Pa)
Albumin Secretion- Day 7 B)
120
and claudin-1 have alternative roles in adapting primary hepatocyte phenotype and
structure to substrate stiffness.
To determine the relationship between cell junction gene expression and integrin
sensing, RT-qPCR was used to analyze cytoplasmic kinase activation on varying
substrate stiffnesses. At early time points, cells expressed FAK and ILK similarly
regardless of substrate stiffness (data not shown). However, at 7 days post-seeding, the
highest expression levels of FAK and ILK were measured on the 600 Pa substrates
0
0.5
1
1.5
2
2.5
600 1200 4600Fo
ld C
ha
ng
e (
Re
lati
ve
to
Liv
er
Ex
pre
ss
ion
)
Gel Stiffness (Pa)
Human Occludin Expression B)
0
1
2
3
4
5
6
7
8
600 1200 4600Fo
ld C
ha
ng
e (
Re
lati
ve
to
Liv
er
Ex
pre
ss
ion
)
Gel Stiffness (Pa)
Human Claudin-1 Expression A)
0
1
2
3
4
5
600 1200 4600
Fo
ld C
ha
ng
e (
Rela
tive
to
Liv
er
Ex
pre
ss
ion
)
Gel Stiffness (Pa)
Human ILK Expression D)
0
0.5
1
1.5
2
600 1200 4600Fo
ld C
ha
ng
e (
Rela
tive
to
Liv
er
Ex
pre
ss
ion
)
Gel Stiffness (Pa)
Human FAK Expression C)
Fig.11) Expression of Cell Junction Molecules and Focal Adhesion Regulators- RT-qPCR
measured expression of A) Claudin-1, B) Occludin, C) FAK, and D) ILK in primary hepatocytes
grown on ECM containing HA gels of varying stiffnesses at 7 days post-seeding. Expression was
standardized relative to normal liver expression (*p<.05, Student’s t test, n=3).
121
Fig.12) A,C,E) FAK and (D,B,F) ILK staining was performed on primary
hepatocytes at day 7 post-seeding at 20x magnification. Mean ECM gel
stiffnesses were; A+B) 600 Pa, C+D) 1200 Pa, or E+F) 4600 Pa. FAK and
ILK stained in red, along with autofluorescent nuclei of dead cells. DAPI
counterstaining stained all nuclei in blue.
(Figure 11C-D). Hepatocytes grown on the 4600 Pa gels expressed FAK and ILK at
levels similar to and greater than physiological liver levels, respectively. On 600 and
1200 Pa stiffnesses, both kinases were expressed at higher than normal liver levels.
122
To correlate gene expression with protein expression, ICC for FAK and ILK was
performed on cells seeded in the same conditions and ECM gel stiffnesses. FAK staining
was most prominent at the leading edge of cells at the 600 Pa gel stiffness (Figure 12A).
Staining was also seen on the 1200 Pa stiffness (Figure 12B). Cell staining for ILK was
less specific, but showed increases on the 600 and 1200 Pa stiffness gels (Figure 12B and
Figure 12D).
3.6 Rho GTPase Expression Correlates with Cytoplasmic Kinase Overexpression and
Hepatocyte Morphology
Next, we quantified expression of vital downstream targets of FAK and ILK that
are responsible for initiating actin polarization and polymerization to alter cell
morphology. RT-qPCR was performed on ECM gels of varying stiffness to measure the
Rho GTPases; Rho, Rac, and CDC42. Similar to FAK and ILK, CDC42 expression was
highest in hepatocytes seeded on the 600 Pa gel stiffness (Figure 13A). In contrast, cells
on the 4600 Pa substrates expressed Rho at the highest levels (Figure 13B). Rac
expression was not altered with stiffness (Figure 13C).
123
4. Discussion
The challenge of maintaining primary hepatocyte phenotype in vitro has led to the
development of more advanced culture systems. Traditional cell culture methods utilizing
collagen coated tissue culture plastic experienced loss of primary hepatocyte viability and
function by 7 days post-seeding (16). Cells grown on substrates supplemented with
natural liver ECM maintain viability and function for extended time periods as compared
to these traditional substrates that contain a limited number of matrix proteins (6).
However, studies have yet to completely optimize methods for incorporating ECM into
culture substrates and the mechanical properties of these substrates. Development of liver
specific gels supplemented with ECM has been slowed by the lack of a reliable method
0
0.5
1
1.5
2
2.5
600 1200 4600Fo
ld C
ha
ng
e (
Re
lati
ve
to
Liv
er
Ex
pre
ss
ion
)
Gel Stiffness (Pa)
Human CDC42 Expression A)
0
0.5
1
1.5
2
2.5
600 1200 4600Fo
ld C
ha
ng
e (
Rela
tive
to
Liv
er
Ex
pre
ss
ion
)
Gel Stiffness (Pa)
Human Rac1 Expression C) Fig.13) Expression of the Cytoskeleton-
Regulating Rho GTPases- RT-qPCR
measured expression of the Rho GTPases;
A) CDC42, B) RhoA, and C) Rac1 in primary
hepatocytes grown on ECM containing HA
gels of varying stiffnesses at 7 days post-
seeding. Expression was standardized relative
to normal liver expression (*p<.05, Student’s t
test, n=3).
0
0.5
1
1.5
2
2.5
600 1200 4600Fo
ld C
ha
ng
e (
Re
lati
ve
to
Liv
er
Ex
pre
ss
ion
)
Gel Stiffness (Pa)
Human RhoA Expression B)
124
for purifying ECM that retained all of the bioregulatory properties of the native matrix.
Perfusion decellularization solved this problem by allowing selective removal of all of
the cellular components of a tissue, while leaving the remaining acellular material
relatively unperturbed and functional as characterized in Figure 1 and Figure 2 (10).
Initial attempts to incorporate cryomilled ECM into substrates yielded suboptimal
results. Powders accumulated in endosomes and compromised the health of the cells
(Figure 3). This build-up of phagocytosed matrix likely caused congestion of intracellular
lysosomes, vacuolization, and induction of apoptosis in highly endocytic hepatocytes (58-
60). Solubilization of the ECM allowed for easy integration into HA hydrogel substrates
and circumvented the negative effect that particulate ECM had on primary hepatocytes
(Figure 3). HA-based gels have also proven to be non-immunogenic following
implantation into experimental animals (11, 14), opening up the possibility to use these
substrates in future cell based therapies.
Preliminary studies with rat hepatocytes also showed greater viability, cell
aggregation, and expression of E-cadherin on ECM supplemented HA gels compared to
standard gels containing only collagen I gels or HA (Figure 4). These features are
especially important in primary hepatocytes, which rely on cell contacts and junction
formation for functional maintenance and cell polarity (61-63). E-cadherin (E-Cad) is a
calcium-dependent glycoprotein vital in epithelial intercellular adhesion, liver tissue
architecture, and hepatocyte functional maintenance (64, 65). This glycoprotein has also
been correlated with reduction in anoikis, or detachment related apoptosis, during culture
of primary hepatocytes (66) . Increased E-cadherin expression on our ECM gels indicates
the abilities of these substrates to support proper hepatocyte phenotype.
125
In our chief study with human cells, ECM supplementation of HA gels was shown
to positively affect primary hepatocyte attachment and viability. Inclusion of solubilized
ECM increased cell coverage of the gels at two of the three stiffnesses tested (Figure 8).
ECM contains several different molecules that promote cell adhesion. Many of these cell
attachment proteins interact with cell membrane integrins through Arg-Gly-Asp (RGD)
motifs. RGD-containing molecules include laminin, fibronectin, vitronectin, and several
of the collagen sub-types. (67). RGD peptides support survival of hepatocytes via the
beta1-integrin-ILK-pAkt pathway (68). The results of the current study indicate that a
strong correlation exists between supplementation of HA substrates with ECM and
integrin beta-1 expression, which may explain why liver ECM mediates efficient
hepatocyte attachment (Supplementary Figure 3) (68). Several other ligands or cell
adhesion molecules contribute to cell-ECM interactions, but promotion of integrin
expression clearly represents a key factor in forming and maintaining cell/substrate
attachment.
The mechanical properties of gel substrates also play a significant role in the
preservation of hepatocyte cell viability and function. Previous studies on hepatocellular
carcinoma cell lines showed increased proliferation on stiff polyacrylamide gels (defined
as 12 kPa) as compared to soft gels (1kPa) (24). A study on primary rat hepatocytes
compared growth on what was defined as soft (11 kPa), medium (27 kPa), and stiff (116
kPa) heparin gels (69). In that study, the softest gel was well above the stiffness of
normal liver, which has been reported by various groups as ranging anywhere from 1.5 to
8 kPa (19-24). The study concluded that “soft” gels promoted greater cell attachment,
better morphology, higher albumin output, and more robust cell aggregation (69).The
126
lack of consensus on normal liver stiffness likely arises from variations in tissue
preparation, instrumentation, and how the measurement was obtained. Similar
discrepancies exist in the reported stiffness of fibrotic livers, which have varied from 8
kPa to 75 kPa (20, 70-73). The variance in definitions of soft and stiff gel substrates
complicate the direct comparison of results among various studies. An internal control of
normal liver ECM was used in the current study, allowing for a point of reference that
takes into account the equipment and methods used.
The gels employed in the current study use different crosslinker formulations to
adjust gel stiffness, rather than varying the matrix concentration. This minimizes
potential confounding effects such as the number of bioactive sites, diffusion rates, and
gel swelling properties. The study’s goal was to determine an optimal stiffness range for
maintaining the viability and function of primary hepatocytes in the presence of normal
liver ECM. The two lower crosslinker concentrations produced sub-normal stiffnesses at
600 and 1200 Pa, which would be representative of matrix undergoing remodeling in
scenarios such as recovery from injury or during development. The highest gel stiffness
at 4600 Pa was marginally higher than what was measured as the average stiffness of
decellularized liver ECM (3000 Pa), but not significantly different. Intact liver tissue had
a significantly higher stiffness, which can be attributed to the presence of cells organized
within the structure. The decellularization process could potentially soften the isolated
liver ECM causing lower values than non-processed ECM (74). However, histological
analysis confirmed the presence of all major structural molecules, suggesting that any
difference in stiffness should be minimal (10). A more precise measurement of the
contribution of ECM to the total stiffness of specific tissues would be beneficial to organ
127
bioengineering strategies that use decellularized natural scaffolds. The three stiffnesses
used in the current study provide a good indication of interactions of primary human
hepatocytes cultured within a narrow, physiologically relevant stiffness range.
In the ECM supplemented gels, primary hepatocyte attachment and growth was
maximized at the 4600 Pa gel stiffness level. Attachment not only correlated with
increased presence of ligand sites, but also the stiffness of the substrate. Stiffer substrates,
in general, provide more stable attachment sites for cells (15). They also provide good
traction for cell motility, but hamper cell detachment. Soft substrates allow cells to detach
more easily, although they provide lower traction for cell migration (75, 76). Cells sense
stiffness through mechanosensing of strain on their cytoskeletons by “outside in
signaling” mechanisms (40, 77). These forces exerted on their cell structure trigger
changes in morphology and phenotype (78-80). Human hepatocytes on the two softer
ECM stiffnesses of 600 and 1200 Pa developed long protrusions and a disorganized
structure. Hepatocytes seeded on 4600 Pa gels exhibited an organized microstructure and
cuboidal morphology. These cells also developed actin stress fibers by day 7, which are
required for cell adhesion, cell junctions, and mechanotransduction (Figure 7). However,
a significant accumulation of stress fibers can eventually lead to phenotypic changes (79,
81). Cells seeded on HA gels without ECM did not display these same stiffness
dependent variations. It is possible that components of the ECM could affect the
activation state of cells and responsiveness to physical cues. In a time dependent manner,
results indicate that ECM supplementation and substrate stiffness both affect primary
hepatocyte attachment and morphology; the latter of which is determined by cytoskeletal
actin organization.
128
Increases in gel stiffness led to early increases in hepatocyte function on the ECM
gels. Greater stiffness improved total and normalized albumin secretion at day 2.
Normalized function of hepatocytes increased across all stiffness groups by day 7 post-
seeding. However, secretion on the 4600 Pa stiffness now matched albumin levels
measured on 1200 Pa gels. This correlated with a decrease on the 4600 Pa stiffness and
increase on 600 and 1200 Pa stiffnesses in gene expression of the hepatocyte markers
albumin and HNF4α. Cells may require time to establish themselves on a substrate before
modifying their gene expression levels and functional output. These data support the
notion that the highest metabolic function occurs on 4600 Pa gels, as shown by
hepatocyte plasma protein secretion at the earliest time point. However, over time a
slightly higher than physiological stiffness can lead to loss of phenotypic marker
expression, and function is surpassed by cells on a more intermediate stiffness. 600 and
1200 Pa stiffnesses could also promote active remodeling of the matrix to better suit
phenotypic maintenance after a certain period of time on the ECM substrates. These
alterations in cell function directly correlated with previously observed changes in
morphology. This association informed the design of experiments to study the
mechanisms of mechanosensing-induced cell signaling and the expression of cell junction
components.
As discussed previously, cell junction maintenance is vital for primary
hepatocytes function and polarity. Tight junctions are types of cell junctions that act as
chemical and electrical barriers, transduce cell/cell signals, and control permeability
across cell layers (82, 83). Claudin-1 is an integral component of epithelial tight
junctions. It has been shown to be critical to the maintenance of biliary function and the
129
prevention of epithelial-to-mesenchymal transition (EMT) (28, 82, 84, 85). Disruption of
claudin-1 expression can cause diseases such as neonatal sclerosing cholangitis, while
overexpression can restore tight junctions in fibrotic cells, resulting in restoration of cell
function (29, 86-88). The 4600 Pa gel stiffness promoted the greatest claudin-1
expression at the 7-day timepoint, correlating with more stable hepatocyte morphology.
Another component of tight junctions, occludin, responded to substrate stiffness
in a different manner. Cells expressed increased levels of occludin relative to normal liver
at all stiffness studied. However, in contrast to claudin-1 expression, occludin levels
decreased with increased gel stiffness. A threshold occludin level is required for
maintenance of cell polarity and morphology (89). Increased expression of occludin
promotes focal reorganization of tight junctions that leads to the formation of cell
protrusions which are critical for directional migration (30). The observed cell elongation
and increased occludin expression profiles at the 600 and 1200 Pa gel stiffnesses suggest
adoption of a pro-migratory phenotype observed previously at low substrate stiffnesses.
Two cytoplasmic kinases linked to cell junction organization and integrin-
mediated signaling, FAK and ILK, were also studied (90). FAK and ILK regulate cell
cytokine responses, cell growth, and cell viability (36, 91-93). They also modulate focal
adhesion turnover and formation of cell protrusion, which affect cell motility and
migration, by regulating the expression of the Rho GTPases: CDC42, Rac1, and RhoA
(30, 40, 77, 94, 95). Both ILK and FAK demonstrated increased gene expression at all
stiffness levels as compared to normal liver, while expression showed an inverse
correlation to gel stiffness. Expression levels were sufficient to allow survival at all
stiffness levels, but ICC showed marked increases in FAK expression and localization at
130
the leading edge of hepatocytes on the 600 Pa gel stiffness (Figure 12A). This result
reinforces previous studies which suggested FAK is required for assembly and direction
of the leading edge of motile cells (96). Greater expression of FAK, ILK, and occludin all
suggest adoption of a migratory phenotype at the lower stiffnesses of 600 and 1200 Pa.
This notion is further supported by measured upregulation of filopodia-initiating CDC42.
Decreases in FAK and increases in RhoA on 4600 Pa gels suggest favorable signaling for
establishment of focal adhesions, stress fiber formation, and cytoskeletal contraction (97).
Taken together, these results indicate stronger attachment and greater cytoskeleton
organization occur on gels at the 4600 Pa stiffness, while greater ability to detach and
move occurs on the softer gels of 600 and 1200 Pa. It is likely that the 1200 Pa gel
stiffness would allow for efficient hepatocyte migration, as both traction and
adhesion/release are in good balance. Understanding how cell morphology, attachment,
and migration are influenced by substrate stiffness and gel formulation are integral to the
design of cell substrates for cell based therapies and tissue engineering. Some
applications would require a static cell component, while others would benefit from
increased migratory potential for promoting cell organization and microarchitecture
remodeling.
Hepatocyte function was maximized at a specific stiffness depending on time in
culture. Future experiments could determine how longer time points or additional liver
cell types impact hepatocyte function in vitro at these particular ECM gel stiffnesses.
Additionally, regional variance in stiffness within a single organ should be taken into
account depending on the requirements of specific applications. There are also
implications for stiffness in models for aging and diseases, as these organs demonstrate
131
an accumulation of certain matrix components. Studies conducted by our group including
data in Figure 2 show that natural liver matrix has the potential to bind beneficial growth
factors such as hepatocyte growth factor (HGF) which can compensate for deficiencies
induced by inappropriate substrate stiffness and regulate function. A preliminary study
using isolated primary rat hepatocytes showed increased viability and cell junction
expression on 600 Pa ECM gels pretreated with EGF and HGF (Supplementary Figure 4).
Cell junction protein expression and localization vary according to cell type,
substrates stiffness, and growth factor microenvironment. Cadherins, integrins, and
growth factor receptors are transmembrane structures that form an interdependent
network of adaptor proteins, effector molecules, and cytoskeletal elements to regulate
signal transduction (98). Mechanical forces sensed by integrins of focal adhesion sites
lead to assembly or disassembly of cytoskeletal structures. Tension on the cytoskeleton
initiates conformational changes in cell-cell adhesions and growth factor receptors to
either promote or block cellular responses (99).
Many downstream cytoplasmic elements are also conserved between the various
transmembrane structures. In the case of HGF signaling, β-catenin complexes with both
the HGF receptor c-Met and E-cadherin. Results have varied on the effects of HGF
binding on E-cadherin expression. HGF induced E-cadherin loss and EMT in liver cells
during development and hepatocellular carcinomas leading to cell scattering or increased
tumor invasiveness (100-102). Studies which cultured primary hepatocytes showed
positive influences of HGF on E-cadherin expression and fibrosis prevention (103-105).
It has been theorized that polarized cells behave differently than other cell types because
of variances in the utilization and localization of β-catenin following HGF binding (106,
132
107). HGF-induced dissociation of β-catenin from c-Met could actually bolster the
formation of independent E-cadherin-β-catenin complexes and cell junctions in
hepatocytes seeded on certain substrate compositions and stiffnesses, which our data
possibly indicated (108).
Separate preliminary assays using human hepatic stellate cells demonstrated the
ability of stiffness to affect growth factor secretion and GAG production. Greater
stiffness led to increased GAG production by HSCs, while cells on lower stiffness levels
released higher quantities of HGF at day 2 post-seeding (Supplementary Figure 5). This
suggests that further refinement of both stiffness and substrate biochemical formulation
may be used to fine tune control of cell phenotype for specific therapeutic applications
and tissue engineering. Other factors such as substrate hydration kinetics and solute/ion
diffusion properties might also be modulated to control cell phenotype. A greater
understanding of how substrates and scaffolds influence cell phenotype would maximize
the efficacy of cell based therapies and would provide greater control of cells in tissue
engineering strategies.
5. Conclusions
HA substrates supplemented with solubilized liver ECM supported improved
growth factor binding capabilities, increased hepatocyte attachment and viability, and
increased phenotypic maintenance. These gels served as a solid foundation for studies of
liver interactions and cell function. Recreating the natural liver microenvironment in HA
hydrogels through supplementation with liver ECM revealed that hepatocyte function and
morphology changed over time based on substrate stiffness. Hepatocyte morphology and
133
expression profiles on sub-physiological stiffnesses of 600 and 1200 Pa were consistent
with cytoskeletal reorganization and filopodia formation. Gels at a higher stiffness of
4600 Pa had high metabolic and functional rates at early time points, but later developed
stress fibers and demonstrated a loss of hepatocyte gene expression. The optimal stiffness
for maintenance of long term primary hepatocyte function, in vitro, and in gel substrates
intended for the delivery of therapeutic cells is an intermediate stiffness between 1200
and 4600 Pa at the lower end of liver physiological limits.
6. Supplementary Figures
Supplementary Fig. 1) Growth factor binding comparison- Decellularized tissue was
incubated with EGF and HGF at a concentration of 500 ng/mL, and ELISAs quantified
growth factor binding. Residual growth factor amounts measured in non-treated samples
were subtracted. Sample measurements were standardized by DNA amount (**p>.01,
Student’s t test, n=6)
134
Supplementary Fig.2) RT-qPCR measured expression of A) HNF4α, B) SNAIL, and C) Claudin-1
in primary hepatocytes grown on ECM containing HA gels of varying stiffnesses at 2 days post-
seeding. No significant differences were found. D) SNAIL expression was also measured at day 7
post-seeding. Expression decreased from day 2, but no differences were found between stiffnesses.
Expression was standardized relative to normal liver expression (p>.05, Student’s t test, n=3).
135
Supplementary Fig. 3) A) RT-qPCR measured expression of integrin beta-1 in primary hepatocytes
grown on ECM containing HA gels of varying stiffnesses at 2 days post-seeding (n=3, Student’s t test,
*p<.05). B) Expression was also measured at day 7 post-seeding, averaged between all stiffnesses,
and compared between gels containing only HA and gels supplemented with ECM (n=9, Student’s t
test, *p<.05). All expression values were standardized relative to normal liver expression.
136
Supplementary Fig. 4) Primary rat hepatocytes were seeded for 2 days on 600 Pa HA gels
supplemented with liver ECM that were untreated or pretreated with combined solutions of
EGF and HGF. Calcein AM stained live cells green, or ethidium homodimer stained dead cells
red. Viability staining on A) untreated or B) pretreated ECM gels showed increased cell
viability with growth factor treatment. C) Cell attachment and D) E-cadherin expression also
increased on growth factor pretreated ECM gels (*p<.05, ***p<.001, DNA n=5, RT qPCR n=3)
137
Supplementary Fig. 5) Human HSCs were seeded for 2 days on HA gels supplemented with liver
ECM at mean stiffnesses of 600, 1200, and 4600 Pa. A) HGF levels per well were measured in the
media by ELISA. B) Total GAG production per well was measured by a colorimetric assay (*p<.05,
**p<.01, ***p<.001, HGF n=3, GAG n=5)
138
Works Cited
1. Atala A. Principles of regenerative medicine. 2nd ed. Amsterdam ; Boston: Elsevier, Academic Press; 2011. xix, 1182 p. p. 2. Bajaj P. Liver-assisting devices. Indian journal of anaesthesia. 2009;53(6):635-6. PubMed PMID: 20640088; PMCID: 2900070. 3. Baptista PM, Siddiqui MM, Lozier G, Rodriguez SR, Atala A, Soker S. The use of whole organ decellularization for the generation of a vascularized liver organoid. Hepatology. 2011:n/a-n/a. doi: 10.1002/hep.24067. 4. Ding YT, Shi XL. Bioartificial liver devices: Perspectives on the state of the art. Frontiers of medicine. 2011;5(1):15-9. doi: 10.1007/s11684-010-0110-x. PubMed PMID: 21088931. 5. Joly A, Desjardins JF, Fremond B, Desille M, Campion JP, Malledant Y, Lebreton Y, Semana G, Edwards-Levy F, Levy MC, Clement B. Survival, proliferation, and functions of porcine hepatocytes encapsulated in coated alginate beads: a step toward a reliable bioartificial liver. Transplantation. 1997;63(6):795-803. PubMed PMID: 9089217. 6. Hansen KC, Kiemele L, Maller O, O'Brien J, Shankar A, Fornetti J, Schedin P. An In-solution Ultrasonication-assisted Digestion Method for Improved Extracellular Matrix Proteome Coverage. Mol Cell Proteomics. 2009;8(7):1648-57. doi: DOI 10.1074/mcp.M900039-MCP200. PubMed PMID: WOS:000267960300015. 7. Zhou P, Lessa N, Estrada DC, Severson EB, Lingala S, Zern MA, Nolta JA, Wu JA. Decellularized Liver Matrix as a Carrier for the Transplantation of Human Fetal and Primary Hepatocytes in Mice. Liver Transplant. 2011;17(4):418-27. doi: Doi 10.1002/Lt.22270. PubMed PMID: WOS:000289004000009. 8. Skardal A, Smith L, Bharadwaj S, Atala A, Soker S, Zhang Y. Tissue specific synthetic ECM hydrogels for 3-D in vitro maintenance of hepatocyte function. Biomaterials. 2012;33(18):4565-75. doi: 10.1016/j.biomaterials.2012.03.034. PubMed PMID: 22475531; PMCID: 3719050. 9. Nakamura S, Ijima H. Solubilized matrix derived from decellularized liver as a growth factor-immobilizable scaffold for hepatocyte culture. J Biosci Bioeng. 2013;116(6):746-53. doi: DOI 10.1016/j.jbiosc.2013.05.031. PubMed PMID: WOS:000329087800016. 10. Shupe T, Williams M, Brown A, Willenberg B, Petersen BE. Method for the decellularization of intact rat liver. Organogenesis. 2010;6(2):134-6. PubMed PMID: 20885860; PMCID: 2901817. 11. Mirmalek-Sani SH, Sullivan DC, Zimmerman C, Shupe TD, Petersen BE. Immunogenicity of decellularized porcine liver for bioengineered hepatic tissue. Am J Pathol. 2013;183(2):558-65. doi: 10.1016/j.ajpath.2013.05.002. PubMed PMID: 23747949; PMCID: 3730770. 12. Zhang Y, He Y, Bharadwaj S, Hammam N, Carnagey K, Myers R, Atala A, Van Dyke M. Tissue-specific extracellular matrix coatings for the promotion of cell proliferation and maintenance of cell phenotype. Biomaterials. 2009;30(23-24):4021-8. doi: 10.1016/j.biomaterials.2009.04.005. PubMed PMID: 19410290. 13. DeQuach JA, Mezzano V, Miglani A, Lange S, Keller GM, Sheikh F, Christman KL. Simple and high yielding method for preparing tissue specific extracellular matrix coatings for cell culture. PLoS One. 2010;5(9):e13039. doi: 10.1371/journal.pone.0013039. PubMed PMID: 20885963; PMCID: 2946408. 14. Murphy SV, Skardal A, Atala A. Evaluation of hydrogels for bio-printing applications. J Biomed Mater Res A. 2013;101(1):272-84. doi: 10.1002/jbm.a.34326. PubMed PMID: 22941807. 15. Wells RG. The role of matrix stiffness in regulating cell behavior. Hepatology. 2008;47(4):1394-400. doi: 10.1002/hep.22193. PubMed PMID: 18307210.
139
16. Hansen LK, Wilhelm J, Fassett JT. Regulation of hepatocyte cell cycle progression and differentiation by type I collagen structure. Current topics in developmental biology. 2006;72:205-36. doi: 10.1016/S0070-2153(05)72004-4. PubMed PMID: 16564336. 17. Semler EJ, Ranucci CS, Moghe PV. Mechanochemical manipulation of hepatocyte aggregation can selectively induce or repress liver-specific function. Biotechnol Bioeng. 2000;69(4):359-69. PubMed PMID: 10862674. 18. Zaman MH, Kamm RD, Matsudaira P, Lauffenburger DA. Computational model for cell migration in three-dimensional matrices. Biophys J. 2005;89(2):1389-97. doi: 10.1529/biophysj.105.060723. PubMed PMID: 15908579; PMCID: 1366623. 19. Fung J, Lee CK, Chan M, Seto WK, Wong DK, Lai CL, Yuen MF. Defining normal liver stiffness range in a normal healthy Chinese population without liver disease. PLoS One. 2013;8(12):e85067. doi: 10.1371/journal.pone.0085067. PubMed PMID: 24386446; PMCID: 3873442. 20. Mueller S, Sandrin L. Liver stiffness: a novel parameter for the diagnosis of liver disease. Hepatic medicine : evidence and research. 2010;2:49-67. PubMed PMID: 24367208; PMCID: 3846375. 21. Arena U, Vizzutti F, Corti G, Ambu S, Stasi C, Bresci S, Moscarella S, Boddi V, Petrarca A, Laffi G, Marra F, Pinzani M. Acute viral hepatitis increases liver stiffness values measured by transient elastography. Hepatology. 2008;47(2):380-4. doi: 10.1002/hep.22007. PubMed PMID: 18095306. 22. Klatt D, Friedrich C, Korth Y, Vogt R, Braun J, Sack I. Viscoelastic properties of liver measured by oscillatory rheometry and multifrequency magnetic resonance elastography. Biorheology. 2010;47(2):133-41. doi: 10.3233/BIR-2010-0565. PubMed PMID: 20683156. 23. Millonig G, Reimann FM, Friedrich S, Fonouni H, Mehrabi A, Buchler MW, Seitz HK, Mueller S. Extrahepatic cholestasis increases liver stiffness (FibroScan) irrespective of fibrosis. Hepatology. 2008;48(5):1718-23. doi: 10.1002/hep.22577. PubMed PMID: 18836992. 24. Schrader J, Gordon-Walker TT, Aucott RL, van Deemter M, Quaas A, Walsh S, Benten D, Forbes SJ, Wells RG, Iredale JP. Matrix stiffness modulates proliferation, chemotherapeutic response, and dormancy in hepatocellular carcinoma cells. Hepatology. 2011;53(4):1192-205. doi: 10.1002/hep.24108. PubMed PMID: 21442631; PMCID: 3076070. 25. Vinken M, Papeleu P, Snykers S, De Rop E, Henkens T, Chipman JK, Rogiers V, Vanhaecke T. Involvement of cell junctions in hepatocyte culture functionality. Crit Rev Toxicol. 2006;36(4):299-318. doi: 10.1080/10408440600599273. PubMed PMID: 16809101. 26. Lee NP, Luk JM. Hepatic tight junctions: from viral entry to cancer metastasis. World journal of gastroenterology : WJG. 2010;16(3):289-95. PubMed PMID: 20082472; PMCID: 2807947. 27. Balda MS, Matter K. Tight junctions at a glance. J Cell Sci. 2008;121(Pt 22):3677-82. doi: 10.1242/jcs.023887. PubMed PMID: 18987354. 28. Kojima T, Sawada N. Expression and function of claudins in hepatocytes. Methods in molecular biology. 2011;762:233-44. doi: 10.1007/978-1-61779-185-7_16. PubMed PMID: 21717360. 29. Gunzel D, Yu AS. Claudins and the modulation of tight junction permeability. Physiol Rev. 2013;93(2):525-69. doi: 10.1152/physrev.00019.2012. PubMed PMID: 23589827; PMCID: 3768107. 30. Du D, Xu F, Yu L, Zhang C, Lu X, Yuan H, Huang Q, Zhang F, Bao H, Jia L, Wu X, Zhu X, Zhang X, Zhang Z, Chen Z. The tight junction protein, occludin, regulates the directional migration of epithelial cells. Dev Cell. 2010;18(1):52-63. doi: 10.1016/j.devcel.2009.12.008. PubMed PMID: 20152177.
140
31. Rao R. Occludin phosphorylation in regulation of epithelial tight junctions. Ann N Y Acad Sci. 2009;1165:62-8. doi: 10.1111/j.1749-6632.2009.04054.x. PubMed PMID: 19538289. 32. Van Itallie CM, Fanning AS, Holmes J, Anderson JM. Occludin is required for cytokine-induced regulation of tight junction barriers. J Cell Sci. 2010;123(Pt 16):2844-52. doi: 10.1242/jcs.065581. PubMed PMID: 20663912; PMCID: 2915885. 33. Ilic D, Almeida EA, Schlaepfer DD, Dazin P, Aizawa S, Damsky CH. Extracellular matrix survival signals transduced by focal adhesion kinase suppress p53-mediated apoptosis. J Cell Biol. 1998;143(2):547-60. PubMed PMID: 9786962; PMCID: 2132850. 34. Dedhar S, Williams B, Hannigan G. Integrin-linked kinase (ILK): a regulator of integrin and growth-factor signalling. Trends Cell Biol. 1999;9(8):319-23. PubMed PMID: 10407411. 35. Su JM, Wang LY, Liang YL, Zha XL. Role of cell adhesion signal molecules in hepatocellular carcinoma cell apoptosis. World journal of gastroenterology : WJG. 2005;11(30):4667-73. PubMed PMID: 16094707. 36. Persad S, Attwell S, Gray V, Delcommenne M, Troussard A, Sanghera J, Dedhar S. Inhibition of integrin-linked kinase (ILK) suppresses activation of protein kinase B/Akt and induces cell cycle arrest and apoptosis of PTEN-mutant prostate cancer cells. Proc Natl Acad Sci U S A. 2000;97(7):3207-12. doi: 10.1073/pnas.060579697. PubMed PMID: 10716737; PMCID: 16217. 37. Wu C, Dedhar S. Integrin-linked kinase (ILK) and its interactors: a new paradigm for the coupling of extracellular matrix to actin cytoskeleton and signaling complexes. J Cell Biol. 2001;155(4):505-10. doi: 10.1083/jcb.200108077. PubMed PMID: 11696562; PMCID: 2198863. 38. Fabry B, Klemm AH, Kienle S, Schaffer TE, Goldmann WH. Focal adhesion kinase stabilizes the cytoskeleton. Biophys J. 2011;101(9):2131-8. doi: 10.1016/j.bpj.2011.09.043. PubMed PMID: 22067150; PMCID: 3207160. 39. Mitra SK, Hanson DA, Schlaepfer DD. Focal adhesion kinase: in command and control of cell motility. Nat Rev Mol Cell Biol. 2005;6(1):56-68. doi: 10.1038/nrm1549. PubMed PMID: 15688067. 40. Li S, Guan JL, Chien S. Biochemistry and biomechanics of cell motility. Annu Rev Biomed Eng. 2005;7:105-50. doi: 10.1146/annurev.bioeng.7.060804.100340. PubMed PMID: 16004568. 41. Nobes CD, Hall A. Rho, rac, and cdc42 GTPases regulate the assembly of multimolecular focal complexes associated with actin stress fibers, lamellipodia, and filopodia. Cell. 1995;81(1):53-62. PubMed PMID: 7536630. 42. Mayor R, Carmona-Fontaine C. Keeping in touch with contact inhibition of locomotion. Trends Cell Biol. 2010;20(6):319-28. doi: 10.1016/j.tcb.2010.03.005. PubMed PMID: 20399659; PMCID: 2927909. 43. Schneider S, Weydig C, Wessler S. Targeting focal adhesions:Helicobacter pylori-host communication in cell migration. Cell Commun Signal. 2008;6:2. doi: 10.1186/1478-811X-6-2. PubMed PMID: 18684322; PMCID: 2517590. 44. Takai Y, Sasaki T, Matozaki T. Small GTP-binding proteins. Physiol Rev. 2001;81(1):153-208. PubMed PMID: 11152757. 45. Shupe TD, Piscaglia AC, Oh SH, Gasbarrini A, Petersen BE. Isolation and characterization of hepatic stem cells, or "oval cells," from rat livers. Methods in molecular biology. 2009;482:387-405. doi: 10.1007/978-1-59745-060-7_24. PubMed PMID: 19089369; PMCID: 3130598. 46. Lison L. Alcian blue 8 G with chlorantine fast red 5 B.A technic for selective staining of mycopolysaccharides. Stain Technol. 1954;29(3):131-8. PubMed PMID: 13156798. 47. Zheng Shu X, Liu Y, Palumbo FS, Luo Y, Prestwich GD. In situ crosslinkable hyaluronan hydrogels for tissue engineering. Biomaterials. 2004;25(7-8):1339-48. PubMed PMID: 14643608.
141
48. Shu XZ, Ahmad S, Liu Y, Prestwich GD. Synthesis and evaluation of injectable, in situ crosslinkable synthetic extracellular matrices for tissue engineering. J Biomed Mater Res A. 2006;79(4):902-12. doi: 10.1002/jbm.a.30831. PubMed PMID: 16941590. 49. Skardal A, Devarasetty M, Kang HW, Mead I, Bishop C, Shupe T, Lee SJ, Jackson J, Yoo J, Soker S, Atala A. A hydrogel bioink toolkit for mimicking native tissue biochemical and mechanical properties in bioprinted tissue constructs. Acta Biomater. 2015. doi: 10.1016/j.actbio.2015.07.030. PubMed PMID: 26210285. 50. Skardal A, Zhang J, McCoard L, Oottamasathien S, Prestwich GD. Dynamically crosslinked gold nanoparticle - hyaluronan hydrogels. Adv Mater. 2010;22(42):4736-40. Epub 2010/08/24. doi: 10.1002/adma.201001436. PubMed PMID: 20730818. 51. Skardal A, Zhang J, McCoard L, Xu X, Oottamasathien S, Prestwich GD. Photocrosslinkable hyaluronan-gelatin hydrogels for two-step bioprinting. Tissue Eng Part A. 2010;16(8):2675-85. doi: 10.1089/ten.TEA.2009.0798. PubMed PMID: 20387987; PMCID: 3135254. 52. Skardal A, Zhang J, Prestwich GD. Bioprinting vessel-like constructs using hyaluronan hydrogels crosslinked with tetrahedral polyethylene glycol tetracrylates. Biomaterials. 2010;31(24):6173-81. Epub 2010/06/16. doi: S0142-9612(10)00561-2 [pii]
10.1016/j.biomaterials.2010.04.045. PubMed PMID: 20546891. 53. Meyers MA, Chawla KK. Mechanical behavior of materials. Upper Saddle River, N.J.: Prentice Hall; 1999. xv, 680 p. p. 54. Swift B, Brouwer KL. Influence of seeding density and extracellular matrix on bile Acid transport and mrp4 expression in sandwich-cultured mouse hepatocytes. Mol Pharm. 2010;7(2):491-500. doi: 10.1021/mp900227a. PubMed PMID: 19968322; PMCID: 3235796. 55. Khetani SR, Bhatia SN. Microscale culture of human liver cells for drug development. Nat Biotechnol. 2008;26(1):120-6. doi: 10.1038/nbt1361. PubMed PMID: 18026090. 56. D'Costa ZJ, Jolly C, Androphy EJ, Mercer A, Matthews CM, Hibma MH. Transcriptional repression of E-cadherin by human papillomavirus type 16 E6. PLoS One. 2012;7(11):e48954. doi: 10.1371/journal.pone.0048954. PubMed PMID: 23189137; PMCID: 3506579. 57. Dingemans AM, van den Boogaart V, Vosse BA, van Suylen RJ, Griffioen AW, Thijssen VL. Integrin expression profiling identifies integrin alpha5 and beta1 as prognostic factors in early stage non-small cell lung cancer. Molecular cancer. 2010;9:152. doi: 10.1186/1476-4598-9-152. PubMed PMID: 20565758; PMCID: 2895598. 58. Pol A, Calvo M, Lu A, Enrich C. The "early-sorting" endocytic compartment of rat hepatocytes is involved in the intracellular pathway of caveolin-1 (VIP-21). Hepatology. 1999;29(6):1848-57. doi: 10.1002/hep.510290602. PubMed PMID: 10347129. 59. Wall DA, Hubbard AL. Receptor-mediated endocytosis of asialoglycoproteins by rat liver hepatocytes: biochemical characterization of the endosomal compartments. J Cell Biol. 1985;101(6):2104-12. PubMed PMID: 2866191; PMCID: 2114009. 60. Cao H, Krueger EW, McNiven MA. Hepatocytes internalize trophic receptors at large endocytic "Hot Spots". Hepatology. 2011;54(5):1819-29. doi: 10.1002/hep.24572. PubMed PMID: 21793030; PMCID: 3205295. 61. Treyer A, Musch A. Hepatocyte polarity. Compr Physiol. 2013;3(1):243-87. doi: 10.1002/cphy.c120009. PubMed PMID: 23720287; PMCID: 3697931. 62. Theard D, Steiner M, Kalicharan D, Hoekstra D, van Ijzendoorn SC. Cell polarity development and protein trafficking in hepatocytes lacking E-cadherin/beta-catenin-based adherens junctions. Mol Biol Cell. 2007;18(6):2313-21. doi: 10.1091/mbc.E06-11-1040. PubMed PMID: 17429067; PMCID: 1877101.
142
63. Giepmans BN, van Ijzendoorn SC. Epithelial cell-cell junctions and plasma membrane domains. Biochim Biophys Acta. 2009;1788(4):820-31. doi: 10.1016/j.bbamem.2008.07.015. PubMed PMID: 18706883. 64. Miyoshi A, Kitajima Y, Sumi K, Sato K, Hagiwara A, Koga Y, Miyazaki K. Snail and SIP1 increase cancer invasion by upregulating MMP family in hepatocellular carcinoma cells. British journal of cancer. 2004;90(6):1265-73. doi: 10.1038/sj.bjc.6601685. PubMed PMID: 15026811; PMCID: 2409652. 65. Brieva TA, Moghe PV. Functional engineering of hepatocytes via heterocellular presentation of a homoadhesive molecule, E-cadherin. Biotechnol Bioeng. 2001;76(4):295-302. PubMed PMID: 11745156. 66. Luebke-Wheeler JL, Nedredal G, Yee L, Amiot BP, Nyberg SL. E-cadherin protects primary hepatocyte spheroids from cell death by a caspase-independent mechanism. Cell transplantation. 2009;18(12):1281-7. doi: 10.3727/096368909X474258. PubMed PMID: 20003757; PMCID: 2920600. 67. Ruoslahti E. RGD and other recognition sequences for integrins. Annual review of cell and developmental biology. 1996;12:697-715. doi: 10.1146/annurev.cellbio.12.1.697. PubMed PMID: 8970741. 68. Pinkse GG, Jiawan-Lalai R, Bruijn JA, de Heer E. RGD peptides confer survival to hepatocytes via the beta1-integrin-ILK-pAkt pathway. J Hepatol. 2005;42(1):87-93. doi: 10.1016/j.jhep.2004.09.010. PubMed PMID: 15629512. 69. You J, Park SA, Shin DS, Patel D, Raghunathan VK, Kim M, Murphy CJ, Tae G, Revzin A. Characterizing the effects of heparin gel stiffness on function of primary hepatocytes. Tissue Eng Part A. 2013;19(23-24):2655-63. doi: 10.1089/ten.TEA.2012.0681. PubMed PMID: 23815179; PMCID: 3856597. 70. Kim M, Lee JY, Jones CN, Revzin A, Tae G. Heparin-based hydrogel as a matrix for encapsulation and cultivation of primary hepatocytes. Biomaterials. 2010;31(13):3596-603. doi: 10.1016/j.biomaterials.2010.01.068. PubMed PMID: 20153045; PMCID: 2837121. 71. Foucher J, Chanteloup E, Vergniol J, Castera L, Le Bail B, Adhoute X, Bertet J, Couzigou P, de Ledinghen V. Diagnosis of cirrhosis by transient elastography (FibroScan): a prospective study. Gut. 2006;55(3):403-8. doi: 10.1136/gut.2005.069153. PubMed PMID: 16020491; PMCID: 1856085. 72. Wong VW, Vergniol J, Wong GL, Foucher J, Chan HL, Le Bail B, Choi PC, Kowo M, Chan AW, Merrouche W, Sung JJ, de Ledinghen V. Diagnosis of fibrosis and cirrhosis using liver stiffness measurement in nonalcoholic fatty liver disease. Hepatology. 2010;51(2):454-62. doi: 10.1002/hep.23312. PubMed PMID: 20101745. 73. Ganne-Carrie N, Ziol M, de Ledinghen V, Douvin C, Marcellin P, Castera L, Dhumeaux D, Trinchet JC, Beaugrand M. Accuracy of liver stiffness measurement for the diagnosis of cirrhosis in patients with chronic liver diseases. Hepatology. 2006;44(6):1511-7. doi: 10.1002/hep.21420. PubMed PMID: 17133503. 74. Evans DW, Moran EC, Baptista PM, Soker S, Sparks JL. Scale-dependent mechanical properties of native and decellularized liver tissue. Biomech Model Mechanobiol. 2013;12(3):569-80. doi: 10.1007/s10237-012-0426-3. PubMed PMID: 22890366. 75. Solon J, Levental I, Sengupta K, Georges PC, Janmey PA. Fibroblast adaptation and stiffness matching to soft elastic substrates. Biophys J. 2007;93(12):4453-61. doi: 10.1529/biophysj.106.101386. PubMed PMID: 18045965; PMCID: 2098710. 76. Lo CM, Wang HB, Dembo M, Wang YL. Cell movement is guided by the rigidity of the substrate. Biophysical Journal. 2000;79(1):144-52. PubMed PMID: WOS:000088048500011.
143
77. Discher DE, Janmey P, Wang YL. Tissue cells feel and respond to the stiffness of their substrate. Science. 2005;310(5751):1139-43. doi: 10.1126/science.1116995. PubMed PMID: 16293750. 78. Engler AJ, Sweeney HL, Discher DE, Schwarzbauer JE. Extracellular matrix elasticity directs stem cell differentiation. J Musculoskelet Neuronal Interact. 2007;7(4):335. PubMed PMID: 18094500. 79. Engler AJ, Sen S, Sweeney HL, Discher DE. Matrix elasticity directs stem cell lineage specification. Cell. 2006;126(4):677-89. doi: 10.1016/j.cell.2006.06.044. PubMed PMID: 16923388. 80. Yeung T, Georges PC, Flanagan LA, Marg B, Ortiz M, Funaki M, Zahir N, Ming W, Weaver V, Janmey PA. Effects of substrate stiffness on cell morphology, cytoskeletal structure, and adhesion. Cell Motil Cytoskeleton. 2005;60(1):24-34. doi: 10.1002/cm.20041. PubMed PMID: 15573414. 81. Kumar S, Maxwell IZ, Heisterkamp A, Polte TR, Lele TP, Salanga M, Mazur E, Ingber DE. Viscoelastic retraction of single living stress fibers and its impact on cell shape, cytoskeletal organization, and extracellular matrix mechanics. Biophys J. 2006;90(10):3762-73. doi: 10.1529/biophysj.105.071506. PubMed PMID: 16500961; PMCID: 1440757. 82. Martin TA, Jiang WG. Loss of tight junction barrier function and its role in cancer metastasis. Biochim Biophys Acta. 2009;1788(4):872-91. doi: 10.1016/j.bbamem.2008.11.005. PubMed PMID: 19059202. 83. Maly IP, Landmann L. Bile duct ligation in the rat causes upregulation of ZO-2 and decreased colocalization of claudins with ZO-1 and occludin. Histochemistry and cell biology. 2008;129(3):289-99. doi: 10.1007/s00418-007-0374-7. PubMed PMID: 18197414. 84. Kojima T, Takano K, Yamamoto T, Murata M, Son S, Imamura M, Yamaguchi H, Osanai M, Chiba H, Himi T, Sawada N. Transforming growth factor-beta induces epithelial to mesenchymal transition by down-regulation of claudin-1 expression and the fence function in adult rat hepatocytes. Liver Int. 2008;28(4):534-45. doi: 10.1111/j.1478-3231.2007.01631.x. PubMed PMID: 18031476. 85. Osanai M, Nishikiori N, Murata M, Chiba H, Kojima T, Sawada N. Occludin expression inhibits tumorigenicity and metastasis. Faseb J. 2006;20(4):A223-A. PubMed PMID: WOS:000236206501489. 86. Hadj-Rabia S, Baala L, Vabres P, Hamel-Teillac D, Jacquemin E, Fabre M, Lyonnet S, De Prost Y, Munnich A, Hadchouel M, Smahi A. Claudin-1 gene mutations in neonatal sclerosing cholangitis associated with ichthyosis: a tight junction disease. Gastroenterology. 2004;127(5):1386-90. PubMed PMID: 15521008. 87. Grosse B, Cassio D, Yousef N, Bernardo C, Jacquemin E, Gonzales E. Claudin-1 involved in neonatal ichthyosis sclerosing cholangitis syndrome regulates hepatic paracellular permeability. Hepatology. 2012;55(4):1249-59. doi: Doi 10.1002/Hep.24761. PubMed PMID: WOS:000302069900028. 88. Ikenouchi J, Matsuda M, Furuse M, Tsukita S. Regulation of tight junctions during the epithelium-mesenchyme transition: direct repression of the gene expression of claudins/occludin by Snail. Journal of Cell Science. 2003;116(10):1959-67. doi: DOI 10.1242/jcs.00389. PubMed PMID: WOS:000183098900011. 89. Ando-Akatsuka Y, Yonemura S, Itoh M, Furuse M, Tsukita S. Differential behavior of E-cadherin and occludin in their colocalization with ZO-1 during the establishment of epithelial cell polarity. J Cell Physiol. 1999;179(2):115-25. doi: 10.1002/(SICI)1097-4652(199905)179:2<115::AID-JCP1>3.0.CO;2-T. PubMed PMID: 10199550.
144
90. Dorfel MJ, Huber O. Modulation of tight junction structure and function by kinases and phosphatases targeting occludin. Journal of biomedicine & biotechnology. 2012;2012:807356. doi: 10.1155/2012/807356. PubMed PMID: 22315516; PMCID: 3270569. 91. Hehlgans S, Haase M, Cordes N. Signalling via integrins: implications for cell survival and anticancer strategies. Biochim Biophys Acta. 2007;1775(1):163-80. doi: 10.1016/j.bbcan.2006.09.001. PubMed PMID: 17084981. 92. Horowitz JC, Rogers DS, Sharma V, Vittal R, White ES, Cui Z, Thannickal VJ. Combinatorial activation of FAK and AKT by transforming growth factor-beta1 confers an anoikis-resistant phenotype to myofibroblasts. Cell Signal. 2007;19(4):761-71. doi: 10.1016/j.cellsig.2006.10.001. PubMed PMID: 17113264; PMCID: 1820832. 93. Frisch SM, Schaller M, Cieply B. Mechanisms that link the oncogenic epithelial-mesenchymal transition to suppression of anoikis. J Cell Sci. 2013;126(Pt 1):21-9. doi: 10.1242/jcs.120907. PubMed PMID: 23516327; PMCID: 3603508. 94. Hsia DA, Mitra SK, Hauck CR, Streblow DN, Nelson JA, Ilic D, Huang S, Li E, Nemerow GR, Leng J, Spencer KS, Cheresh DA, Schlaepfer DD. Differential regulation of cell motility and invasion by FAK. J Cell Biol. 2003;160(5):753-67. doi: 10.1083/jcb.200212114. PubMed PMID: 12615911; PMCID: 2173366. 95. Hildebrand JD, Taylor JM, Parsons JT. An SH3 domain-containing GTPase-activating protein for Rho and Cdc42 associates with focal adhesion kinase. Mol Cell Biol. 1996;16(6):3169-78. PubMed PMID: 8649427; PMCID: 231310. 96. Tilghman RW, Slack-Davis JK, Sergina N, Martin KH, Iwanicki M, Hershey ED, Beggs HE, Reichardt LF, Parsons JT. Focal adhesion kinase is required for the spatial organization of the leading edge in migrating cells. J Cell Sci. 2005;118(Pt 12):2613-23. doi: 10.1242/jcs.02380. PubMed PMID: 15914540. 97. Ren XD, Kiosses WB, Sieg DJ, Otey CA, Schlaepfer DD, Schwartz MA. Focal adhesion kinase suppresses Rho activity to promote focal adhesion turnover. J Cell Sci. 2000;113 ( Pt 20):3673-8. PubMed PMID: 11017882. 98. Weber GF, Bjerke MA, DeSimone DW. Integrins and cadherins join forces to form adhesive networks. J Cell Sci. 2011;124(Pt 8):1183-93. doi: 10.1242/jcs.064618. PubMed PMID: 21444749; PMCID: 3115772. 99. Ingber DE. Cellular mechanotransduction: putting all the pieces together again. Faseb J. 2006;20(7):811-27. doi: 10.1096/fj.05-5424rev. PubMed PMID: 16675838. 100. Ogunwobi OO, Liu C. Hepatocyte growth factor upregulation promotes carcinogenesis and epithelial-mesenchymal transition in hepatocellular carcinoma via Akt and COX-2 pathways. Clinical & experimental metastasis. 2011;28(8):721-31. doi: 10.1007/s10585-011-9404-x. PubMed PMID: 21744257; PMCID: 3732749. 101. Nagai T, Arao T, Furuta K, Sakai K, Kudo K, Kaneda H, Tamura D, Aomatsu K, Kimura H, Fujita Y, Matsumoto K, Saijo N, Kudo M, Nishio K. Sorafenib inhibits the hepatocyte growth factor-mediated epithelial mesenchymal transition in hepatocellular carcinoma. Molecular cancer therapeutics. 2011;10(1):169-77. doi: 10.1158/1535-7163.MCT-10-0544. PubMed PMID: 21220499. 102. Mangold S, Wu SK, Norwood SJ, Collins BM, Hamilton NA, Thorn P, Yap AS. Hepatocyte growth factor acutely perturbs actin filament anchorage at the epithelial zonula adherens. Current biology : CB. 2011;21(6):503-7. doi: 10.1016/j.cub.2011.02.018. PubMed PMID: 21396819. 103. Jones CN, Tuleuova N, Lee JY, Ramanculov E, Reddi AH, Zern MA, Revzin A. Cultivating hepatocytes on printed arrays of HGF and BMP7 to characterize protective effects of these
145
growth factors during in vitro alcohol injury. Biomaterials. 2010;31(23):5936-44. doi: 10.1016/j.biomaterials.2010.04.006. PubMed PMID: 20488537; PMCID: 2893270. 104. Liu Y. Hepatocyte growth factor in kidney fibrosis: therapeutic potential and mechanisms of action. American journal of physiology Renal physiology. 2004;287(1):F7-16. doi: 10.1152/ajprenal.00451.2003. PubMed PMID: 15180923. 105. Yang J, Dai C, Liu Y. A novel mechanism by which hepatocyte growth factor blocks tubular epithelial to mesenchymal transition. Journal of the American Society of Nephrology : JASN. 2005;16(1):68-78. doi: 10.1681/ASN.2003090795. PubMed PMID: 15537870. 106. Monga SP, Mars WM, Pediaditakis P, Bell A, Mule K, Bowen WC, Wang X, Zarnegar R, Michalopoulos GK. Hepatocyte growth factor induces Wnt-independent nuclear translocation of beta-catenin after Met-beta-catenin dissociation in hepatocytes. Cancer Res. 2002;62(7):2064-71. PubMed PMID: 11929826. 107. Balkovetz DF, Pollack AL, Mostov KE. Hepatocyte growth factor alters the polarity of Madin-Darby canine kidney cell monolayers. J Biol Chem. 1997;272(6):3471-7. PubMed PMID: 9013593. 108. Apte U, Zeng G, Muller P, Tan X, Micsenyi A, Cieply B, Dai C, Liu Y, Kaestner KH, Monga SP. Activation of Wnt/beta-catenin pathway during hepatocyte growth factor-induced hepatomegaly in mice. Hepatology. 2006;44(4):992-1002. doi: 10.1002/hep.21317. PubMed PMID: 17006939.
147
The field of regenerative medicine uses a variety of tools, technologies, and
advanced models to gain knowledge of disease mechanisms and to develop new therapies
and treatments. Each approach has a set of advantages and disadvantages depending on
the field of research. Three-dimensional liver spheroids, or organoids, recreate entire
organ systems on a miniature scale (1-5). Spheroids can be maintained in a variety of
substrates like hydrogels. In microfluidic devices, organoids are arranged in a circuit and
interact in a way that mimics how organ systems work together (6, 7). Consequently, they
are a great platform for toxicology and drug testing. Because of their longevity and long
term maintenance of function, testing can be carried out in vitro without the need for
complex in vivo models. However, organoids are not always suitable for ECM based
studies. These cell structures form tight junctions which prevent manipulation of ECM
composition and mechanical properties at the core of the organoids. Since cells only
sense ECM composition and mechanical forces in adjacent areas, these cell centers would
be relatively unaffected by substrate alterations.
Decellularized tissue is an excellent biomaterial for tissue engineering. Optimized
decellularization methods preserve ECM components in their native architecture and
maintain physiological proportions in the tissue. This scaffold material enables cells to
home to specific sites of the matrix and either sustain function or, in the cases of stem
cells, differentiate into appropriate cell phenotypes (8-10). However, challenges remain
in producing consistent decellularized products. The success of cell removal or the
remaining composition of the decellularized tissue can vary because of differences
between animals and differences in the size or vascularity of the organ. When designing
studies on small tissue segments, it can be hard to control for structural variances in the
148
tissue that may exist even within the same organ. Matrix samples used for cellular assays
that are excised from different regions of the organ, certain areas that are not fully
decellularized, or sections damaged by the perfusion process can create error or deviation
in the results. It is also difficult to precisely manipulate physical and chemical properties
of the intact matrix. Without a complete analysis and mapping of the tissue, methods to
initiate these changes could be unpredictable.
Hydrogel substrates are viable alternatives to intact decellularized tissue
constructs. Although no longer possessing the native microarchitecture, gels enable better
control and manipulation of individual substrate properties. The concentration of specific
molecules in the hydrogels can be controlled and altered. In the case of this thesis study,
ECM from decellularized tissue was solubilized and incorporated into HA gel substrates
to replicate a natural liver microenvironment. Since the matrix of the entire organs was
solubilized in a homogenous solution, there was little variability within the material. All
regions of the ECM and multiple liver samples were combined, stored, and utilized
across multiple assays, allowing for consistent composition of the substrates. ECM
molecules were also quantified to ensure that all assays used the same total concentration
of solubilized material, and there were similar ratios of structural and bioregulatory
molecules. This ability standardized substrate production and resulted in more consistent
results and less experimental error.
This thesis work demonstrates that HA gels supplemented with ECM provided
some of the same functional advantages for seeded cells as fully intact matrix scaffolds.
Use of ECM containing gels in past studies were shown to greatly improve long term
viability and function of culture of cells, as compared to traditional in vitro methods (11,
149
12). Our human hepatocyte assays showed increases in viability, adhesion, and cell
junction formation. To determine the mechanism for some of the improvements seen in
primary hepatocyte attachment and viability, expression of integrin β-1 was analyzed in
cells seeded on ECM supplemented gels versus cells on HA only gels. At day 7,
hepatocyte expression increased on gel substrates with the addition of liver matrix.
Although only one integrin class was measured, integrin β-1 may be the most
critical type for maintenance of cell function. Integrin β-1 is the most highly expressed
beta-integrin, dimerizing with more than 10 alpha-integrin subunits (13). These integrin
complexes directly interact with the actin cytoskeleton (14). Integrin β-1 expression
likely correlates with overall integrin expression in the presence of liver ECM and
explains the increases in hepatocyte attachment and viability relative to cells seeded on
simpler substrates. To establish this conclusion, expression of a greater variety of integrin
types could be analyzed. Varying concentrations of solubilized liver matrix could also be
used in substrate formation to determine if concentration dependent changes were
observed. As controls, certain integrins could also be blocked with increasing
concentrations of integrin-specific antibodies or RGD peptides in order to measure
reduction in binding or survival of the hepatocytes. Manipulations of the cells and the
adaptable gel substrate could help determine the most likely mechanisms for differences
observed.
Greater control of the composition of the substrate also correlates with better
control of the chemistry and physical properties of the material. These characteristics can
be adjusted by adding chemically-modified compounds to the substrate preparation.
Thiol-modified HA, heparin, and collagen were used for gel formation in our study. PEG-
150
acrylate crosslinker reacted with these moieties to solidify hydrogels and alter stiffness
within a narrow range. Uniform composition of our ECM supplemented gels allowed
creation of sample replicates at equivalent stiffness levels. Different concentrations and
configurations of the crosslinker also generated adjustable degrees of stiffness to
establish separate experimental groups. Crosslinking can vary the property of stiffness
without modifying the concentration of functional elements in the ECM substrates. These
advantages make ECM supplemented gels optimal for use in cell culture assays to
identify the effects of substrate stiffness variations.
Following the development and creation of liver ECM supplemented gels, this
thesis evaluated the effects of substrate stiffness on primary hepatocyte function. Results
revealed hepatocyte gene marker expression and the functional output of albumin varied
with stiffness and time in culture. In early time points, higher gel stiffness stimulated
greater metabolic activity and albumin output. However, after metabolic functions
stabilized for 7 days, hepatocytes grown on an intermediate stiffness of 1200 Pa produced
albumin at the greatest rate. This observation was correlated to a significant drop off in
gene expression of hepatic markers by cells on the highest stiffness of 4600 Pa. To test
the hypothesis that the highest 4600 Pa stiffness causes a loss in long term phenotype,
experiments could utilize time points past day 7. These assays would indicate if observed
variances in function and gene expression widen between cells seeded on the different
stiffnesses in the long term. It would also be relevant to examine if markers for EMT like
SNAIL were upregulated by cells on the highest stiffness group.
The balance between EMT and MET is vital in liver development and could
possibly have an integral role in liver repair and regeneration. It is theorized that
151
following injury, epithelial cells undergo EMT and travel to the mesenchyme to
proliferate and produce ECM molecules. These cells then differentiate into hepatocytes or
cholangiocytes to repopulate and restore the liver parenchyma (15). Imbalances in this
process can lead to increases in fibroblasts and a progression of disease (16). During liver
disease, many experts theorize that the stiffening ECM microenvironment promotes
additional EMT of hepatocytes and contributes to worsening fibrosis and eventual organ
failure (16). Our work has helped delineate a proper substrate stiffness for long term
maintenance of hepatocyte phenotype as well as a stiffness range where EMT could be
initiated. Future experiments could be designed using the liver ECM gels to further
replicate events of liver repair or liver disease in culture and study the delicate balance
between EMT and MET.
While studying the effects of stiffness on hepatocyte phenotype, assays were
designed to study the mechanisms behind observed changes in attachment, viability, and
morphology. Experiments monitored gene and protein expression of FAK and ILK, two
integrin-localized kinases with vital roles in mechanotransduction and cytoskeletal
remodeling. As previously discussed, hepatocytes on all gel stiffness expressed the FAK
and ILK genes at levels higher than cells of normal liver tissue. However, at day 7 post-
seeding, FAK and ILK gene and protein expression levels were highest in the lower
stiffness levels, with FAK concentrated at the leading edge of epithelial cell layers.
Although FAK and ILK have been shown to have effects on viability, trends we observed
seemed to correlate most closely with morphological changes in the hepatocytes,
specifically with formation of cellular projections.
152
Previous studies have shown FAK and ILK expression stimulates pathways
controlling cytoskeletal structures. In addition, overexpression of these kinases has been
observed in migrating cells. With this knowledge, expression of the downstream targets
and actin regulating Rho GTPases were measured to determine if differences in
cytoskeleton activation and regulation were occurring in human hepatocytes seeded on
ECM gels of varying stiffnesses. Cells expressed RhoA, which controls stress fiber
formation and contractility, on stiffer ECM gels and exhibited greater attachment and
assembly of actin fibers. CDC42, which initiates polarization and filopodia formation,
decreased with substrate stiffness. This result demonstrated how FAK, ILK, and CDC42
overexpression correlates with lower substrate stiffness, morphological changes, and
most likely, cell migration.
Further experimentation could help confirm detected morphological and
migratory trends and help strengthen our hypotheses. Determining exact mechanisms and
sites of FAK and ILK phosphorylation induced by substrate stiffness could help explain
the numerous functions of these kinases. Previous studies have shown blocking FAK and
ILK in cell culture assays may not be ideal because deficiencies in FAK and ILK lead to
reductions in viability. However, plasmid transfections could be used to upregulate FAK,
ILK, or CDC42 in primary hepatocytes, as has been successful in previous studies (17).
Site specific mutations could also be used to pinpoint the importance of specific
phosphorylation sites. RhoA could also be upregulated to determine if increased
expression initiates development of a more anchored phenotype with greater actin
expression. Induction of morphological effects similar to what was observed on the
varying gel stiffnesses would support our hypotheses and further implicate FAK, ILK,
153
and the Rho GTPases in the changes we observed. Hepatocyte culture experiments and
imaging specifically designed to track and map cell motility and migration throughout
time points past 7 days would also more definitively validate our theories.
In addition to analyzing the expression and localization of FAK, ILK, and the
Rho GTPases, additional targets related to mechanotransduction could be included. Src
tyrosine kinase binds to and phosphorylates various FAK sites. Several adaptor proteins
including PI3K, CAS, GRB7, and N-WASP associate with FAK and act as intermediates
between FAK and the Rho GTPases. Similar to FAK, ILK also stimulates the PI3K
pathway. Unique adaptor proteins of ILK include PINCH1 and ELMO2 which aid in
cellular polarization. Other focal adhesion components important to mechanosensing,
integrin attachment, and actin modification include talin, vinculin, and paxillin. Assays
testing expression and localization of all these proteins would provide a clearer
understanding of the interactions between ECM gel stiffness, substrate sensing, and the
regulation of the hepatocyte cytoskeleton and focal adhesion formation.
ECM composition and substrate stiffness play important roles in hepatocyte
phenotype; however, growth factor presence also represents a factor in determining the
fates of cells. Thesis studies have shown the ability of ECM gel substrates to bind active
growth factor for cellular use (Chapter 3- Figure 2). Future assays like phospho-labeling
will be required to pinpoint exact binding and interaction sites in the gels. Nevertheless,
the presented studies demonstrate the ability to manipulate the growth factor
concentration in the liver cell substrates. With these ECM supplemented gels, future
assays could be designed to measure how stiffness intensifies or reduces growth factor
effects on liver cells.
154
Preliminary studies exhibited the ability of HGF to bind to our liver hydrogels and
improve cell junction formation and hepatocyte function on low stiffness gels (Chapter 3-
Supplementary Figure 4). A previous publication showed similar results with HGF and
EGF-treated hepatocytes seeded on Matrigel of varying stiffnesses. Growth factor co-
stimulation improved aggregation and function on the lower stiffness, while the converse
occurred at the higher stiffness level (18). Additional experiments on HGF and EGF
stimulated epithelial cells implicated stiffness in cell polarization and migratory
responses (19-22). Other research has also shown the direct effects of substrate stiffness
on transforming growth factor beta (TGF-β)-induced dedifferentiation of hepatocytes and
possible EMT (23). Studies could be conducted to determine if these same changes occur
on our liver ECM gels. Further data could define the exact stiffness ranges that induce
these responses and the molecular mechanisms behind these behaviors.
Opportunities exist to improve both the complexity and overall function of the
hepatocyte cell culture system we have developed. Other cell types could be individually
seeded on the ECM supplemented substrates to test effects of stiffness on different cells
of the liver, or co-culture of multiple cell types could determine how they interact and
function in varied mechanical environments. Hepatic stellate cells (HSCs) are one of
most important liver cell types involved in ECM production and maintenance. HSCs exist
in either a quiescent or myofibroblast form. In vivo these cells are vital to both hepatocyte
and stem cell mediated regeneration and repair. They secrete and assemble ECM
molecules including fibronectin, collagens, and GAGs (24, 25). During a fibrotic disease
state, hepatic stellate cells deposit excess amounts of collagen, a process worsened by
myofibroblastic activation and proliferation on stiffening ECM. Several fields from
155
cancer biology to regenerative medicine also explored their ability to secrete growth
factors. Assays revealed HSCs produce HGF, EGF, TGFα, and multiple insulin-like
growth factor binding proteins within the liver (26-28).
Previous studies have shown the abilities of stiff substrates to stimulate
myofibroblastic differentiation of HSCs (29, 30). Less is known about the function of
quiescent cells in the normal liver state and the exact stiffness range where
myofibroblastic activation occurs. Results on the liver ECM gels determined that hepatic
stellate cell production of GAGs and HGF were time and stiffness dependent. HGF
production was highest in the lowest gel stiffness, while ECM production was greatest on
the highest gel stiffness (Chapter 3- Supplementary Figure 5). More detailed studies
would reveal if these cells could substantially modify the composition or physical
properties of the substrate microenvironment. Co-culture with primary hepatocytes
should improve long term viability of the cells and provide insight into the relationships
between cell types and certain substrate properties. In addition, modeling a fibrotic
stiffness and determining methods of controlling excess ECM production and
maintenance of hepatocyte function could help in the understanding and treatment of
liver diseases like cirrhosis.
Besides increasing diversity of cell types grown on the substrate, increasing the
three dimensionality of this system could improve capabilities to model disease states and
enable advances in tissue engineering. Hepatocytes are arranged in epithelial sheets in the
liver. 2D systems replicate individual cell layers, but not the complex interactions of the
entire tissue. New 3D bioprinting technology could provide a method of creating these
3D structures. This tool enables careful layering of cells and gel substrate. Using
156
knowledge gained from this thesis, a bioprinter could alternate printing cell sheets and
ECM gel layers at a set stiffness level to produce a tissue-like system. In this way, in
vitro modeling could better mimic an in vivo environment, while still providing an
adjustable platform to study specific ECM properties and mechanisms.
Despite the advantages of a fully 3D gel system, new challenges and
complications are created that are non-existent in 2D culture. Gel swelling, pore size, and
the flow of cell nutrients through the substrate all become important. Crosslinking affects
stiffness on a gel layer, but could have confounding effects in 3D. Substrate fabrication
and multiple gel properties would need to be monitored, tested, and studied to prevent the
introduction of unwanted variables. Although several areas still need to be addressed, the
abilities and flexibility of ECM gel substrates provide a promising future.
Overall, liver ECM gels improve long term cell function and allow isolation and
testing of single substrate-related properties. This thesis work elucidated specific
mechanisms of mechanotransduction and cytoskeletal regulation and determined stiffness
ranges where hepatocyte function was optimized in vitro. Future advances and
development of this substrate system could further clarify molecular interactions and
produce more accurate disease models. Liver ECM gels could also be utilized in cell
encapsulation technologies or cell therapies to repair liver injuries or disorders. The thesis
established methods to integrate a natural liver ECM microenvironment in gel substrates
at a physiological stiffness range. This new knowledge and innovation has facilitated
numerous possibilities to pursue and can ultimately lead to future advances and practical
therapies or treatments.
157
Works Cited
1. Sato T, Stange DE, Ferrante M, Vries RG, Van Es JH, Van den Brink S, Van Houdt WJ, Pronk A, Van Gorp J, Siersema PD, Clevers H. Long-term expansion of epithelial organoids from human colon, adenoma, adenocarcinoma, and Barrett's epithelium. Gastroenterology. 2011;141(5):1762-72. doi: 10.1053/j.gastro.2011.07.050. PubMed PMID: 21889923. 2. Nguyen-Ngoc KV, Cheung KJ, Brenot A, Shamir ER, Gray RS, Hines WC, Yaswen P, Werb Z, Ewald AJ. ECM microenvironment regulates collective migration and local dissemination in normal and malignant mammary epithelium. Proc Natl Acad Sci U S A. 2012;109(39):E2595-604. doi: 10.1073/pnas.1212834109. PubMed PMID: 22923691; PMCID: 3465416. 3. Mroue R, Bissell MJ. Three-dimensional cultures of mouse mammary epithelial cells. Methods in molecular biology. 2013;945:221-50. doi: 10.1007/978-1-62703-125-7_14. PubMed PMID: 23097110; PMCID: 3666564. 4. Alcaraz J, Xu R, Mori H, Nelson CM, Mroue R, Spencer VA, Brownfield D, Radisky DC, Bustamante C, Bissell MJ. Laminin and biomimetic extracellular elasticity enhance functional differentiation in mammary epithelia. EMBO J. 2008;27(21):2829-38. doi: 10.1038/emboj.2008.206. PubMed PMID: 18843297; PMCID: 2569873. 5. Whitehead RH, Robinson PS, Williams JA, Bie W, Tyner AL, Franklin JL. Conditionally immortalized colonic epithelial cell line from a Ptk6 null mouse that polarizes and differentiates in vitro. J Gastroenterol Hepatol. 2008;23(7 Pt 1):1119-24. doi: 10.1111/j.1440-1746.2008.05308.x. PubMed PMID: 18205771; PMCID: 3005200. 6. Bhatia SN, Ingber DE. Microfluidic organs-on-chips. Nat Biotechnol. 2014;32(8):760-72. doi: 10.1038/nbt.2989. PubMed PMID: 25093883. 7. Khademhosseini A, Eng G, Yeh J, Kucharczyk PA, Langer R, Vunjak-Novakovic G, Radisic M. Microfluidic patterning for fabrication of contractile cardiac organoids. Biomed Microdevices. 2007;9(2):149-57. doi: 10.1007/s10544-006-9013-7. PubMed PMID: 17146728. 8. Park JS, Chu JS, Tsou AD, Diop R, Tang Z, Wang A, Li S. The effect of matrix stiffness on the differentiation of mesenchymal stem cells in response to TGF-beta. Biomaterials. 2011;32(16):3921-30. doi: 10.1016/j.biomaterials.2011.02.019. PubMed PMID: 21397942; PMCID: 3073995. 9. Gattazzo F, Urciuolo A, Bonaldo P. Extracellular matrix: a dynamic microenvironment for stem cell niche. Biochim Biophys Acta. 2014;1840(8):2506-19. doi: 10.1016/j.bbagen.2014.01.010. PubMed PMID: 24418517; PMCID: 4081568. 10. Malinen MM, Kanninen LK, Corlu A, Isoniemi HM, Lou YR, Yliperttula ML, Urtti AO. Differentiation of liver progenitor cell line to functional organotypic cultures in 3D nanofibrillar cellulose and hyaluronan-gelatin hydrogels. Biomaterials. 2014;35(19):5110-21. doi: 10.1016/j.biomaterials.2014.03.020. PubMed PMID: 24698520. 11. Skardal A, Smith L, Bharadwaj S, Atala A, Soker S, Zhang Y. Tissue specific synthetic ECM hydrogels for 3-D in vitro maintenance of hepatocyte function. Biomaterials. 2012;33(18):4565-75. doi: 10.1016/j.biomaterials.2012.03.034. PubMed PMID: 22475531; PMCID: 3719050. 12. Nakamura S, Ijima H. Solubilized matrix derived from decellularized liver as a growth factor-immobilizable scaffold for hepatocyte culture. J Biosci Bioeng. 2013;116(6):746-53. doi: DOI 10.1016/j.jbiosc.2013.05.031. PubMed PMID: WOS:000329087800016. 13. Hynes RO. Integrins: versatility, modulation, and signaling in cell adhesion. Cell. 1992;69(1):11-25. PubMed PMID: 1555235. 14. Burridge K, Chrzanowska-Wodnicka M. Focal adhesions, contractility, and signaling. Annual review of cell and developmental biology. 1996;12:463-518. doi: 10.1146/annurev.cellbio.12.1.463. PubMed PMID: 8970735.
158
15. Choi SS, Diehl AM. Epithelial-to-mesenchymal transitions in the liver. Hepatology. 2009;50(6):2007-13. doi: 10.1002/hep.23196. PubMed PMID: 19824076; PMCID: 2787916. 16. Wells RG. The epithelial-to-mesenchymal transition in liver fibrosis: here today, gone tomorrow? Hepatology. 2010;51(3):737-40. doi: 10.1002/hep.23529. PubMed PMID: 20198628; PMCID: 3247701. 17. Cary LA, Chang JF, Guan JL. Stimulation of cell migration by overexpression of focal adhesion kinase and its association with Src and Fyn. J Cell Sci. 1996;109 ( Pt 7):1787-94. PubMed PMID: 8832401. 18. Semler EJ, Ranucci CS, Moghe PV. Mechanochemical manipulation of hepatocyte aggregation can selectively induce or repress liver-specific function. Biotechnol Bioeng. 2000;69(4):359-69. PubMed PMID: 10862674. 19. Chan AY, Bailly M, Zebda N, Segall JE, Condeelis JS. Role of cofilin in epidermal growth factor-stimulated actin polymerization and lamellipod protrusion. J Cell Biol. 2000;148(3):531-42. PubMed PMID: 10662778; PMCID: 2174812. 20. Tanimura S, Nomura K, Ozaki K, Tsujimoto M, Kondo T, Kohno M. Prolonged nuclear retention of activated extracellular signal-regulated kinase 1/2 is required for hepatocyte growth factor-induced cell motility. J Biol Chem. 2002;277(31):28256-64. doi: 10.1074/jbc.M202866200. PubMed PMID: 12032150. 21. Ridley AJ, Hall A. The small GTP-binding protein rho regulates the assembly of focal adhesions and actin stress fibers in response to growth factors. Cell. 1992;70(3):389-99. PubMed PMID: 1643657. 22. He W, Rose DW, Olefsky JM, Gustafson TA. Grb10 interacts differentially with the insulin receptor, insulin-like growth factor I receptor, and epidermal growth factor receptor via the Grb10 Src homology 2 (SH2) domain and a second novel domain located between the pleckstrin homology and SH2 domains. J Biol Chem. 1998;273(12):6860-7. PubMed PMID: 9506989. 23. Godoy P, Hengstler JG, Ilkavets I, Meyer C, Bachmann A, Muller A, Tuschl G, Mueller SO, Dooley S. Extracellular matrix modulates sensitivity of hepatocytes to fibroblastoid dedifferentiation and transforming growth factor beta-induced apoptosis. Hepatology. 2009;49(6):2031-43. doi: 10.1002/hep.22880. PubMed PMID: 19274752. 24. Wang P, Liu T, Cong M, Wu X, Bai Y, Yin C, An W, Wang B, Jia J, You H. Expression of extracellular matrix genes in cultured hepatic oval cells: an origin of hepatic stellate cells through transforming growth factor beta? Liver international : official journal of the International Association for the Study of the Liver. 2009;29(4):575-84. doi: 10.1111/j.1478-3231.2009.01992.x. PubMed PMID: 19323784. 25. Zhu NL, Asahina K, Wang J, Ueno A, Lazaro R, Miyaoka Y, Miyajima A, Tsukamoto H. Hepatic stellate cell-derived delta-like homolog 1 (DLK1) protein in liver regeneration. J Biol Chem. 2012;287(13):10355-67. doi: 10.1074/jbc.M111.312751. PubMed PMID: 22298767; PMCID: 3322997. 26. Mullhaupt B, Feren A, Fodor E, Jones A. Liver expression of epidermal growth factor RNA. Rapid increases in immediate-early phase of liver regeneration. J Biol Chem. 1994;269(31):19667-70. PubMed PMID: 7519599. 27. Bachem MG, Meyer D, Melchior R, Sell KM, Gressner AM. Activation of rat liver perisinusoidal lipocytes by transforming growth factors derived from myofibroblastlike cells. A potential mechanism of self perpetuation in liver fibrogenesis. The Journal of clinical investigation. 1992;89(1):19-27. doi: 10.1172/JCI115561. PubMed PMID: 1729271; PMCID: 442814.
159
28. Pinzani M, Abboud HE, Aron DC. Secretion of insulin-like growth factor-I and binding proteins by rat liver fat-storing cells: regulatory role of platelet-derived growth factor. Endocrinology. 1990;127(5):2343-9. PubMed PMID: 1699746. 29. Olsen AL, Bloomer SA, Chan EP, Gaca MD, Georges PC, Sackey B, Uemura M, Janmey PA, Wells RG. Hepatic stellate cells require a stiff environment for myofibroblastic differentiation. Am J Physiol Gastrointest Liver Physiol. 2011;301(1):G110-8. doi: 10.1152/ajpgi.00412.2010. PubMed PMID: 21527725; PMCID: 3129929. 30. Guvendiren M, Perepelyuk M, Wells RG, Burdick JA. Hydrogels with differential and patterned mechanics to study stiffness-mediated myofibroblastic differentiation of hepatic stellate cells. J Mech Behav Biomed Mater. 2014;38:198-208. doi: 10.1016/j.jmbbm.2013.11.008. PubMed PMID: 24361340.
160
Daniel Deegan
1310 Brookstown Ave. Apt. 4 (908)-627-2216
Winston-Salem, NC 27101 [email protected]
Education
Wake Forest University Graduate School of Arts and Sciences
Winston-Salem, NC
Doctorate of Philosophy in Molecular Medicine and Translational Sciences (December
2015)
Area of Specialization: Regenerative Medicine
Dissertation: “Functional Effects of Liver Specific ECM Stiffness on Primary
Hepatocytes in Culture”
Faculty Advisor: Thomas Shupe
GPA: 3.67
Virginia Polytechnic Institute and State University (Virginia Tech)
Blacksburg, VA
Bachelor of Science in Biological Sciences, Minor in Economics (May 2009)
GPA: 3.9
Research Experience
Wake Forest Institute of Regenerative Medicine - PhD Thesis Lab
Research Advisor: Dr. Thomas Shupe (2010-Present)
- Researched the recellularization of decellularized kidneys and livers for transplant
- Involved in protein analysis of the ECM and cell culture on ECM gels
- Analyzed influences of ECM bound growth factors on primary hepatocyte culture
- Analyzed effects of altered substrate stiffness on cell culture
- Experience in submitting grants, poster presentations, and seminar lecturing
- Author on publications
Wake Forest University- Molecular Medicine and Translational Science - 1st Year
Rotations
Research Advisors: Dr. Peter Antinozzi, Dr. Richard Loeser (2009 – 2010)
- Translational research performed on topics including diabetes, molecular pathways of
osteoarthritis
- Experience shadowing clinicians related to fields of research
- Abstract for American Society of Nephrology
Virginia Tech - Plant Pathology Lab, Honors Undergraduate Research
Research Advisor: Dr. John McDowell (2007 – 2009)
- Assisted graduate student and professor with research on plant disease in Arabidopsis
- Focused on GUS and trypan blue staining of plants to see when auxin responses and
cell death are triggered after infection of Hyaloperonospora parasitica
- Pictures of microscope slides used for publication
161
Rutgers University - Research Experience for Undergraduates Program, Camden, NJ
Research Advisor: Dr. Heike Bücking (Summer
2008)
- Ten week program sponsored by the National Science Foundation involved research,
reporting of findings, creating a professional poster, and writing a paper
- Explored uptake of nutrients by mycorrhizal fungi under sterile conditions using
radioactive isotopes
- Lab skills and techniques were also presented and taught through several seminars
Laboratory and Computer Skills
Real-time PCR; DNA/RNA analysis; gel electrophoresis; Western blotting; protein
quantification; ELISA; organ collection and perfusion; decellularization; surgical
techniques; immunohistochemistry; microscopy; primary human cell culture; 3D
organoid culture
Microsoft Word, Excel, Powerpoint; GraphPad
Graduate School Positions
Student Representative of the Molecular Medicine and Translational Sciences Executive
Committee
Member of Wake Forest University Honors Council
Honors and Awards
Outstanding Poster at North Carolina Tissue Engineering and Regenerative Medicine
Annual Conference and Innovation Summit, 2013
Member of Phi Sigma Biology Honors Society
Graduated summa cum laude with degree in biological sciences
Honors Scholar at Virginia Tech
Graduated with an additional 18 credit hours of coursework and research
Member of the University Honors Program at Virginia Tech (Freshman through Senior
years)
Virginia Tech Dean’s List (every semester as undergraduate)
Member of the Biological Life Sciences Community at Virginia Tech
- Freshman year program where Biology majors lived together in student housing on
campus, interacted
socially and academically, participated in seminars, and completed a community
service project
- Service project included Stroubles Creek watershed clean-up
162
Publications:
Manuscripts:
Deegan D.B., Zimmerman C., Skardal A., Atala A., Shupe T., Stiffness of Hyaluronic
Acid Gels Containing Liver Extracellular Matrix Supports Hepatocyte Function and
Alters Cell Morphology. Journal of the Mechanical Behavior of Biomedical Materials.
2015; pp. 87-103 (online)
Ko IK, Peng L., Peloso A., Smith C.J., Dhal A., Deegan D.B., Zimmerman C., Clouse C.,
Zhao W., Shupe T.D., Soker S., Yoo J.J., Atala A., Bioengineered transplantable porcine
livers with re-endothelialized vasculature. Biomaterials. 2015 Feb; 40:72-9.
Sullivan D.C., Mirmalek-Sani S.H., Deegan D.B., Aboushwareb T., Baptista P.M., Atala
A., Yoo J.J., Decellularization Methods of Porcine Kidneys for Whole Organ
Engineering Using a High-Throughput System. Biomaterials. 2012 Nov 28;
33(31):7756-64.
Antinozzi P.A., Barnes B.C., Deegan D., Ma L., Freedman B.I., Murea M. Digital
deconstruction of heterogeneous renal cell preparations with high-content imaging and
cell specific biomarkers [abstract]. Journal of the American Society of Nephrology.
2010; 21(Suppl):409A.
McDowell J.M., Hoff, T., Anderson, R., Deegan, D., Propagation, Storage, and Assays
with Hyaloperonospora arabidopsidis, a model oomycete pathogen of Arabidopsis,
Methods in Molecular Biology. 2011; 712:137-51.
Book Chapter:
Dhal, A., Vyas, D., Moran, E.C., Deegan, D.B., Soker, S., Baptista, P.M. (2014). Hepatic
Systems. In Anthony Atala (Ed.), Translational Regenerative Medicine (pp. 469-479).
Boston, MA: Academic Press.
Presentations:
Deegan D., “Effects of Liver Specific Extracellular Matrix Gels of Varying Stiffnesses
and Growth Factor Presence on Primary Hepatocyte Function.” Molecular Medicine
Seminar. Wake Forest University, Winston-Salem. 19 Sep. 2014.
Deegan D., “Effects of ECM Bound Growth Factors on Primary Hepatocyte Culture.”
Molecular Medicine Seminar. Wake Forest University, Winston-Salem. 9 Sep. 2013.
Deegan D., "Effects of ECM Bound Growth Factors on Liver Cell Function." Molecular
Medicine Seminar. Wake Forest University, Winston-Salem. 1 Mar. 2013.
Deegan D., "Growth Factor Retention on Decellularized Liver Scaffolds." Molecular
Medicine Seminar. Wake Forest University, Winston-Salem. 11 May 2012.
Deegan D., "Characterization of a Decellularized Porcine Kidney Scaffold for Cellular
Reattachment." Molecular Medicine Seminar. Wake Forest University, Winston-Salem.
25 Mar. 2011.
163
Deegan D., "Effects of SOD1 and MKK7 Expression on IGF Induced AKT and MAPK
Signaling." Molecular Medicine Seminar. Wake Forest University, Winston-Salem. 7
May 2010.
Deegan D., "Development of a High Content Cell Based Assay for Renal Disease."
Molecular Medicine Seminar. Wake Forest University, Winston-Salem. 13 Nov. 2009.
Lectures:
Guest lecture, Winthrop University, BIOL523X: Stem Cell Biology. Liver
Bioengineering. April 10, 2015.
Posters:
Deegan D., Zimmerman C., Shupe T., Effects of Liver Specific Extracellular Matrix
Hyaluronic Acid Gels of Varying Stiffnesses on Primary Hepatocyte Function. Wake
Forest Institute for Regenerative Medicine, 2015
Deegan D., Zimmerman C., Shupe T., Effects of Increased Hydrogel Crosslinking and
ECM Bound Growth Factors on Primary Hepatocyte Culture. Wake Forest Institute for
Regenerative Medicine Retreat, 2013
Deegan D., Zimmerman C., Shupe T., Effects of Extracellular Matrix Bound Growth
Factors on Primary Hepatocyte Culture. North Carolina Tissue Engineering and
Regenerative Medicine Annual Conference and Innovation Summit, 2013
Deegan D., Zimmerman C., Brown
A., Petersen
B., Shupe T., Growth factor retention on
decellularized liver matrices derived from varying growth states and serum treatment.
Wake Forest Institute for Regenerative Medicine Retreat, 2012
Deegan D.B., Sullivan D.C., Mirmalek-Sani S.H., Atala A., Yoo J.J., Aboushwareb T.,
Effects of Detergent Concentrations on Decellularized Porcine Kidney Scaffold, Wake
Forest Institute for Regenerative Medicine Retreat, 2011