Figure S1. Related to Figure 1.europepmc.org/articles/PMC5023502/bin/NIHMS807940-supplemen… ·...
Transcript of Figure S1. Related to Figure 1.europepmc.org/articles/PMC5023502/bin/NIHMS807940-supplemen… ·...

Setdb2
DM
SO
DEX
DEX+R
U48
60
5
10
15
rela
tive
mR
NA
Low High
Cpt1
Contr
ol
Wy1
4653
0
1
2
3
4
rela
tiv
e m
RN
A
Setdb2
Contr
ol
WY14
653
0.0
0.5
1.0
1.5
rela
tiv
e m
RN
A
G6pc
Contr
olIS
OFS
KDEX
Glu
cagon
Insu
lin0
5
10
15250300350400450500
rela
tiv
e m
RN
A
B C
Setdb2
TSS
-5kbTSS
ChIPseq at Setdb2 locus
DNase
DEX DNase
Vehicle
GR
DEX
GR
Vehicle
The EMBO Journal (2013) 32, 1568–1583 E
IgG GR0.0
0.1
0.2
0.3
0.4
Setdb2 (-5kbTSS)
%in
pu
t
shCTRL-PBSshCTRL-DEXshGR-DEX
F
shCTRL+PBS
shCTRL+DEX
shGR+DEX
D A *
*
*
*
IgG GR0.00
0.02
0.04
0.06
%in
pu
t
REFEDFAST
IgG SETDB20.000
0.005
0.010
0.015
Insig2a +41kb TSS
%in
pu
t
Refed
Fast
*
* *
*
*
*
Figure S1. Related to Figure 1.
G
IgG GR0.0
0.1
0.2
0.3
0.4
Setdb2 (-5kbTSS)
%in
pu
t
shCTRL-PBSshCTRL-DEXshGR-DEX
Dm
soDex
0.0
0.5
1.0
1.5
2.0
rela
tive m
RN
A
Human HepatoCellsSetdb2H
HepG2Setdb2
Dm
so
Dex
0.0
0.5
1.0
1.5
2.0
rela
tive m
RN
A
DEX (1uM)
- +
HepG2
1 2
*
SETDB2
b-actin
*
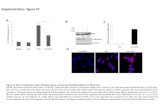
Figure S1. SETDB2 is a glucocorticoid responsive gene. A) Heat map of mouse liver gene expression profile comparing fed versus fasted state. B) mRNA
levels of mouse primary hepatocytes (MPH) treated with various stress/fasting-related compounds. C) PPAR-a agonist increases Cpt1a but not Setdb2 in
MPH; G6Pase and Cpt1a were used as treatment controls. D) Setdb2 mRNA in MPH treated with DEX in presence or absence of RU486. E) Setdb2 gene
track showing DEX-responsive DNase and GR binding site at -5kb of the Setdb2 TSS. F) GR ChIP-qPCR at -5kbTSS of Setdb2 gene locus +/- DEX. G) GR
enrichment at Setdb2 -5kbTSS with fasting. (n=3,*p<0.05 relative to control). H) Setdb2 mRNA in human HepatoCells and HepG2 treated with and without
Dex (1uM) for 6h, and nuclear SETDB2 immunoblot of Dex-treated HepG2. *p<0.05 relative to vehicle treatment. Data are represented as mean ± SEM.

IgG GR0.000
0.005
0.010
0.015
0.020
0.025
%in
pu
t
Sult1e1
IgG GR0.00
0.02
0.04
0.06
0.08
0.10
%in
pu
t
Mt2
b
a
b
b
a
b
shCtrl
shCtrl
shDB2
0
1
2
3
4
5Mt2
rela
tive m
RN
A
shCtrl
shCtrl
shDB2
0
5
10
15
20
25Sult1e1
rela
tive m
RN
A PBS
DEXDEX
a
b
a
b
b b PBSDEXDEX
A
B
shCtrl+PBS
shCtrl+DEX
shDB2+DEX
shCtrl+PBS
shCtrl+DEX
shDB2+DEX
Figure S2. SETDB2 is required for DEX-mediated activation of
select GR targets. Non-SETDB2 regulated GR targets in liver. A)
Liver mRNA of Mt2 and Sult1e1 in Dex-treated mice. B) GR-ChIP
qPCR at promoters of GR targets not regulated by SETDB2. n=3.
Groups with different letters are statistically different, p<0.05. Data are
represented as mean ± SEM.
Figure S2. Related to Figure 2.

shCtrl
shDB2
0
5
10
15
ug
/mg
tis
su
e
Liver TAG
IgG GR SETDB20.00
0.01
0.02
0.04
0.06
0.08
0.10
%in
pu
t
Insig2a (-1.6kbTSS)
Contr
ol
Isopro
tere
nol
Forsko
linDex
Glu
cagon
Insu
lin
WY14
653
0
1
2
3
4
rela
tiv
e m
RN
A
Insig2a
Contr
olDex
Dex
+Ru48
6
Insu
lin
Insu
lin+D
ex0
5
10
15
rela
tive m
RN
A
Insig2a
* *
*
A B
Refed Fast0
2
4
6
8
rela
tive m
RN
A
Insig2a
shCtrlshDB2
Refed Fast0.0
0.5
1.0
1.5
2.0
2.5
rela
tive m
RN
A
Setdb2
shCtrlshDB2
*
C
*
*
IgG GR SETDB20.00
0.05
0.10
%in
pu
t
Insig2a (-3.2kbTSS)
IgG GR SETDB20.00
0.05
0.10
0.15
0.20
%in
pu
tInsig2a (exon1)
shCtrl+PBSshCtrl+DEXshDB2+DEXshGR+DEX
IgG GR0.00
0.02
0.04
0.06
0.08
0.10
%in
pu
t
Insig2 promoter
Refed
Fast
IgG
-3.2
kb
-1.6
kbTS
S
Exo
n10.00
0.05
0.10
0.15
0.20
0.25
%in
pu
t
SETDB2shCtrl+PBSshCtrl+DEXshDB2+DEXshGR+DEX
IgG
-3.2
kb
-1.6
kbTS
S
Exo
n10.00
0.05
0.10
0.15
0.20
0.25
%in
pu
t
SETDB2shCtrl+PBSshCtrl+DEXshDB2+DEXshGR+DEX
E F G
*
* b
a
a,b
c
b
a a
c
b
a
c
d b
a a
b
*
D
Figure S3. SETDB2 and GR are required for Dex-mediated activation of Insig2a. A) Insig2a mRNA in mouse primary hepatocytes
(MPH) treated with various stress/fasting-related compounds (4hr). B) MPH Insig2a mRNA induction by Dex is inhibited by RU486 or
Insulin (18hr). C) Liver Insig2a mRNA induction by fasting is SETDB2 dependent. D) Hepatic triglycerides in adlib chow fed mice infected
with control or shSETDB2 adenovirus. GR and SETDB2 ChIP-qPCR at E) -3.2kbTSS and F) exon1 of livers with and without SETDB2 or
GR. G) Fasting-induced GR and SETDB2 binding at Insig2a promoter. *p<0.05 relative to vehicle treatment or Refed state. Groups with
different letter are statistically different (p<0.05). Data are represented as mean ± SEM.
Figure S3. Related to Figure 3.

Tat
Insi
g2
Cyp
2b10
0.0
0.5
1.0
1.5
%in
pu
t
Tat
Insi
g2
Cyp
2b10
0.00
0.25
0.50
0.75
1.00
1.25
%in
pu
t
H3K9me1 H3K9me2 B A
Tat
Insi
g2
Cyp
2b10
0.00
0.02
0.04
0.06
0.08
0.10
%in
pu
t
IgG D
PBSDEXDEX
C H3
Figure S4. Related to Figure 4.
shCtrl
shCtrl
shDB2
0
1
2
3Tat promoter
H3K
9m
e3/H
3 r
ati
o
shCtrl
shCtrl
shDB2
0.0
0.5
1.0
1.5
2.0
H3K
9m
e3/H
3 r
ati
o
Insig2a promoterF
shCtrl
shCtrl
shDB2
0.0
0.5
1.0
1.5
2.0
Cyp2b10 promoter
H3K
9m
e3/H
3 r
ati
o
shCtrl
shCtrl
shDB2
0.0
0.5
1.0
1.5
Lcn2 promoter
H3K
9m
e3/H
3 r
ati
o
G H
Figure S4. SETDB2 does not correlate with H3K9 methylation at promoters of Dex-induced GR targets. Liver ChIP-qPCR at promoters of
SETDB2-GR regulated genes. A) H3K9me1. B) H3K9me2. C) Total H3. D) IgG. H3K9me3/Total H3 ratios at promoters of SETDB2-GR regulated
genes in livers treated with shControl or shDB2 in presence or absence of Dex. E) Insig2a. F) Tat. G) Cyp2b10. H) Lcn2. Data are represented as mean
± SEM.
Tat
Insi
g2
Cyp
2b10
0.0
0.2
0.4
0.6
0.8
%in
pu
t
PBS
DEX
E

IgG GR0.0
0.5
1.0
1.5Lcn2 promoter
%in
pu
t
KO
FLP
FRT
loxP loxP
loxP
LacZ/
STOP
Neo/
STOP Exon5 Exon6 Exon7
loxP loxP Exon5 Exon6 Exon7
DB2
(GT)
DB2
(FLP)
A
Figure S5. Related to Figure 5.
FRT
DB2(
GT)
DB2(
FLP)
0
1
2
3
rela
tive m
RN
A
Lcn2
WT
WT
DB2(
GT)
0
2
4
6
8
10re
lative m
RN
A
Setdb2
WT
WT
DB2(
GT)
0
200
400
600
800
rela
tive m
RN
A
Lcn2B
DB2(
GT)
DB2(
FLP)
0
1
2
3
4
5Setdb2
rela
tive m
RN
A
C
* * PBSDEXDEX
* *
*
IgG Rad210.00
0.02
0.04
0.06
0.08
0.10Lcn2 promoter
%in
pu
t
KO
FLP
IgG SETDB20.00
0.01
0.02
0.03
0.04Lcn2 promoter
%in
pu
t
KO
FLP
IgG GR0.00
0.01
0.02
0.03Insig2 enhancer
%in
pu
t
DB2(GT)
DB2(FLP)
D
IgG GR0.00
0.02
0.04
0.06
0.08Lcn2 enhancer
%in
pu
t
* * * *
Figure S5. Long range chromatin interaction of enhancer-promoter at Lcn2 locus after Flp-FRT recombination. A) DB2 (GT): Setdb2
hypomorphic gene trap allele results in SETDB2 deficiency. DB2 (FLP): Setdb2 floxed allele without gene trap and restored SETDB2 expression due to
Flp-FRT recombination. Liver Setdb2 and Lcn2 mRNA after Dex treatment in B) DB2 (GT) and C) FLP mice. D) GR, SETDB2, RAD21 ChIP-qPCR at
Lcn2 promoter and GR-ChIP at Lcn2 enhancer. *p<0.05 relative to control. Data are represented as mean ± SEM.
PBSDEXDEX
IgG GR0.00
0.01
0.02
0.03Insig2 enhancer
%in
pu
t
DB2(GT)
DB2(FLP)
IgG GR0.00
0.01
0.02
0.03Insig2 enhancer
%in
pu
t
DB2(GT)
DB2(FLP)
IgG GR0.00
0.01
0.02
0.03Insig2 enhancer
%in
pu
t
DB2(GT)
DB2(FLP)

IgG GR0.0
0.1
0.2
0.3
0.4
0.5Insig2 promoter
%in
pu
t
KO
FLP
IgG GR0.00
0.01
0.02
0.03Insig2 enhancer
%in
pu
tDB2(GT)
DB2(FLP)
IgG SETDB20.00
0.01
0.02
0.03
0.04Insig2 enhancer
%in
pu
t
KO
FLP
IgG SETDB20.00
0.01
0.02
0.03
0.04Insig2 promoter
%in
pu
t
KO
FLP
IgG GR0.00
0.01
0.02
0.03Insig2 enhancer
%in
pu
t
DB2(GT)
DB2(FLP)
E
IgG GR0.00
0.01
0.02
0.03Insig2 enhancer
%in
pu
t
DB2(GT)
DB2(FLP)
IgG GR0.00
0.01
0.02
0.03Insig2 enhancer
%in
pu
t
DB2(GT)
DB2(FLP)
IgG GR0.00
0.01
0.02
0.03Insig2 enhancer
%in
pu
t
DB2(GT)
DB2(FLP)* * * *
Figure S6. Related to Figure 6.
IgG GR0.00
0.01
0.02
0.03
%in
pu
t
Insig2a enhancer +41kbTSS
Refed
Fast
IgG SETDB20.000
0.005
0.010
0.015
Insig2a +41kb TSS
%in
pu
t
Refed
Fast
Ref
edFas
t0
5
10
15
rela
tive e
RN
A
Insig2a enhancer
+41kbTSS
IgG SETDB20.000
0.005
0.010
0.015
Insig2a +41kb TSS
%in
pu
t
Refed
Fast IgG SETDB20.000
0.005
0.010
0.015
Insig2a +41kb TSS
%in
pu
t
Refed
Fast
A B
WT
WT
DB2(
GT)
DB2(
FLP)
0
2
4
6
rela
tive m
RN
A
Insig2aD
IgG SETDB20.000
0.005
0.010
0.015
Insig2a enhancer +41kb TSS
%in
pu
t
Refed
Fast
PBSDEXDEX
* * *
IgG SETDB20.000
0.005
0.010
0.015
Insig2a +41kb TSS
%in
pu
t
Refed
Fast
C
*
* *
IgG SETDB20.000
0.005
0.010
0.015
Insig2a +41kb TSS
%in
pu
t
Refed
Fast
GFP
hDB2M
UT
0.0
0.5
1.0
1.5
2.0
2.5Insig2a
rela
tive m
RN
A
GFP
hDB2M
UT
0.0
0.5
1.0
1.5Insig2b
rela
tive m
RN
A
GFP
hDB2M
UT
0.0
0.5
1.0
1.5
2.0
2.5Lcn2
rela
tive m
RN
A
GFP
hDB2M
UT
0
1
2
3
4Cyp2b10
rela
tive m
RN
A
GFP
hDB2M
UT
0.0
0.5
1.0
1.5
2.0
2.5Tat
rela
tive m
RN
A
GFP
hDB2M
UT
0.0
0.5
1.0
1.5hSetdb2
rela
tive m
RN
A *
* * *
* *
ND
*
*
F
Figure S6. Long range chromatin interaction of enhancer-promoter at Insig2. A) Insig2 enhancer RNA of mouse liver in response to 24h fasting. B) Liver ChIP-qPCR of
GR and C) SETDB2 at Insig2 enhancer of livers in refed and fasted state. D) Insig2a liver mRNA +/- 2-hr Dex (10mg/kg). E) Liver ChIP-qPCR of GR and SETDB2 at Insig2
promoter (-1.6kbTSS) and enhancer (+41kbTSS). F) mRNA of SETDB2-GR gene targets in DB2GT primary hepatocytes with restored Setdb2 expression via human wild type
or mutant SETDB2 adenoviral infection (48hr) and treated with Dex (1uM) for 16h. *p<0.05 relative to control. Data are represented as mean ± SEM. ND=Not Detected.

0 1.0
Refed 8hr
0.1 10 100 DEX (mg/kg)
N-SREBP-1
YY-1
N-SREBP-2
M-INSIG2
M-Transferrin
Receptor
1 2 3 4 5 6 7 8 9 10
Figure S7. Dex inhibition of SREBP processing during refeeding. Liver immunoblot of nuclear
SREBP1, SREBP2, and membrane INSIG2 at different DEX doses during a refed state. N-Nuclear, M-
Membrane. Each lane represents protein from individual mice.
Figure S7. Related to Figure 7.

Table S1. Related to Figure 1.
Table S1. Microarray of fed versus fasted (24h) mouse livers. Submitted as excel file.

Gene EntrezID N_TPM F_TPM R_TPM F/R
Setdb2 239122 4.49 7.65 1.03 7.43
Setd4 224440 1.34 2.87 1.44 1.99
Setd7 73251 3.72 4.50 2.98 1.51
Setd2 235626 23.96 23.15 21.39 1.08
Setdb1 84505 7.54 7.75 8.95 0.87
Smyd5 232187 2.58 3.83 4.53 0.85
G9a 110147 6.59 5.07 6.79 0.75
Smyd2 226830 2.10 3.35 6.48 0.52
Table S2. Gene expression of SET domain containing proteins in liver during normal feeding
(N), refed (R) or fasted (F) state. (Table created from published RNA-seq analysis by Zhang F.
et, al. PLoS One. 2011;6(11))
Table S2. Related to Figure 1.

Table S3. Related to Figure 2.
Table S3. RNA-seq of mouse primary hepatocytes treated with shControl or shSETDB2 adenovirus for 72h prior to 4h Dex
treatment. Each treatment represents a pool of 3 mice. Submitted as excel file.

SETDB2/GR
regulated GR regulated
Aass Nnmt Agpat6 G0s2 Nedd4l Smox
Ahr Phf8 Ankhd1 Gadd45g Npm3 Snx10
Arhgef3 Pqlc1 Anubl1 Got1 Nup98 Spata2L
Ccng2 Prkd3 Apbb3 Hisppd1 Pctk2 Sult1e1
Chd1l Sec22c Apobec1 Fmo5 Pex11a Thbd
Cldn3 Serpina5 Atp6v0a4 Lcn13 Ppargc1a Tm6sf1
Cyp2b10 Slc10a2 Bcl2l1 Lpin1 Ppp1r3c Tpst2
Gdf15 Slc45a3 Btg2 Lrrc58 Pptc7 Uap1
Herpud1 Smox Cab39l Map3k5 Pxmp4 Ube2g2
Igfbp1 Snx10 Cdc37l1 Mbd1 Rbbp8 Zh2c2
Insig2 Tat Cdkn1a Mcart1 S3-12
Lcn2 Tesk2 Dusp14 Mdm1 Sec14l4
Lyve1 Tff3 EG545306 Megf9 Sgk1
Mical3 Tgm1 Egln3 Mrap Slc25a32
Nedd4l Tiparp Ell2 Ms4a6d Slc38a2
Nek7 Trp53inp1 Fkbp5 Mt1 Slc43a1
Nfil3 Tsc22d3 Fmo5 Mt2 Slc45a3
Table S4. DEX-induced GR-dependent genes (Endocrinology. 2010 Jan;151(1):185-94.) that
are SETDB2 co-regulated and not co-regulated. Table was generated comparing Table S3 to
microarray from (Wong et al., 2010).
Table S4. Related to Figure 2.

Table S5. Related to Experimental Procedures.
Table S5. A) Primers for mRNA and eRNA. B) Primers for ChIP-qPCR. C) Primers for 3C-qPCR. Submitted as excel file.

SUPPLEMENTAL EXPERIMENTAL PROCEDURES
Cell Culture
Mouse primary hepatocytes (MPH) were isolated from 10-week old male mice by liver collagenase perfusion as described
previously (Li et al., 2010) with minor modifications. Mice were anesthetized using a ketamine/xylazine mixture and the liver
was perfused with Earle's balanced salt solution (EBSS) (Life Technologies) supplemented with 0.5 mM EGTA through the
inferior vena cava at a flow rate of 4 ml/min followed by perfusion with EBSS supplemented with 0.3 mg/ml collagenase
(Sigma C2139) and 4.8 mM CaCl2. The liver was dissected from the mouse and cells were dispersed gently in Medium 199
(Life Technologies) supplemented with 10% heat-inactivated fetal bovine serum (FBS; Atlanta Biologicals). Cells were
filtered through a 100-μm-nylon cell strainer (BD Falcon), washed, and hepatocytes were separated by 45% Percoll solution
(Sigma) density gradient centrifugation. Hepatocyte viability was monitored by trypan blue exclusion and more than 90% of
cells were consistently viable. Isolated MPH were plated in collagen-coated 6-well dishes (BD Biocoat) at a density of 6 ×
105 cells per well.
After a 4-hour attachment, MPH was maintained overnight in serum-free Medium199 prior to treatment or adenoviral
infection. On day 2, MPH were treated with either 100 nM insulin (Sigma, I9278), 1.0 uM isoproterenol (Sigma, I5627), 1.0
uM forskolin (Sigma, F3917), 10 uM WY14653, 1.0 uM dexamethasone (Sigma, D4902) or 100 nM glucagon (Sigma,
G2044) for 4 hours prior to harvest for RNA analysis. For adenovirus-mediated SETDB2 knockdown or overexpression
studies, MPH were infected with adenoviral constructs in medium 199 supplemented with 2% heat-inactivated FBS and 10
nM Dex. Duration of adenoviral treatment was 48 hours at a multiplicity of infection of 40. Infection media was replaced
with serum-free Medium 199 24 hours prior to Dex (1.0 uM) treatment. In SETDB2 knockdown studies, MPH were treated
with either an shControl adenoviral construct or an shSETDB2 construct targeting the 3’UTR of mouse SETDB2
(Invitrogen). In SETDB2 overexpression studies, MPH from DB2GT mice were infected with an adenoviral construct
encoding a control GFP, a human SETDB2 protein, or mutant form of human SETDB2 that contains point mutations at
critical regions within the SET domain that are required for S-adenosylmethionine binding and catalytic activity (N639A and
H640A).
Human HepatoCells (Corning) were plated in a 96-well collagen-coated plate according to manufacture’s instructions.
Plating media consisted of Corning Culture Medium for HepatoCells supplemented with 10%FBS. After 6h, media was
removed and cell monolayer was overlayed with matrigel solution (0.25mg/ml). Cells were incubated overnight. On days 2
and 3, media was replaced with sera-free HepatoCell medium. On Day 4, cells were treated with vehicle or Dex (1uM) for
6h.
HepG2 (ATCC) cells were plated in DMEM supplemented with 10%FBS and 1% penicillin-streptomycin. Cells were seeded
in 6-well plates at 5x10^5 cells per well. On day2, media was replaced with sera-free DMEM for 24h prior to vehicle or Dex
treatment (1uM) for 6h.
Co-immunoprecipitation
Co-immunoprecipitation of GR and SETDB2 was performed using 500 ug total protein isolated from MPH treated with 1.0
uM Dex for 2 hours. Immunoprecipitation reactions were incubated overnight in phosphate-buffered saline (PBS) containing
protease inhibitors and either Rabbit IgG, GR antibody (Santa Cruz Biotechnology, SC-8892), or SETDB2 antibody (Abcam,
ab13712/clone33-1). Antibody-protein complexes were precipitated using Dynabeads Protein G magnetic beads (Life
Technologies) for 6 hours. Beads were washed with RIPA buffer followed by sequential PBS washes prior to eluting the
protein in SDS-PAGE loading buffer by boiling for 5 min.
Generation of SETDB2 antibody
Custom rabbit polyclonal SETDB2 antibodies were produced by New England Peptide, LLC using a recombinant protein
fragment corresponding to residues 3-60 of the mouse SETDB2. We chose this motif because it is uniquely encoded in
SETDB2 proteins and it is also the most conserved domain between the human and mouse SETDB2 proteins. The peptide
fragment was fused to an N-terminal (6x) histidine tag and over-expressed using a bacterial expression system. The fusion
peptide was purified at high yield using Ni-affinity followed by Superdex 75 size exclusion chromatography. The pure
fusion protein was dissolved at 3.6 mg/ml in 10 mM HEPES buffer (pH 7.5) containing 150 mM NaCl for use as an antigen
for injection into rabbits by New England Peptide. Antibodies were affinity-purified against the antigen and judged as pure
and homogeneous when analyzed by superose 6 size exclusion column and SDS-PAGE. Clone 33-1 was used in
immunoprecipitation and immunoblot experiments.

Protein Analysis
Membrane protein from mouse liver was isolated as described previously (Engelking et al., 2004) with minor modifications.
Briefly, 100mg of frozen liver was dounced in buffer (20 mM Tris-HCl at pH 7.4, 2 mM MgCl2, 0.25 mM sucrose, 10 mM
sodium EDTA, and 10 mM sodium EGTA) followed by low-speed centrifugation (1000 x g, 4°C, 5 min). The supernatant
was used to isolate membrane protein and the pellet was lysed for nuclear protein analysis. For membrane fraction,
supernatants were transferred to 1.5 mL Beckman tubes (catalog #357448) and centrifuged at 100,000 x g for 45 min at 4°C.
The resulting membrane pellets were lysed with SDS-containing buffer, protein was quantified, and samples were diluted
with urea buffer to a final concentration of 2.0 ug/uL; membrane protein was incubated at 37°C for 20 minutes prior to SDS-
PAGE. Supernatants from membrane isolations were used for cytoplasmic protein analysis.
Protein samples were subject to 7.5% or 10% SDS-PAGE and transferred to nitrocellulose membranes. The membranes were
incubated with appropriate primary antibody followed by the corresponding secondary antibody conjugated to horseradish
peroxidase. Reactivity was detected with the SuperSignal West enhanced chemiluminescent kit (Pierce). The primary
antibodies used are as follows: GR (Santa Cruz, SC-8892), pGRser211 (Cell Signaling, 4161), SETDB2 (Abcam, 13712),
YY1 (Santa Cruz, SC281), SREBP1 (IgG-2A4), SREBP2 (Seo et al., 2011), Transferrin Receptor (Life Technologies),
Calnexin (Abcam, ab22595). The INSIG2 antibody was described previously (Haas et al., 2012).
Immunofluorescence
MPH or Hepa1-6 cells were seeded in 4-well chamber slides (Thermo Scientific Nunc) at a density of 1x105 cells per well,
washed twice with PBS, fixed with 4% paraformaldehyde (PFA) for 20 min, washed 3x with PBS, and blocked using 3%
BSA/0.5% Triton X-100/5% Signal FX enhancer (Life Technologies) for 1 hour at room temperature. Primary antibody
incubations were performed overnight at 4°C in blocking buffer. Cells were washed 3x in PBS/0.5% Tween-20 followed by
secondary antibody incubation for 2 hours in the dark at room temperature in blocking buffer with slow shaking. Cells were
washed 3x in PBS/0.5% Tween-20 and stained with Hoechst 33258 (1.0 ug/ml). All images were captured using a Nikon
A1R VAAS confocal microscope. Primary antibodies used for immunofluorescence: GR (SC-8892) and SETDB2
(abcam13712). Secondary antibodies: donkey anti-goat Alexa Fluor-488 and donkey anti-rabbit Alexa Fluor-647.
RNA sequencing and analysis
Total RNA from MPH treated with either shControl or shSETDB2 adenovirus (pool of 3 mice per group) was isolated by the
TRIzol method (Life Technologies) according to manufacturer’s instructions. Total RNA was DNase-treated and RNA
integrity was evaluated by Biolanalyzer (Agilent). RNA sequencing was performed by the SBP Genomics Core Facility in
Lake Nona (Orlando, FL) using the Illumina HiSeq platform and the following parameters: paired-end read length: 100 BP;
number of output reads: 300m reads. The Tophat+Cufflinks pipeline was used for the RNA-seq analysis as described
previously (Haas et al., 2013; Trapnell et al., 2012). Differentially expressed genes in shSETDB2 vs. shControl MPH were
determined using a 2-fold cutoff, p-value of less than 0.05, and FPKM (fragments per kilobase of transcript per million
mapped reads) cutoff of 1. The shSETDB2 and shControl RNA-seq reads were mapped to the mm10 genome using Tophat
(Trapnell et al., 2009). The mapping results were further analyzed by Cufflinks, which outputs the final differential gene list.
Cufflinks directly samples from the beta negative binomial model for each transcript per treatment in order to estimate the
null distribution of its log fold change under the null hypothesis.
Liver microarray
Heatmap was generated using a previously published microarray from our lab comparing fed versus fasted mouse liver
(n=3mice), using a 2-fold change cutoff and p-value of less than 0.05. Statistical analyses of this data set was described
previously (Shin et al., 2012).
Real Time PCR
Total RNA was prepared from mouse tissues using a 2010 Geno/Grinder (Spex Sample Prep) and the TRIzol method (Life
Technologies) according to the manufacturer’s instructions. cDNA was synthesized from 1.0 μg of DNase I–treated total
RNA using the iScript cDNA synthesis kit (Bio-Rad). Specific primers for each gene were designed using Primer Blast
(NCBI resources). Quantitative real-time PCR (qPCR) reactions (10 uL) contained 20 ng of reverse-transcribed total RNA,
forward and reverse primers, and 2x SsoFast Evagreen Supermix (Bio-Rad). qPCR reactions were performed using the Bio-
Rad CFX96 Real-Time PCR Detection System. All reactions were performed in triplicate. Relative mRNA quantities were
calculated using the comparative threshold cycle (CT) method and normalized to L32 expression. The primers used for qPCR

are provided in Table S5.
Chromatin immunoprecipitation analysis
Chromatin immunoprecipitation (ChIP) assays were performed as follows: 1.0 gram liver tissue was finely minced and fixed
in a 1.0 mM ethylene glycol-bis (succinimidylsuccinate) (EGS; Thermo Scientific, #21565)/PBS solution for 45 min at room
temperature. Following EGS fixation, samples were washed with PBS, fixed in 1% formaldehyde (Thermo Scientific,
#28908) for 10 min at room temperature, treated with 125 mM glycine/PBS for 5 min, and washed a final time with PBS.
The fixed liver tissue pellet was dounced in 6.0 ml of homogenization buffer (10 mM HEPES, pH 7.6, 25 mM KCl, 1.0 mM
EDTA, 1.0 mM EGTA, 2.0 M sucrose, 10% glycerol, 0.15 mM spermine) and overlayed on 3.0 ml of homogenization buffer
in Beckman ultracentrifuge tubes (No. 331372). Samples were centrifuged at 26K rpm using a Beckman SW41 rotor. After
separation, the nuclei pellet was lysed and sonicated until DNA fragments of 200-500 base pairs were achieved. 50 ug of
chromatin was used for each ChIP. Antibodies used were the following: GR (SC-8892), RAD21 (Abcam, ab-992), H3K9me1
(Active Motif, 39888), H3K9me2 (Diagenode Inc., C15410060), H3K9me3 (Active Motif, 61013), Total H3 (abcam1791),
SETDB2 (Abcam13712 / clone33-1).
Triglyceride extraction and quantification
Triglycerides were extracted from whole liver lysates following the Folch method (Folch, 1957). Briefly, approximately
50mg of liver was homogenized in 1ml of chloroform: methanol (2:1) solution in a glass tube and vortexed. Each sample
then received 300ul of water and was vortexed, followed by a 20 minute centrifugation (1000g, 4C) step. The lower phase
was collected, evaporated, and resuspended in 1ml of chloroform. An aliquot of the chloroform sample was used to measure
triglycerides content according to a standard assay kit from Thermo Scientific (Thermo Scientific, TR22421).
Statistics
The data are presented as mean ± SEM, as detailed in the figure legends. Differences between the means of the individual
groups were assessed by one-way ANOVA or a Student’s t test. Differences were considered significant at p < 0.05. The
statistical software package Prism 6.0 (GraphPad) was used for these analyses.
SUPPLEMENTAL REFERENCES
Engelking, L.J., Kuriyama, H., Hammer, R.E., Horton, J.D., Brown, M.S., Goldstein, J.L., and Liang, G. (2004).
Overexpression of Insig-1 in the livers of transgenic mice inhibits SREBP processing and reduces insulin-stimulated
lipogenesis. J Clin Invest 113, 1168-1175.
Folch, J.L., M; Sloane Stanley GH (1957). A simple method for the isolation and purification of total lipides from animal
tissues. J Biol Chem 226, 497-509.
Haas, B.J., Papanicolaou, A., Yassour, M., Grabherr, M., Blood, P.D., Bowden, J., Couger, M.B., Eccles, D., Li, B., Lieber,
M., et al. (2013). De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference
generation and analysis. Nat Protoc 8, 1494-1512.
Haas, J.T., Miao, J., Chanda, D., Wang, Y., Zhao, E., Haas, M.E., Hirschey, M., Vaitheesvaran, B., Farese, R.V., Jr.,
Kurland, I.J., et al. (2012). Hepatic insulin signaling is required for obesity-dependent expression of SREBP-1c mRNA but
not for feeding-dependent expression. Cell Metab 15, 873-884.
Li, W.C., Ralphs, K.L., and Tosh, D. (2010). Isolation and culture of adult mouse hepatocytes. Methods Mol Biol 633, 185-
196.
Seo, Y.K., Jeon, T.I., Chong, H.K., Biesinger, J., Xie, X., and Osborne, T.F. (2011). Genome-wide localization of SREBP-2
in hepatic chromatin predicts a role in autophagy. Cell Metab 13, 367-375.
Shin, D.J., Joshi, P., Hong, S.H., Mosure, K., Shin, D.G., and Osborne, T.F. (2012). Genome-wide analysis of FoxO1 binding
in hepatic chromatin: potential involvement of FoxO1 in linking retinoid signaling to hepatic gluconeogenesis. Nucleic Acids
Res 40, 11499-11509.
Trapnell, C., Pachter, L., and Salzberg, S.L. (2009). TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25,
1105-1111.
Trapnell, C., Roberts, A., Goff, L., Pertea, G., Kim, D., Kelley, D.R., Pimentel, H., Salzberg, S.L., Rinn, J.L., and Pachter, L.
(2012). Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc
7, 562-578.