Sequence alignment
description
Transcript of Sequence alignment

Sequence alignment
School B&I TCD Bioinformatics
May 2010

What is an alignment?• CENTRAL concept in bioinformatics• Easy if straight-forward, similar seqs
– THISTHESAME or THISTHESAME– | |||| ||| | ||| ||– TOSSTHEGAME TRANTHELPME
• Hard and CPU-intensive if seqs v. diff.– THISTHESAME vs THATGAMETHE– THISTHESAME--- or THIS----THESAME– || ||| || |||– THAT---GAMETHE THATGAMETHE
< better

Why align?
• Trying to establish homology by similarity
• Homology – having a common ancestor– whale fin, bat wing, human hand (Cuvier)– human beta globin, dog beta globin– human beta globin, human alpha globin
• You can have % similarity, % identity
• Can’t have % homology
paralog
ortholog

Why homology?
• homologous structures/molecules have similar function.
• related by evolution. – more similar seqs = more recent common
ancestor = more likely similar function
• human hand not for locomotion
• bricolage – evolutionary tinkering

Define terms
• Indel– Insertion or deletion– May get a better alignment if you put a gap in
one sequence
• Implies a mutation in one of the seqs– Not clear if insert in one or delete in the other

Optimal alignment• Best guess at evolutionary relationship
– Which residues/bases are homologous
• Depends on model of evolution and parameters of alignment– Is a gap more likely than a substitution– Is one substitution more likely than another– Transition (Y-Y or R-R) vs transversion (R-Y)– Similar shape amino acid or different
• No “correct” answer.

Global alignment• Needleman & Wunsch• Tries to align two sequences from 5’ to 3’ or C
terminus to N terminus• Assumes (only works well if) seqs are similar
over their entire length• So less good if there are large indels (but can
identify such features)• Assesses overall (functional) similarity) LARGGHYFGKISTGREFDN L FGKI T E LNAHILSFGKISTSLEDA • Identify (and count) every difference/mutation

Local alignment
• Smith & Waterman
• Ignores whole and focuses on region or domain
• Use to make high quality alignments
• …that has good similarity ----------FGKI----------
||||
----------FGKI----------
BLAST:Basic local alignment search tool

Algorithm
• Both local and global alignment programs use “dynamic programming” (wikipedia that)
• … to make optimal alignment– the alignment that tells evolutionary story
• True story unknown without time-travel
– the alignment that has the highest score
• Choose/change parameters to maximise score

2 sequence alignment
aligning GARFIELDTHECAT & GARFIELDTHERAT is easy
GARFIELDTHECAT
||||||||||| ||
GARFIELDTHERAT

Scoring systems DNA
• In an alignment add 1 if bases identical 0 if they are different
• Transition/transversion?– AG purines CUT pyrimidines
A T C G
A 1 0 0 0
T 0 1 0 0
C 0 0 1 0
G 0 0 0 1
A T C G
A 2 0 0 1
T 0 2 1 0
C 0 1 2 0
G 1 0 0 2

Scoring comparison DNA
• CTAGCGATGC• CGAACGACAC• 1010111001 1/0 Score = 6/10• 2021222112 Ts/Tv score = 15/20
• transitions 5x more common that tranversions

Insert gaps
Sometimes, you can get a better overall alignment if you insert gaps
GARFIELDTHECAT |||||||| ||| GARFIELDA--CAT is better (scores higher) than GARFIELDTHECAT |||||||| GARFIELDACAT

No gap penalty
But there must be some sort of a gap-penalty or you can align ANY two sequences:
G-R--E------AT
| | | ||
GARFIELDTHECAT

Gap penalty
• Could set a –ve score for each indel– Linear gap penalty
• But mutation could be point or deletion– latter is a single event
• Advise to use affine (open + extend)– Open –10, extend -0.05
• How choose penalty? – Start with program defaults– Use good judgment - trial and error– Investigate statistical distribution of indels

Scoring for similarities: proteins
• Gap penalty?
• Traded vs positive scores for matches in aligned residues
• Could, as with DNA, use – match=1 mismatch=0
• Or …

Scoring system proteins
• When doing a similarity search against a database you are trying to decide which of many sequences is the CLOSEST match to your search sequence. Which of the following alignment pairs is better?:
FGDERTHHSFGD--DHRS
FGDERTHHSFGDD--HRS
FGDERTHHSFGD-D-HRS
Where put the gap?

3 Garfield relatives
GARFIELDTHECAT |||| |||||||GARFRIEDTHECAT GARFIELDTHECAT ||| ||| ||||| GARWIELESHECAT GARFIELDTHECAT || ||||||| || GAVGIELDTHEMAT

Substitution matricesTop left part of a BLOSUM 90 matrix A R N D C Q E G H I LA 5 -2 -2 -3 -1 -1 -1 0 -2 -2 -2R -2 6 -1 -3 -5 1 -1 -3 0 -4 -3N -2 -1 7 1 -4 0 -1 -1 0 -4 -4D -3 -3 1 7 -5 -1 1 -2 -2 -5 -5C -1 -5 -4 -5 9 -4 -6 -4 -5 -2 -2Q -1 1 0 -1 -4 7 2 -3 1 -4 -3E -1 -1 -1 1 -6 2 6 -3 -1 -4 -4G 0 -3 -1 -2 -4 -3 -3 6 -3 -5 -5H -2 0 0 -2 -5 1 -1 -3 8 -4 -4I -2 -4 -4 -5 -2 -4 -4 -5 -4 5 1L -2 -3 -4 -5 -2 -3 -4 -5 -4 1 5
Symmetrical!
conservative subst.

Willie Taylor’s AA Venn Diagram

Substitution matrices
• Plenty of choice– Identical = 1.0; similar (K/R, F/Y) = 0.5; rest 0.0– PAM series, BLOSUM series, others
• Based on observations and counting in real seqs• Blosum 90 made from aligned seqs 90% identical
• Main diagonal elements positive– Some more positive than others– More highly conserved (C, F etc.)
• Off-diagonal elements mostly negative– Some more negative than others (less likely)– Some positive score (K-R, D-E etc.)

Dotplot theory
A T G A T A T T C T T A . . . . . . . . . . . T . . . . . . . . . . . T . . . . . . . . . . . G . . . . . . . . . . . T . . . . . . . . . . . T . . . . . . . . . . . C . . . . . . . . . . .
Task: align ATGATATTCTT and ATTGTTC
Another way of comparing 2 sequences

A T G A T A T T C T T A . . . . . . . . . . . T . + . . + . + . . + . T . . . . . . . . . . . G . . . . . . . . . . . T . . . . . . . . . . . T . . . . . . . . . . . C . . . . . . . . . . .
Go along the first seq inserting a + wherever 2/3 bases in a moving window match. The first seq is compared to ATT (the first 3 bases in the vertical sequence)
Windowsize = 3Threshold = 2

A T G A T A T T C T T A . . . . . . . . . . . T . + . . + . + . . + . T . + . . . . . + . . . G . . . . . . . . . . . T . . . . . . . . . . . T . . . . . . . . . . . C . . . . . . . . . . .
Then go along the first seq inserting a + wherever 2/3 bases in a moving window match. The first seq is compared to TTG (the next 3 in the vertical sequence).

A T G A T A T T C T T A . . . . . . . . . . . T . + . . + . + . . + . T . + . . . . . + . . . G . . + . . . . . + . . T . . . + . . . . . + . T . . . . . . . + . . . C . . . . . . . . . . .
Iterate until

A T G A T A T T C T T A T + + + + T + + G + + T + + T + C
The human eye is particularly good at picking up structure from the pattern of dots. You might see a hint of a duplicated region in the horizontal sequence that is not so clear from the sequence itself



Jurassic DotplotMark Boguski1st smartass

Dinosaur DNA 2
(GAT1_CHICK sw:P17678 Erythroid Transcription Factor)scoring matrix: BLOSUM50, gap penalties: -12/-295.6% identity; Global alignment score: 2144
• New seq published in Jurassic Park II
• Search database with “dinosaur” DNA
• Top hit
But alignment not perfect – gaps inserted

TPQWVPPATQMEPPHYLELLQPPRGSPPHPSSGPLLPLSSGPPPCEARECVMARKNCGAT::::::::::::::::::::::::::::::::::::::::::::::::::: ::::: TPQWVPPATQMEPPHYLELLQPPRGSPPHPSSGPLLPLSSGPPPCEARECV----NCGAT
ATPLWRRDGTGHYLCNWASACGLYHRLNGQNRPLIRPKKRLLVSKRAGTVCSHERENCQT :::::::::::::::: ::::::::::::::::::::::::::::::::: :::: ATPLWRRDGTGHYLCN---ACGLYHRLNGQNRPLIRPKKRLLVSKRAGTVCS----NCQT
STTTLWRRSPMGDPVCNNIHACGLYYKLHQVNRPLTMRKDGIQTRNRKVSSKGKKRRPPG::::::::::::::::: ::::::::::::::::::::::::::::::::::::::::STTTLWRRSPMGDPVCN---ACGLYYKLHQVNRPLTMRKDGIQTRNRKVSSKGKKRRPPG
Dinosaur Boguski Alignment
Aligning the “dinosaur” DNA (upper) with the chicken (lower)

When global fails
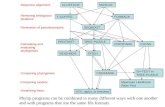
F1
E F1 EF2
E K CatalyticK
K CatalyticF12
PLAT
Two blood clotting genes Factor 12 and Plasminogen Activatorhave F, E, K and Catalytic domains typical of pathway
By aligning PLAT’s F1 domain with F12’s F2 domain, you miss a better alignment (in grey) between the two F1 domains
The alignment doesn’t recognise the second E domain in F12 but just puts a gap in the other sequence
The alignment doesn’t recognise the second K domain in PLAT but forces an alignment to the other sequence

Alignment protocol• What should real biologists do?1. Dotplot against self to identify internal repeats2. Dotplot against other sequence
• Alter windowsize and stringency3. If similarity along whole seq do global
alignment• Take default parameters• Then change parameters to check effect
4. If local/domain similarity only then do local alignment
5. If in doubt do local alignment6. LOOK at the alignment and see if you can
improve it: by hand – use good judgment

LALIGN
• Internal repeats really confuse global alignment• Local alignment reports only BEST alignment• What about sub-optimal, second best hits?• If you do a dotplot repeats will be clear• Use LALIGN to report not only the best
alignment but also any other repeated elements– And show you the aligned sequences there

2 sequence alignment
Finally, some sequences are similar even if they have no recent common ancestor. Huntington's disease is caused by repeated CAG tracks in the DNA which results in polyGlutamine (Gln, Q) tracks in the protein. If you do a homology search with QQQQQQQQQQ you get hits to other proteins that have a lot of glutamines but have totally different function.

2 sequence alignment
Huntingtin: MATLEKLMKA FESLKSFQQQ QQQQQQQQQQQQQQQQQQQQ PPPPPPPPPP PQLPQPPPQA Search against database hits:
>MM16_MOUSE MATRIX METALLOPROTEINASE-16 Score = 34.4 bits (78), Expect = 0.18 Identities = 21/65 (32%), Positives = 25/65 (38%), Gaps = 2/65 (3%):
FQQQQQQQQQQQQQQQQQQQQQQQPPPPPPPPPPPQLPQPPPQ--AQPLLPQPQPPPPPPF Q + + Q Q+ PP PPP LP PP P P+ P PPFYQYMETDNFKLPNDDLQGIQKIYGPPDKIPPPTRPLPTVPPHRSVPPADPRRHDRPKPP
But not because it is involved in microtubule mediated transport!
PRPs (proline-rich protein) have same problem