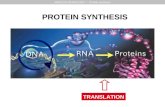
Protein Synthesis
description
Transcript of Protein Synthesis

E P A E P A
Protein Synthesis

E. coli Ribosome
-70S particle, MW ~2.5 x 106
-dissociable into small (30S) and large (50S) subunits-30S contains 16S RNA, 21 polypeptides-50S contains 5S, 23S RNA + 31 polypeptides
“Although the ribosome has been crystallized…it is such a complex entity that it will be many years before its structure is known in molecular detail” - Voet and Voet, Biochemistry 1995

Ribosome X-ray Structure
Science (2000) 289:920-30

Puromycin
“…is known to interfere with protein formation by interfering with the function of RNA in the cells involved. In goldfish studied by Bernard W. Agranoff at the University of Michigan long term memory was obliterated when the fish were given minute injections of puromycin. Since short-term memory is not much affected, it is concluded that the antibiotic interferes with the process by which memory becomes fixed in the brain” - Merck Index 1968
O
N
NN
N
CH3N
H3C
HO
NH OH
CH3O
O
NH3+
Tyrosyl-tRNA
ON
NN
N
HN
H
tRNA-O3PO
O OH
HO
O
NH3+
Puromycin Inhibition of Protein Synthesis

Science (2000) 289:920-30

Science (2000) 289:920-30

Chemistry of Peptidyl Transfer
Science (2000) 289:947-50
BB
B

Split Genes and RNA Splicing
Nobel Prize - Medicine or Physiology - 1993
P.A. Sharp (Biology, MIT)
Proc. Natl. Acad. Sci. U.S.A. (1977) 74, 3171-5
studies on genetic structure of adenovirus 2
intronexon exon

Split Gene Structure
-eukaryotes from yeast to humans-90% of human genes-intron length highly variable-exon length ~200 nt-Dystrophin

mRNA
template DNA
?
Gene Structure Analysis by EM

Discovery of Split Genes (1977)
Voet and Voet Biochemistry
5’
mRNA
DNA
3’
I
II
IIIIV
V
VI
VII1 23 45 6
7
chicken ovalbumin
123 5 764
1 2 3 5 764
I II III IV V VI VII
7700 bp
1872 nucleotides
< 20% coding

Pre-mRNA Splicing
-splicing is nuclear (HeLa nuclear extracts)-requires Mg2+
-requires ATP-(is co-transcriptional)

IVS-E2
IVS
E1-IVS-E2
E1-E2
E1
Analysis of in vitro Splicing of 32P-Labeled pre-mRNA

A PPTOH2'
A
OH
A CAG-OH
step one
step two
CAGGU
CAGPPT
PPT
UG
UG
Pre-mRNA Splicing

Chemistry of Splicing
1st step 2nd step
P
OO
OG
OHO
O
-
OG
OH
O
O
OG
OHOH
O
P
OO
OG
OHO
O
-
OG
OHOH
O
OG
OH
O
O
5’ exon
5’ exon
5’ exon
5’ exon
3’ exon
3’ exon
OG
OHO
O
P
O
OO
OG
OHO
-
OA
OH
O
O
P
OO
OG
OHO
O
-
OA
O
O
OG
OHOH
O
O

Splicing Time Course
IVS-E2
IVS
E1-IVS-E2

Complex Formation in HeLa Extract
H
A
B
C

Discovery of the Spliceosome
Cell (1985) 42, 345-53
-60S particle required for pre-mRNA splicing-spliceosome contains ribonucleoprotein particles(snRNPs - small nuclear)-U1, U2, U4/U6 U5-each snRNP contains respective snRNA (U1, U2,U4/U6 U5) + associated proteins

snRNP Composition

Protein Components of the Spliceosome
-~10-220 kD-structural roles, functional roles-conserved (core), unique-non-snRNP
“Comprehensive proteomic analysis of the human spliceosome.”Nature 419, 182-185 (2002).

Cell (1999) 96, 375-87
snRNP Core Proteins
B,D1,D2,D3,E,F,G

U1
U2
U1
U1
U4
U4 U1
U1
U1
U1
U1U1
U2
U2
U4
U4
U2
U2
U4
U2
U2
U6
U6
U6
A
U5
U5
U5
U5
U5
U5
U5
-3'
-5' A
-5'
UG
-3'A AG
-5'
A
5'- GU -3'A AG
-3'A
U4/U6
U4/U6U4/U6
-5'
B1
C1
B2
C2
I
A
CC
ATP
ATP
ATP
b p
e x o n 15 '- e x o n 2 - 3 'G U A A G
P y
5 ' S S 3 ' S S
Pre-mRNA
A
- 3 '5 '- e x o n 1 e x o n 2
mRNA
A AG-5'
-3'
U6
-3'
U6
U4
U2
U6 U5
A AG
The spliceosome cycle

GU CAGPPTUACUAAC
5’ splice site 3’ splice site
poly-pyrimidine tract
branch region
Splicing Directed by Conserved Intron Sequences

Early Steps in Spliceosome AssemblyRecognition of the Pyrimidine Tract
. .
HeLa
poly-U
guanidine1 M KClΔ HeLa 2U AF
U2AF required for A complex

U2 Auxiliary Factor
-heterodimer, 65 kD, 35 kD subunits-U2AF65 required splicing factor-U2AF35, 3’ splice site
RNA BINDEFFECTORN C
U2AF65 Domain Structure
MSDFDEFERQLNENKQERDKENRHRKR S HSR S R S RDRKRR S R S RDRRNRDQR S ASRDRRRR S K-
U2AF65
Pyr tractBranch Region

.BBP
SR
AG
SR
U2AF65
U1snRNP
A
Bridged Commitment Complex
Py
EBCU2AF35
Bridged Commitment ComplexE Complex

Selection of 5’ Splice Site and Branch
GUAUGU5' exon
CAUUCAU1
••• ••
5' splice site
5'
UACUAC 3' exon
AUGAUGU2
• • • • • 5'
A
•34
UACUAAC 3' exon
3' splice siteBBP

RNA Rearrangements in the Spliceosome-extensive U4/U6 interaction is replaced with a U2/U6 structure-U1 displaced at 5’ splice site by U6

RNA Rearrangment at 5’ Splice Site
GUAUGU5' exon
CAUUCAU1
••• ••
5' splice site
5'
GUAUGU5' exon
GACACAU6 47
• • •5'
3

Unusual Classes of Introns
AU ACA
“AT-AC”
minor spliceosome
U11 U1
U12 U2
U4atac/U6atac U4/U6
U5 common