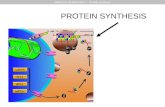
Jessica Hawley PROTEIN SYNTHESIS. Protein Synthesis Protein Synthesis PROTEIN SYNTHESIS.
Protein Synthesis
description
Transcript of Protein Synthesis


Genetic codes • A genetic code=3 consecutive nucleotides: triplet
– Why 3 bases are required to code for 1 amino acid?– Letter =4, No. of words = 20, min. length of letter =?
• Universal• redundant/ degenerate:
3 genetic codes (i.e. UAA, UAG and UGA) are meaningless Remaining 61 genetic codes represent 20 different amino acids
• Non-overlapping– Advantage?– Disadvantage?


RNA transcriptionRNA transcription
1. Structure of RNA- 3 main differences between RNA and DNA
a. Sugar in RNA is riboseribose
b. RNA is single strandedsingle stranded
c. RNA contains uraciluracil in place of thymine

3 3 Types of RNATypes of RNAa. Messenger RNAMessenger RNA (mRNAmRNA)- disposable copy of DNA to carry instructions to rest of cellb. Ribosomal RNARibosomal RNA (rRNArRNA)- helps to assemble proteins on ribosomesc. Transfer RNATransfer RNA (tRNAtRNA)- transfers amino acids to ribosomes to contruct protein molecules

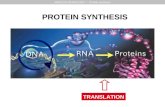
TranscriptionTranscription- process by which DNA makes complementary sequence of RNA
1. Enzyme (RNA Polymerase) separates DNA strand
2. One strand of DNA used as template to assemble strand of RNA. Takes place in nucleus
3. Transcription begins at specific locations on DNA (promoters)
http://www.twgss.edu.hk/bio/A-level/animation/transcription.mov

• After mRNA is transcribed from DNA, then the mRNA has a different fate in prokaryotes and eukaryotes
• Prokaryotes immediately begin translatingtranslating the mRNA. Eukaryotes must process it first.

Enkaryotes: mRNA Processing:• intron/exon• methyl cap• poly-A tail
Prokayotes: No mRNA Processing

• Viral DNA injected into cells. Cells evolve nucleasesnucleases in cytoplasm that chomp up any RNA or DNA out there
• Nucleases can’t get through the nuclear envelope so DNA is safe
• mRNA sent out into the cytoplasm must be protected– Methyl cap is a block– Poly A tail is a fuse
• mRNA is still chomped up into NTP’s and recycled, but the Poly A tail gives it some time

• Eukaryotic DNA is composed mostly of “non-“non-coding DNA”coding DNA” (or “junk DNA”)– We’re still not entirely sure what it does– Was probably inserted by different viruses over time
• The intronsintrons are the sections of DNA not expressed, the exonsexons are the sections that are expressed
• So now we’ve got some mRNA that codes for a protein

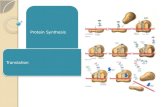
TRANSLATION

U CAU G A U UA
mRNA
CODON CODON CODON
Triplet Codon

Translation
1. mRNA attaches to ribosome. Translation begins AUGAUG, the start codon start codon

A A U
Leucine
AC U
Aspartic
Acid
AU G
Threonine
Amino acids combine with the tRNAs using the energy from the splitting of ADP.
tRNA and AMINO ACID ACTIVITION
ANTICODON

U CAU G A U UA
mRNA
CYTOLPLASM
Let’s look at another example:

U CAU G A U UA
A A U
Leucine
AC U
Aspartic
Acid
RIBOSOME
2. Complementary anticodon of the tRNA is attracted to the codon of the mRNA .

U CAU G A U UAA A U
Leucine
AC U
Aspartic
Acid
2. Complementary anticodon of the tRNA is attracted to the codon of the mRNA .

U CAU G A U UA
A A U
Leucine
AC U
Aspartic
Acid
AU G
Threonine
3. The next amino acid-tRNA attaches to the adjacent mRNA codon (Asp in this case).

U CAU G A U UA
A A U
Leucine
AC U
Aspartic
Acid
AUG
Threonine
4. Peptide bond forms between the two adjacent amino acids.

U CAU G A U UA
A A U
Leucine
AC U
Aspartic
Acid
AU G
Threonine
5. Ribosome moves along the mRNA. Another complementary tRNA is attracted to the codon. The first tRNA is released back to cytoplasm.

U CAU G A U UA
A A U
Leucine
AC U
Aspartic
Acid
AU G
Threonine
Amino acid is linked to the previous amino acids with peptide bond.

6. Polypeptide chain (protein) grows until ribosome reaches stop codonstop codon
stop codonstop codon
Protein moleculeProtein molecule

Polysome – a group of ribosomes all attached to one piece of mRNA.

Post-translational modification
Chiefly in the Golgi Apparatus
(refer to cell biology)

A classical view of negative control of the lac operon; oversimplified
Repressor is an allosteric protein:‘allos’=the other, ‘stereos’=shape

The lac Operon
• The first operon discovered• Three genes that code for the enzymes for E. coli to use lactose (galactoside) - Galactoside permease (lacY): enzyme to transport the lactose
into the cells - ß-Galactosidase (lacZ): enzyme to break the ß–
galactosidic bond - Galactoside transacetylase (lacA): its function still unclear

How was the lac operon discovered?
Monod began in 1940Inducibility of ß-galactosidase • inducibility of lactose metabolism in E. coli• an important role of ß-galactosidase in lactose metabolism• increase of the amount of ß–galactosidase upon induction
Jacob and Monod were successful in the discovery of the lac operon in combination of biochemical and genetic analysis