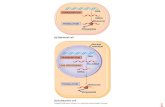
LECTURE 5 (Chapter 13) Translation of mRNA 1. INTRODUCTION The translation of the mRNA codons into...
-
Upload
catherine-willis -
Category
Documents
-
view
223 -
download
4
Transcript of LECTURE 5 (Chapter 13) Translation of mRNA 1. INTRODUCTION The translation of the mRNA codons into...

LECTURE 5
(Chapter 13)
Translation of mRNA
1

INTRODUCTION
The translation of the mRNA codons into amino acid sequences leads to the synthesis of proteins
A variety of cellular components play important roles in translation These include proteins, RNAs and small molecules
In this chapter we will discuss the current state of knowledge regarding the molecular features of mRNA translation
2

Proteins are the active participants in cell structure and function
Genes that encode polypeptides are termed structural genes These are transcribed into messenger RNA (mRNA)
The main function of the genetic material is to encode the production of cellular proteins In the correct cell, at the proper time, and in suitable
amounts
13.1 THE GENETIC BASIS FOR PROTEIN SYNTHESIS
3

4

First to propose (at the beginning of the 20th century) a relationship between genes and protein production
Garrod studied patients who had defects in their ability to metabolize certain compounds Urine chemist
He was particularly interested in alkaptonuria Patients bodies accumulate abnormal levels of
homogentisic acid (alkapton) Disease characterized by
Black urine and bluish black discoloration of cartilage and skin
Archibald Garrod
5

6

He proposed that alkaptonuria was due to a missing enzyme, namely homogentisic acid oxidase
Garrod also knew that alkaptonuria follows an autosomal recessive pattern of inheritance
He proposed that a relationship exists between the inheritance of the trait and the inheritance of a defective enzyme
Archibald Garrod
7

8
Inheritance of alkaptonuria

Metabolic pathway of phenylalanine metabolism and related genetic diseases
Figure 13.1
Dietaryprotein
CH2
NH2
Phenylalanine
Tyrosine
Phenylalaninehydroxylase
Tyrosineaminotransferase
Hydroxyphenylpyruvateoxidase
Homogentisicacid oxidase
p-hydroxyphenylpyruvicacid
Homogentisicacid
Maleylacetoaceticacid
Phenylketonuria
Tyrosinosis
Alkaptonuria
COOHC
CH2HO COOHC
H
H
NH2
Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
9

In the early 1940s, George Beadle and Edward Tatum were also interested in the relationship between genes, enzymes and traits Experiments supported Garrod’s idea that each gene =
one enzyme
Their genetic model was Neurospora crassa (a common bread mold) Their studies involved the analysis of simple nutritional
requirements
Beadle and Tatum’s Experiments
10

They analyzed more than 2,000 strains that had been irradiated to produce mutations At this point, DNA identified as probable carrier of genetic
information Does DNA somehow “code” for enzymes?
They analyzed enzyme pathways for synthesis of vitamins and amino acids
Figure 13.2 shows an example of their findings on the synthesis of the amino acid methionine
Beadle and Tatum’s Experiments
11

12

Figure 13.2
Every mutant strain was blocked at one (and only one) particular step in the synthesis pathway, showing that each gene encoded one enzyme
1
3
4
1
3
1
3
1
3
1
2
3
Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
Neurosporagrowth
WT WT WT WT WT
2
Minimal +O–acetylhomoserine +Cystathionine +Homocysteine +Methionine
(a) Growth of strains on minimal and supplemented growth media
(b) Simplified pathway for methionine biosynthesis
Homoserine O–acetylhomoserine Cystathionine Homocysteine MethionineEnzyme 1 Enzyme 2 Enzyme 3 Enzyme 4
4 2 4 2 4 2 4
13

Beadle and Tatum’s conclusion: A single “gene” in DNA controls the synthesis of a single enzyme This was referred to as the one gene–one enzyme
hypothesis
14

In later decades, this theory was progressively modified by new research 1. Enzymes are only one category of proteins 2. Some proteins are composed of two or more different
polypeptides The term polypeptide denotes structure The term protein denotes function So it is more accurate to say a structural gene encodes a
polypeptide In eukaryotes, alternative splicing means that a structural gene
can encode many different polypeptides 3. Many genes have been identified that do not encode
polypeptides For instance, functional RNA molecules (tRNA, rRNA, etc.)
15

Degenerate: (Adj) Having declined or become less specialized
Adaptor (Noun) A device that converts attributes of one device or system to those of an otherwise incompatible device or system.
Charge (Verb) To give a task to something or someone (Last slide Quiz 6, Sec 7)
16

Translation involves an interpretation of one language into another In genetics, the nucleotide language of mRNA is
translated into the amino acid language of proteins
Translation relies on the genetic code Refer to Table 13.1
The genetic information is coded within mRNA in groups of three nucleotides known as codons
The Genetic Code (first slide quiz 8, Sec 7)
17

Triplet codons correspond to a specific amino acid
Multiple codons may encode the same amino acid.
These are known as synonymous codons
Three codons do not encode an amino acid. These are read
as STOP signals for translation
18

Special codons: AUG (which specifies methionine) = start codon
This defines the reading frame for all following codons AUG specifies additional methionines within the coding sequence
UAA, UAG and UGA = termination, or stop, codons
The code is degenerate More than one codon can specify the same amino acid
For example: GGU, GGC, GGA and GGG all code for glycine In most instances, the third base is the variable base
It is sometime referred to as the wobble base
The code is nearly universal Only a few rare exceptions have been noted
Refer to Table 13.319

Figure 13.3
Figure 13.3 provides an overview of gene expression
Note that the start codon sets the reading frame for all remaining codons
5′
Template strand
Coding strand
Transcription
3′
Translation
DNA
mRNA
tRNAPolypeptide
5′ − untranslated region
3′ − untranslated region
Startcodon
Codons Anticodons
3′
3′
5′
5′
A C T G C C C A T G G G G C TC G A CA G GC G G G A A T A A C C G T C G A G G
G G C A G C T C C
C C G U C G A G G
T T GC A C
T G A C G G G T A C C C C G AG C T GT C CG C C C T T A T TA A CG T G
5′ 3′A C U G C C C A U G G G G C UC G A CA G GC G G G A A U A AU U GC A C
Met Gly LeuSer Asp Gly GluHis Leu
Stopcodon
UAC CCC GAGUCG CUG CCC CUUGUG A AC
Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
20

Sample Problem(only one answer is correct)
A tRNA has the anticodon 5’-CAU-3’. What amino acid does it carry?
a. Histidine
b. Methionine
c. Phenyalanine
d. Valine
e. None of the above
21

Polypeptide synthesis has a directionality that parallels the 5’ to 3’ orientation of mRNA
During each cycle of elongation, a peptide bond is formed between the carboxyl group of the last amino acid in the polypeptide chain and the amino group in the amino acid being added
The first amino acid has an exposed amino group Said to be N-terminal or amino terminal end
The last amino acid has an exposed carboxyl group Said to be C-terminal or carboxy terminal end
Refer to Figure 13.6
A Polypeptide Chain Has Directionality
22

Figure 13.6
(a) Attachment of an amino acid to a peptide chain
(b) Directionality in a polypeptide and mRNA
H H H H H
H3N+ H3N+
H3N+
H3N+
C C C CN C C C+
+
N
R1 R2O O
O– O–
R3 R4O
C
O
H H H H H H
Last peptide bond formed in thegrowing chain of amino acids
H O–
O–
H2OC C C CN C CN C CN
R1 R2O O R3 R4O O
H HO
H3C
Aminoterminalend
Carboxylterminalend
Methionine Serine
Peptide bonds
Sequence in mRNA
Valine
CH2
CH3
CH3
CH2
CH2
OH
CH
S
C C CN
H
O
C CN C
H O H
Cysteine
CH2
SH
CN
H
O
C
Tyrosine
CH2
OH
H
CN C
H O
H
5′ 3′A U G A G C GU U U A C U G C
Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
H
23

Figure 13.7
There are 20 amino acids that may be found in polypeptides Each contains a different side chain, or R group Each R group has its own particular chemical properties
Nonpolar amino acids are hydrophobic
They are often buried within the interior of a folded protein or in a cell membrane
H
HGlycine (Gly) G
(a) Nonpolar, aliphatic amino acids
H3N C COO–
CH3 CH3
CH
HAlanine (Ala) A
H3N COO–
CH3 CH3
CH
CH2
HValine (Val) V
H3N C COO–+
CH2CH2
CH2
HProline (Pro) P
H2N C COO–+
CH2
CH3
CH3 CH
HLeucine (Leu) L Methionine (Met) M
H3N C COO–+
Cysteine (Cys) C
+
CH2
SH
H
H3N C COO–
CH2
CH2
CH3
S
H
H3N C COO–+
HIsoleucine (Ile) I
H3N C COO–+
(b) Aromatic amino acids
Phenylalanine (Phe) F Tyrosine (Tyr) YH
H3N C COO–+
CH2
H
H3N C COO–+
CH2
OH
Tryptophan (Trp) WH
H3N C COO–+
CH2
N
H
Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
+
CH3
C+
24

Figure 13.7
Polar and charged amino acids are hydrophilic They are more likely to be on the surface of a protein
(c) Polar, neutral amino acids
Serine (Ser) S Threonine (Thr) TH
H3N C COO–+
CH2
OH
H
HCOH
H3N C
CH3
COO–+
HGlutamine (Gln) Q
H3N C COO–+
CH2
C
O NH2
HAsparagine (Asn) N
H3N C COO–+
CH2
CH2
C
O NH2
HGlutamic acid (Glu) E
H3N C COO–+
CH2
C
O O–
HAspartic acid (Asp) D
H3N C COO–+
CH2
CH2
C
O O–
(d) Polar, acidic amino acids (e) Polar, basic amino acids
Histidine (His) HH
H3N C COO–+
+
+ +
CH2
NHHN
Lysine (Lys) KH
H3N C COO–+
CH2
CH2
CH2
CH2
NH3
Arginine (Arg) RH
H3N C COO–+
CH2
CH2
CH2
C
NH
NH2
NH2
(f) Nonstandard amino acids
Selenocysteine (Sec)H
H3N C COO–+
CH2
SeH
N
CH3
Pyrrolysine (Pyl)H
H3N C COO–+
CH2
CH2
CH2
CH2
NH
C O
Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
25

There are four levels of structure in proteins 1. Primary 2. Secondary 3. Tertiary 4. Quaternary
A protein’s primary structure is its amino acid sequence Refer to Figure 13.8
Levels of Structure in Proteins
26

Lys
NH3+
110
20
30
40
5060
70
8090
100
110
120
129
ValPhe Gly Arg Cys Glu
LeuAla
AlaAla
Met
Lys
Arg
HisGlyLeuAspAsnTyrArgGlyTyr
Ser
Thr
AspTyr Gly
Leu
Asn
SerGluPheLysAlaAlaCysValTrpAsn
Leu
Gly
Phe
Asn
ThrGinAla
ThrAsnArgAsnThr
Asp
Gly
Ser
lleGln
lleAsn
SerArg Trp
Trp
Cys
Asn
AspGlyArgThrProGlySerArgAsnLeu
Cys
Asn
lle
Pro
CysSer Ala Leu
LeuSer
SerAsp
lleThr
Arg AsnArg Cys
Lys
Gly
Thr
Asp
AlaTrp ValAlaAsn
Met
GlyAsp
GlyAsp Ser Val lle Lys Lys Ala Cys
AsnVal
Ser
Ala
ValGlnAlaTrplleArgGlyCysArg
Leu
Trp
COO–
Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
Figure 13.8
The amino acid sequence of the enzyme lysozyme
129 amino acids long
Within the cell, the protein will not be found in this linear state Rather, it will adopt
a compact 3-D structure
Indeed, this folding can begin during translation
The progression from the primary structure to the 3-D structure is dictated by the amino acid sequence within the polypeptide
Copyright ©The McGraw-Hill Companies, Inc. Permission required for reproduction or display 27

The primary structure of a protein folds to form regular, repeating shapes known as secondary structures
There are two types of secondary structures helix sheet Certain amino acids are good candidates for each structure These secondary structures are stabilized by the
formation of hydrogen bonds between atoms located in the polypeptide backbone
Refer to Figure 13.9
Levels of Structures in Proteins
28

The short regions of secondary structure in a protein fold into a three-dimensional tertiary structure Refer to Figure 13.9 This is the final conformation of proteins that are composed of a
single polypeptide Structure determined by hydrophobic and ionic interactions as well as
hydrogen bonds and Van der Waals interactions
Proteins made up of two or more polypeptides have a quaternary structure This is formed when the various polypeptides associate with one
another to make a functional protein Refer to Figure 13.9
Levels of Structures in Proteins
29

Figure 13.9Copyright ©The McGraw-Hill Companies, Inc. Permission required for reproduction or display
α helix
β sheet
Primarystructure
Secondarystructure
Quaternarystructure
Tertiarystructure
Proteinsubunit
Ala C
O
C
C
C
C
O
O
Val
Phe
Glu
Tyr
Leu
Iso
Ala
H
NNH3
+
NH3+
COO–
COO–
NH3+
COO–
H
N
CC
CC O
O
HH
NN
H
N
CC
C
CC
CO
OC
OH
H
N
NN
Depending onthe amino acidsequence,some regionsmay fold intoan α helix orβ sheet.
Two or morepolypeptidesmay associatewith each other.
Regions ofsecondarystructure andirregularly shapedregions fold into athree-dimensionalconformation.
C
C
C C
O
H
H
N
NN
C
C
C
C CC
O
O
H
H
NC C C
O
NC C C
O
NC
O
HC
C
C
O
O
H
H
NCHC
C
O
H
NO
CC
HC
C
O
H
CC
O
H
C CO
H
(a)
(b)
(c)
(d)
H
COO C
HH
H
O C
Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
30

To a great extent, the characteristics of a cell depend on the types of proteins its makes
Proteins can perform a variety of functions Refer to Table 13.5
A key category of proteins are enzymes Accelerate chemical reactions within a cell Can be divided into two main categories
Anabolic enzymes Synthesize molecules and macromolecules Catabolic enzymes Break down large molecules into small ones
Important in generating cellular energy
Functions of Proteins
31

13-3932

In the 1950s, Francis Crick and Mahon Hoagland proposed the adaptor hypothesis tRNAs play a direct role in the recognition of codons in
the mRNA
In particular, the hypothesis proposed that tRNA has two functions 1. Recognizing a 3-base codon in mRNA 2. Carrying an amino acid that is specific for that codon
13.2 STRUCTURE AND FUNCTION OF tRNA
33

During mRNA-tRNA recognition, the anticodon in tRNA binds to a complementary codon in mRNA
Recognition Between tRNA and mRNA
Figure 13.10
tRNAs are named according to the
amino acid they bear
The anticodon is anti-parallel to the codon
Phenylalanine
tRNAPhetRNAPro
Phenylalanineanticodon
Phenylalaninecodon
Prolinecodon
A G
Proline
Prolineanticodon
U C
3′ mRNA5′
Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
G CA G
U C C G
34

The secondary structure of tRNAs exhibits a cloverleaf pattern It contains
Three stem-loop structures A few variable sites An acceptor stem with a 3’ single strand region
The actual three-dimensional or tertiary structure involves additional folding
In addition to the normal A, U, G and C nucleotides, tRNAs commonly contain modified nucleotides More than 80 of these can occur
tRNAs Share Common Structural Features
35

Anticodon
U
GG
C
GA
A
UH2
UH2 UH2
30
10
19
40
60
70
Acceptor stem
50
U
I C
mI
P
G
PO4
OH
UU
A
G
CPT
m2G
A
C
C
3′
5′
A
C
C
NH3+
C RC O
H
O CovalentbondbetweentRNAand anaminoacid
U
Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
Stem–loop
Structure of tRNAFigure 13.12
Found in all tRNAs
Not found in all tRNAsOther variable sites are
shown in blue as well
The modified bases are:I = inosinemI = methylinosineT = ribothymidineUH2 = dihydrouridinem2G = dimethylguanosine= pseudouridine
36

The enzymes that attach amino acids to tRNAs are known as aminoacyl-tRNA synthetases There are 20 types
One for each amino acid
Aminoacyl-tRNA synthetases catalyze a two-step reaction involving three different molecules Amino acid, tRNA and ATP
Refer to Figure 13.13
Charging of tRNAs
37

The aminoacyl-tRNA synthetases are responsible for the “second genetic code” The selection of the correct amino acid must be highly
accurate or the polypeptides may be nonfunctional Error rate is less than one in every 100,000 Sequences throughout the tRNA including but not limited
to the anticodon are used as recognition sites Modified bases may affect
translation rates recognition by aminoacyl-tRNA synthetases Codon-anticodon recognition
Charging of tRNAs
38

Figure 13.13
The amino acid is attached to the 3’ end of the tRNA by an ester bond
P
P P
P PPyrophosphate
Specificamino acid
Aminoacyl-tRNAsynthetase
A
PA
PA
3′
3′
5′
3′5′
5′
AMP
ATP An amino acid and ATP bind tothe enzyme. AMP is covalentlybound to the amino acid, andpyrophosphate is released.
The correct tRNA binds to theenzyme. The amino acidbecomes covalently attached tothe 3′ end of the tRNA. AMP isreleased.
The “charged” tRNA isreleased.
tRNA
Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
39

40
Please note that due to differing operating systems, some animations will not appear until the presentation is viewed in Presentation Mode (Slide Show view). You may see blank slides in the “Normal” or “Slide Sorter” views. All animations will appear after viewing in Presentation Mode and playing each animation. Most animations will require the latest version of the Flash Player, which is available at http://get.adobe.com/flashplayer.

As mentioned earlier, the genetic code is degenerate With the exception of serine, arginine and leucine, this
degeneracy always occurs at the codon’s third position
To explain this pattern of degeneracy, Francis Crick proposed in 1966 the wobble hypothesis In the codon-anticodon recognition process, the first two
positions pair strictly according to the A – U /G – C rule However, the third position can actually “wobble” or move
a bit Thus tolerating certain types of mismatches
tRNAs and the Wobble Rule
41

U
3′5′
5′
Wobbleposition
Nucleotide of tRNA anticodon
Third nucleotideof mRNA codon
GCAUI
xm5s2Uxm5Um
C, UGU, C, G, (A)A, U, G, (C)U, C, A
A, (G)
U, A, GA
(a) Location of wobble position
(b) Revised wobble rules
Phenylalanine
Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
3′
Umxm5U
xo5Uk2C
A A G
U U
Wobble position and base pairing rulesFigure 13.14
tRNAs that can recognize the same codon are termed isoacceptor tRNAs
Recognized very poorly
by the tRNA
5-methyl-2-thiouridine
inosine
5-methyl-2’-O-methyluridine
5-methyluridine
lysidine
2’-O-methyluridine
5-hydroxyuridine
42
You don’t need to memorize these rules

Translation occurs on the surface of a large macromolecular complex termed the ribosome
Bacterial cells have one type of ribosome Found in their cytoplasm
Eukaryotic cells have two types of ribosomes One type is found in the cytoplasm The other is found in organelles
Mitochondria ; Chloroplasts
13.3 RIBOSOME STRUCTURE AND ASSEMBLY
43

Unless otherwise noted the term eukaryotic ribosome refers to the ribosomes in the cytosol
A ribosome is composed of structures called the large and small subunits Each subunit is formed from the assembly of
Proteins rRNA
Table 13.6 presents the composition of bacterial and eukaryotic ribosomes
13.3 RIBOSOME STRUCTURE AND ASSEMBLY
44

45

During bacterial translation, the mRNA lies on the surface of the 30S subunit As a polypeptide is being synthesized, it exits through a
channel within the 50S subunit
Ribosomes contain three discrete sites Peptidyl site (P site) Aminoacyl site (A site) Exit site (E site)
Ribosomal structure is shown in Figure 13.15
Functional Sites of Ribosomes
46

Figure 13.15
(c) Model for ribosome structure
Polypeptide
30S
50S
35
tRNA
mRNA
E P A
Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
47

Translation can be viewed as occurring in three stages Initiation Elongation Termination
Refer to 13.16 for an overview of translation
13.4 STAGES OF TRANSLATION
48

mRNA
UACAnticodon
InitiatortRNA – tRNAwith firstamino acid
AUGStart codon
AUGStart codon
UAGStop codon
UAGStop codon
Completedpolypeptide
Termination
Elongation(This stepoccurs manytimes.)
Recycling of translationalcomponents
Releasefactor
Small
Large
Ribosomalsubunits
EEA
EAP
aa1
aa2
aa3
aa4
aa5
aa1
aa1
3′3′ 5′5′
3′5′
3′5′
Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
P P A
Figure 13.16
Initiator tRNA
Initiation
49

The mRNA, initiator tRNA, and ribosomal subunits associate to form an initiation complex This process requires three Initiation Factors
The initiator tRNA recognizes the start codon in mRNA In bacteria, this tRNA is designated tRNAfmet
It carries a methionine that has been covalently modified to N-formylmethionine
The start codon is AUG, but in some cases GUG or UUG In all three cases, the first amino acid is N-formylmethionine
The Translation Initiation Stage
50

Shine-Dalgarnosequence
mRNA
5′ 3′A U C U A G U A A G U U C A GG G U CG A GU C A C G C A GU G GG U A
3′
Startcodon
A U U C C C A C A GC 16S rRNAU
Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
The binding of mRNA to the 30S subunit is facilitated by a ribosomal-binding site or Shine-Dalgarno sequence
This is complementary to a sequence in the 16S rRNA
Figure 13.17 outlines the steps that occur during translational initiation in bacteria
Figure 13.18
Hydrogen bonding
Component of the 30S subunit
51

Figure 13.17
IF2, which uses GTP, promotesthe binding of the initiator tRNAto the start codon in the P site.
Portion of16S rRNA
The mRNA binds to the 30S subunit.The Shine-Dalgarno sequence iscomplementary to a portion of the16S rRNA.
IF1 and IF3 bind to the 30S subunit.
3′5′
30S subunit
Shine-Dalgarnosequence(actually 9nucleotides long)
Startcodon
IF3 IF1
IF1IF3
Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
52

Figure 13.17
70S initiation complex
This marks the end of the initiation
stage
IF1 and IF3 are released.
IF2 hydrolyzes its GTP and is released.
The 50S subunit associates.
tRNAfMet
IF2GTP
E AP
3′5′
3′5′
70Sinitiationcomplex
IF1IF3
Initiator tRNA
tRNAfMet
53

54
Please note that due to differing operating systems, some animations will not appear until the presentation is viewed in Presentation Mode (Slide Show view). You may see blank slides in the “Normal” or “Slide Sorter” views. All animations will appear after viewing in Presentation Mode and playing each animation. Most animations will require the latest version of the Flash Player, which is available at http://get.adobe.com/flashplayer.

In eukaryotes, the assembly of the initiation complex is similar to that in bacteria However, additional factors are required
Note that eukaryotic Initiation Factors are denoted eIF
Refer to Table 13.7
The initiator tRNA is designated tRNAmet It carries a methionine rather than a formylmethionine
The Translation Initiation Stage
55

The start codon for eukaryotic translation is AUG Ribosome scans from the 5’ end of mRNA until it finds
the AUG start codon (not all AUGs can act as a start) The consensus sequence for optimal start codon
recognition is show here
Start codon
G C C (A/G) C C A U G G
-6 -5 -4 -3 -2 -1 +1 +2 +3 +4
Most important positions for codon selection
These rules are called Kozak’s rules After Marilyn Kozak who first proposed them
With that in mind, the start codon for eukaryotic translation is usually the first AUG after the 5’ Cap!
56

Translational initiation in eukaryotes can be summarized as such:
An initiation factor protein complex (eIF4) binds to the 5’ cap in mRNA
These are joined by a complex consisting of the 40S subunit, tRNAmet, and other initiation factors
The entire assembly moves along the mRNA scanning for the right start codon
Once it finds this AUG, the 40S subunit binds to it The 60S subunit joins This forms the 80S initiation complex
57

During this stage, amino acids are added to the polypeptide chain, one at a time
The addition of each amino acid occurs via a series of steps outlined in Figure 13.19
This process, though complex, can occur at a remarkable rate In bacteria 15-20 amino acids per second In eukaryotes 2-6 amino acids per second
The Translation Elongation Stage
58

Figure 13.19
The 23S rRNA (a component of the large subunit) is the
actual peptidyl transferase
Thus, the ribosome is a ribozyme!
Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
3′
P site
Codon 3
Codon 4
mRNA
E siteA site
aa1
aa2
aa3Ribosome
aa1
aa2
aa3
EAP
aa4
A charged tRNA bindsto the A site. EF-Tufacilitates tRNA bindingand hydrolyzes GTP.
Peptidyltransferase, whichis a component of the 50Ssubunit, catalyzes peptidebond formation between thepolypeptide and the aminoacid in the A site.Thepolypeptide is transferredto the A site.
5′
5′
3′
59

Figure 13.19
tRNAs at the P and A sites move into
the E and P sites, respectively
Codon 4
Codon 5Codon 3
3′5′
aa1
aa2
aa3
aa4
aa1aa2
aa3
E A
A
Codon 4
Codon 5Codon 33′
5′
aa1aa2aa3
aa4
EA
P
P
aa4
This process is repeated, again andagain, until a stop codon is reached.
An unchargedtRNA is releasedfrom the E site.
The ribosome translocates1 codon to the right. Thistranslocation is promotedby EF-G, which hydrolyzesGTP.
5′3′
EP
Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
60

61
Please note that due to differing operating systems, some animations will not appear until the presentation is viewed in Presentation Mode (Slide Show view). You may see blank slides in the “Normal” or “Slide Sorter” views. All animations will appear after viewing in Presentation Mode and playing each animation. Most animations will require the latest version of the Flash Player, which is available at http://get.adobe.com/flashplayer.

The final stage occurs when a stop codon is reached in the mRNA In most species there are three stop or nonsense codons
UAG UAA UGA
These codons are not recognized by tRNAs, but by proteins called release factors
Indeed, the 3-D structure of release factors mimics that of tRNAs
The Translation Termination Stage
62

Bacteria have three release factors
RF1, which recognizes UAA and UAG RF2, which recognizes UAA and UGA RF3, which does not recognize any of the three codons
It binds GTP and helps facilitate the termination process
Eukaryotes only have one release factor eRF, which recognizes all three stop codons
The Translation Termination Stage
63

Figure 13.20
3′5′
Stop codonin A site
tRNA in Psite carriescompletedpolypeptide
E A
3′5′
E A
mRNAA release factor (RF) binds to the A site.
The polypeptide is cleaved from the tRNAin the P site. The tRNA is then released.
The ribosomal subunits, mRNA, andrelease factor dissociate.
Releasefactor
3′
+
3′
5′
5′
50S subunit 30S subunit
mRNA
Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
P
P
64

65
Please note that due to differing operating systems, some animations will not appear until the presentation is viewed in Presentation Mode (Slide Show view). You may see blank slides in the “Normal” or “Slide Sorter” views. All animations will appear after viewing in Presentation Mode and playing each animation. Most animations will require the latest version of the Flash Player, which is available at http://get.adobe.com/flashplayer.

66

Bacteria lack a nucleus Therefore, both transcription and translation occur in the cytoplasm
As soon an mRNA strand is long enough, a ribosome will attach to its 5’ end
So translation begins before transcription ends This phenomenon is termed coupling Refer to Figure 13.21
Bacterial Translation Can Begin Before Transcription Is Completed
67

Figure 13.21
Coupling between transcription and translation in bacteria
68

69