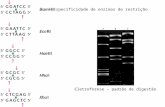
Enzimas de restriccion
-
Upload
cibergeneticaunam -
Category
Education
-
view
2.694 -
download
0
Transcript of Enzimas de restriccion

1
http://en.wikibooks.org/wiki/Structural_Biochemistry/Enzyme_Catalytic_Mechanism/Restriction_
Enzyme
Structural Biochemistry/
Enzyme Catalytic Mechanism/
Restriction Enzyme
General information
Restriction enzymes (also called restriction endonucleases) recognizes and
cleaves DNA at specific sequences. Found in a wide range of bacterial species,
these enzymes are valuable tools in the field of bioengineering as they are nature's
scapple. Restriction enzymes serve the purpose of defending bacteria from virus
where it degrades and digest foreign DNA. Restriction enzymes can also be used
for cloning DNA fragments in the field of bioinformatics through the process of
Bacterial Artificial Chromosomes (BAC) and Yeast Artificial Chromosomes (YAC).
One example of such restriction enzyme is BamHI. This process is of high
specificity and uses restriction endonucleases to degrade the viral DNA. Specific
base sequences, recognition sites, are recognized by these enzymes. These
enzymes usually recognize some specific sequence of DNA which is known
as recognition sites for the enzyme and cleave the DNA at certain positions.
There are three different types of restriction enzymes: Types I, II and III. Types I
and III are large, multi-subunit complexes containing both the endonuclease and
methylase activities. Type I restriction enzymes cleave DNA at sites that can be
1,000 base pairs away from the recognition site. Type III cleaves DNA at sites
about 25 base pairs from the recognition sites. The most commonly used type is
the type II restriction enzyme. This type of restriction enzyme cleaves the DNA at
the site of recognition.

2
Each restriction enzyme cleaves DNA in a specific way. Some cuts the two DNA
strands asymmetrically, leaving unpaired bases at each end. This type of unpaired
strands of DNA is called a sticky end. Sticky ends of different but complementary
DNA strands can ligate to form a recombinant DNA. Restriction enzymes that
cleave directly at the opposing phosphodiester bond, leaving no unpaired bases on
the ends, are called blunt ends.
The most widely studied restriction endonuclease is called "BamHI." This
restriction endonuclease provides an excellent example of how most
endonucleases behave in order to perform their assigned tasks. Like many other
restriction endonucleases, BamHI binds to two divalent metals (such as Ca2+ or
Mg2+) to form a coordination compound that will increase its binding ability to DNA.
However, instead of "searching" the entire DNA double-helix for the specific
sequence it requires, BamHI uses a cellular shortcut: it latches onto the DNA
molecule at any convenient point, and then begins to slide along the DNA
backbone until it finds the sequence of interest. This behavior is not unique to
BamHI; restriction endonucleases routinely perform such "tricks" in order to do
their jobs more efficiently. This tactic is quite effective: instead of having to sort
through the messy twists and turns of an entire DNA double-helix, the restriction
endonucleases only need to slide along one strand until they reach their target
sequence. They effectively turn a 3-D problem (searching through the bends and
folds of the DNA molecule) into a 2-D problem (sliding one way along a DNA
strand). For instance, restriction enzymes may choose to methylated nucleotide
sequences that are not intended to be displaced through the addition of a methyl
group. This methylation marks the DNA sequence that will not be cut out from the
original sequence which could disrupt the whole cell. These methylated nucleotides
are formed as tags for the restriction enzymes to identify.
Mechanisms for Cleavage by In-Line Displacement
Restriction enzymes catalyze the hydrolysis of the of the phosphodiester bonds in
the backbone of DNA. It specifically hydrolyzes the bond between 3’ oxygen and
phosphorus to cause the bond to break. Resulting products are a DNA strand that

3
has a free 3’ hydroxyl group and another strand with a 5’ phosphoryl group. There
are two methods that restriction enzymes can use to cleave the phosphodiester
bond in the DNA back bone. One way is to form covalent intermediates that can do
nucleophilic attacks to the Phosphate group at the phosphodiester linkages. The
other way is to simply hydrolyze the complex by adding water. The two methods
are shown below.
Mechanism type 1 (Covalent intermediate)
Nucleophile attacks the phosphoryl group and forms an intermediate. Then the
intermediate is hydrolyzed and produces two separate DNA strands.
Stereochemistry is retained in this mechanism - in the first step it is inverted but
also in the second step. Therefore, it retains the original stereo chemistry of the
phosphorous atom.
Mechanism type 2 (Direct Hydrolysis)
An activated water molecule attacks the phosphorus atom directly. There is no
intermediate step in this kind of mechanism, such reactions are known as
concerted reactions. With this kind of mechanism, the stereochemistry of molecule
is inverted. This mechanism is almost identical to the second order nucleophilic

4
substitution (SN2) reaction. The attack of water on the phosphorus atom is driven
by the fact that the electronegative oxygen, equipped with 2 lone pairs of electrons,
is attracted to the phosphorus which is relatively electropositive because much of
its electron density is drawn away by the oxygen containing groups attached to it.
The reason the reverse reaction (the R1OH alcohol product attacking the
phosphorus and kicking off the water) does not take place is first, the alcohol is a
bit more bulky and will thus encounter more steric hindrance from the groups
attached to the phosphorus and second, and more importantly, this reaction
usually takes place in an aqueous solvent where the is a lot of water. Thus,
according to LeChatlier's Principle, the excess of one of the reactant of the reaction
shown below will push the equilibrium towards the products.
Magnesium
Enzymes acting on phosphate containing substrates usually require divalent
cations such as magnesium for activity. There are several ways to determine the
presence of metal ions. One way is to prepare crystals of EcoRV bound to
oligonucleotides that contain the enzyme's recognition sequence without the
magnesium to prevent cleavage. After the crystals have been formed, they are
soaked in a metal ion solution. Another way to prepare the crystals is to use a
mutated form of an enzyme that is less active. In this way, no cleavage takes place
so binding sites can be determined since it is impossible to view the complex in
EcoRV and cognated DNA molecules with magnesium present as the enzymes
cleaves the substrate under this condition. A maximum of three metal ions can be
found in each active site. The metal ions are coordinated to the protein through two
aspartate residues and to one of the phosphoryl group oxygen atoms near the
cleavage site which helps water attack the phosphorous atom like the zinc in
carbonic anhydrase.

5
Specificity
Most restriction enzymes have recognition sequences that are inverted repeats
that gives a 3D structure of the recognition site a twofold rotational symmetry. The
symmetry between the recognition sequence and the enzyme facilitates the
recognition of cognate DNA. The binding affinity for substrates can help to
determine specificity as well. The enzyme only cleaves cognate sequences
because within the GATATC sequence, the G and A bases at the end of the 5 end
of each strand and their base partners are in direct contact with the enzyme
through hydrogen bonds with residues located on the loops. This sequence has a
kink in the center due to the TA base pairs at the center allowing for the distortion
of DNA. Noncognate sequences, on the other hand, are not distorted enough.
Thus, the lack of distortion means that there is no phosphate positioned close
enough to the active site aspartate residues to complete a magnesium binding site
so nonspecific complexes are unable to bind magnesium ions and complete the
catalytic apparatus. Distortion of the substrate and magnesium binding are the
main factors that contribute to catalytic specificity. Distorted DNA increases binding
energy since it makes additional contacts with the enzyme but it is cancelled out by
the energy it takes to distort the DNA. In EcoRV this causes a slight difference in
binding affinity between cognate and nonspecific DNA but it cognate complexes, it
affects catalysis through the completion of magnesium binding. Distortion also
helps protect host-cell DNA by blocking catalysis through methylation since the
presence of methyl groups block the formation of hydrogen bonds between the
amino group of an adenine nucleotide of the recognition site and the side chain of
the asparagine residue that is closely linked to the DNA.
Restriction enzymes bind indiscriminately to any DNA sequence but only specific
non-methylated cognate sequences are cleaved.
Take EcoRV endonuclease for example,
the enzyme recognises the sequence
5'–GATATC–3'
3'–CTATAG–5'
and cuts specifically at the center.

6
This recognition site has special features: It is made of inverted repeats, known as
palindromic sequences; and it is symmetric around axis of rotation, giving a 2-fold
rotational symmetry.
The dimeric enzyme matches this 2-fold symmetry of the substrate DNA and
surrounds the DNA in tight embrace. On binding with the DNA, the enzyme forms
hydrogen-bonds to the G and A bases at the 5' end of each strand and their
corresponding Watson-Crick partners. The central TA base pairs do not form
hydrogen bonds but are essential to the distortion of the DNA recognition site as
they are most easily deformed.
The Eco RI restriction enzyme
An example of post-reactive cognate DNA-Eco RI complex at 2.5A in the presence
of Mn2+ ion.

7
Although the additional interactions made by the distorted cognate DNA with
EcoRV have led to an increase in its binding energy, this additional energy is
canceled as the DNA distorts itself from the relaxed conformation. Therefore, both
cognate and non-cognate complexes have about the same free energy in the end,
accounting for the little difference in their binding affinity. In the non-cognate
complex, however, DNA phosphates are too far away from the Asp residues to
make up a Mg-binding site. Thus, the DNA will be bound but not cleaved.
BamHI
BamHI is the most studied restriction endonuclease. It is derived from Bacillus
amyloliquefaciens. G'GATCC is its recognition site, and leaves a sticky end. It's a
dimer and each monomer binds to its recognition site. Persistent issues with this
specific enzyme spontaneously failing to generate cloneable fragments necessitate
aliquoting prior to use. Questions such as 'How does BamHI cleave DNA? How
can BamHI find its specific site among the thousands of nonspecific sites? Dr.
Hector co-crystallied the complex BamHI DNA with divalent cations(missing some
e's) such as Ca2+ and Mn2+. Ca2+ binds to active site but reaction does not carry

8
out catalysis. It's catalytic mechanism consist of pre-reactive state, transition state,
post-reactive state. Dr. Hector also crystallized BamHI with a non-specific DNA.
DNA is far from active site of enzyme. Restriction endonuclease first binds to any
DNA and until find right side they cleave it. Three main thermodynamic properties
of a protein/non-specific DNA complex: 1. More hydrated at teh protein DNA
interface. 2. Stabilized primarily by electrostatic interactions. 3. Experience low-
heat capacity change upon binding.
Mechanism: The mechanism that BamHI uses is essentially the Sn2 mechanism.
The amino acid E113 activates a water molecule by drawing a proton away from it
and making the water oxygen atom more like a hydroxide oxygen atom. The
activated water attacks the phosphorous atom of the phosphodiester bond that is
to be broken. The transition state that forms is trigonal bipyrimadal and carries an
additional negative charge that is stabilized by the two divalent metal ions. The
leaving group is the alkoxide attached to the C3' carbon. A second water molecule
donates a proton to neutralize the alkoxide and form the free 3' end of the DNA
strand. The proton that the first water molecule lost to E113 can be proton shuttled
to the hydroxide ion formed when the second water molecule donated its proton
and the enzyme is back to where it started. The final result is a strand with a free 3'
end and a phosphorylated 5' end.
Type II Restriction Enzymes
Type II restriction enzymes have a structure containing beta strands with aspartate
residues forming the magnesium binding sites present in Archaea and Bacteria.
The sequences show that bacteria most likely obtained the genes encoding the
enzymes from other species through horizontal gene transfer which is the passing
of DNA fragments that provide selective advantage in a certain environment
between different species.