The Molecular Basis of Life Chapter 16. Historical Note T.H. Morgan showed that genes are located on...
-
Upload
noreen-harrison -
Category
Documents
-
view
216 -
download
0
Transcript of The Molecular Basis of Life Chapter 16. Historical Note T.H. Morgan showed that genes are located on...

The Molecular Basis of Life
Chapter 16

Historical Note
T.H. Morgan showed that genes are located on chromosomes
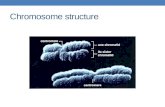
Chromosomes are made of DNA and Protein
Scientists thought proteins were the genetic material DNA was too simple.
It took half a century to disprove this.

Genes are made of DNA
The following scientists proved that
DNA is the genetic material Fredrick Griffith (1928) Oswald Avery ( 1944) Hershey and Chase (1952)

Fredrick Griffith (1928) The discovery of the genetic role of DNA
began with research by Frederick Griffith. Griffith was an English military scientist
(microbiologist) hired to find the #1 cause of death in WWI (pneumonia)
He studied Streptococcus pneumoniae, a bacterium that causes pneumonia in mammals. One strain, the R strain, was harmless. The other strain, the S strain, was pathogenic.
He used mice – fits the human model

Griffith’s Experiment

When Griffith mixed heat-killed S strain with live R strain bacteria and injected this into a mouse it died.
He recovered the pathogenic strain from the dead mouse’s blood.
Some harmless bacteria had been “transformed” into the deadly strain
He called this phenomenon transformation, a change in genotype and phenotype due to the assimilation of a foreign substance (now known to be DNA) by a cell
For the next 14 years scientists tried to identify the transforming substance.

Transforming Substance? Could be:
DNAProteins
Because scientists already knew chromosomes consist of these substances. So the debate started.

Oswald Avery ( 1944) Treated Griffiths mixture with
Protein digesting enzymes remove all proteins
DNA digesting enzymes remove all DNA.
Still, many biologists were skeptical. In part, this reflected a belief that the genes of
bacteria could not be similar in composition and function to those of more complex organisms

Avery contd. Is Protein the transforming factor?
treated Griffith’s mixture of heat treated deadly strain and live harmless strains with protein-destroying enzymes grew the strains
The bacterial colonies were still transformed
Concluded that protein could not be the transforming factor

Avery contd. Is DNA the transforming factor?
treated the mixture with DNA-destroying enzymes grew the strains
The bacterial colonies failed to transform
Concluded that DNA is the genetic material of the cell
Finally in 1944, Oswald Avery, Maclyn McCarty and Colin MacLeod announced that the transforming substance was DNA

The Debate Continues
Scientists were still skeptical proteins made of 20 AAs, DNA only 4 bases
Major reason was so little was known about DNA

Hershey and Chase (1952) Used viruses to prove that DNA is the genetic material.Viruses consist of a DNA (sometimes
RNA) enclosed by a protective coat of protein.
To replicate, a virus infects a host cell and takes over the cell’s metabolic machinery.
Viruses that specifically attack bacteria are called bacteriophages or just phages.


Viruses Infecting a Bacterial Cell
Colorized TEM


Conclusion
Phage DNA entered the bacterial cell, proteins did not
DNA carries the genetic information. Published his work in 1953 In 1969 Delbruck, Luria, Hershey
won the Nobel Prize

Additional Evidence
Came from a biochemist Erwin Chargaff (1947)
The composition of DNA was known

Building Blocks of DNANucleotides A ring-shaped sugar called
deoxyribose A phosphate group (a
phosphorus atom surrounded by four oxygen atoms)
A nitrogenous base ("nitrogen-containing") : a single or double ring of carbon and nitrogen atoms with functional groups

Nitrogenous Bases
The four nucleotides in DNA differ only in their nitrogenous bases
Bases:Thymine (T) single ringCytosine (C) single ringAdenine (A) double ringGuanine (G) double ring

Bases

Base Pairing Chargaff’s Rule Adenine forms a base pair with Thymine
(%T = %A). Guanine forms a base pair with Cytosine
(%G = %C). Chargaff noted that the DNA composition varies
from species to species. In any one species, the four bases are found in
characteristic, but not necessarily equal, ratios Human DNA is 30.9% adenine, 29.4% thymine,
19.9% guanine and 19.8% cytosine.


The Double Helix

Building a Structural Model of DNA By the beginning of the 1950’s, the
race was on to build a three-dimensional structure of DNA.
Scientists working on the problem were Linus Pauling, in California, and Maurice Wilkins and Rosalind Franklin, in London.

Maurice Wilkins Rosalind Franklin
Used X-ray crystallography to study the structure of DNA
In this technique, X-rays are diffracted as they passed through aligned fibers of purified DNA.
The diffraction pattern can be used to deduce the three-dimensional shape of molecules

James Watson and Francis Crick
Learned from their research that DNA was helical in shape and he deduced the width of the helix and the spacing of bases
Watson and Crick began to work on a model of DNA with two strands, the double helix.
Using molecular models made of wire, they first tried to place the sugar-phosphate chains on the inside.
However, this did not fit the X-ray measurements and other information on the chemistry of DNA

Breakthrough!! The key breakthrough came when
Watson put the sugar-phosphate chain on the outside and the nitrogen bases on the inside of the double helix.The sugar-phosphate chains of each strand
are like the side ropes of a rope ladder.Pairs of nitrogen bases, one from each
strand, form rungs.The ladder forms a twist every ten bases.

Copyright © 2002 Pearson Education, Inc., publishing as Benjamin Cummings
Fig. 16.5

The nitrogenous bases are paired in specific combinations: adenine with thymine and guanine with cytosine.
Pairing like nucleotides did not fit the uniform diameter indicated by the X-ray data. A purine-purine pair would be too wide and a
pyrimidine-pyrimidine pairing would be too short. Only a pyrimidine-
purine pairing would produce the 2-nm diameter indicated by the X-ray data

Watson and Crick determined that chemical side groups off the nitrogen bases would form hydrogen bonds, connecting the two strands.
Based on details of their structure, adenine would form two hydrogen bonds only with thymine and guanine would form three hydrogen bonds only with cytosine.
This finding explained Chargaff’s rules.

The base-pairing rules dictate the combinations of nitrogenous bases that form the “rungs” of DNA.
In April 1953, Watson and Crick published a short, one-page paper in Nature reporting their double helix model of DNA.
Their structure of DNA suggested a possible mechanism of replication

DNA Replication and Repair Watson and Crick published their
hypothesis for how DNA replicates.Essentially, because each strand is
complementary to each other, each can form a template when separated.
The order of bases on one strand can be used to add in complementary bases and therefore duplicate the pairs of bases exactly.

The Basic Principle: Base Pairing to a Template Strand
Since the two strands of DNA are complementary Each strand acts as a template for
building a new strand in replication

DNA Replication The parent molecule unwinds, and
two new daughter strands are built based on base-pairing rules

DNA replication semiconservative
Each of the two new daughter molecules will have one old strand, derived from the parent molecule, and one newly made strand

Meselson and Stahl
Supported the semiconservative model of DNA replication

Mechanism of Replication More than a dozen enzymes and other
proteins participate in DNA replication The replication of a DNA molecule begins
at special sites, origins of replication. In bacteria a single specific sequence of
nucleotides that is recognized by the replication enzymes. These enzymes separate the strands, forming
a replication “bubble”. Replication proceeds in both directions until
the entire molecule is copied.

Eukaryotes There may be hundreds or thousands of origin
sites per chromosome. At the origin sites, the DNA strands separate
forming a replication “bubble” with replication forks at each end.
The replication bubbles elongate as the DNA is replicated and eventually fuse.


Players Primase adds the primer molecule to
DNA to initiate DNA replication DNA polymerases adds nucleotides to the
DNA molecule DNA ligases joins the ends of nucleotides A helicase untwists and separates the
template DNA strands at the replication fork.
Single-strand binding proteins keep the unpaired template strands apart during replication

Priming DNA Synthesis
DNA polymerases cannot initiate the synthesis of a polynucleotide They can only add nucleotides to the 3
end The initial nucleotide strand
Is an RNA or DNA primer

Priming DNA Synthesis To start a new chain requires a
primer, a short segment of RNA.The primer is about 10 nucleotides long
in eukaryotes. Primase, an RNA polymerase, links
ribonucleotides that are complementary to the DNA template into the primer.RNA polymerases can start an RNA
chain from a single template strand

After formation of the primer, DNA polymerases can add deoxyribonucleotides to the 3’ end of the ribonucleotide chain
Another DNA polymerase later replaces the primer ribonucleotides with deoxyribonucleotides complimentary to the template

Elongating a New DNA Strand DNA polymerases catalyze the
elongation of new DNA at a replication fork. As nucleotides align with complementary
bases along the template strand, they are added to the growing end of the new strand by the polymerase. The rate of elongation is about 500 nucleotides
per second in bacteria and 50 per second in human cells.
The raw nucleotides are nucleoside triphosphates. Each has a nitrogen base, deoxyribose, and a
triphosphate tail.

As each nucleotide is added, the last two phosphate groups are hydrolyzed to form pyrophosphate.The exergonic
hydrolysis of pyrophosphate to two inorganic phosphate molecules drives the polymerization of the nucleotide to the new strand

Strands in the double helix are antiparallel
The sugar-phosphate backbones run in opposite directions. Each DNA strand has a 3’
end with a free hydroxyl group attached to deoxyribose and a 5’ end with a free phosphate group attached to deoxyribose.
The 5’ -> 3’ direction of one strand runs counter to the 3’ -> 5’ direction of the other strand

Antiparallel Elongation DNA polymerases add nucleotides to the
free 3’ end of a growing DNA strand. A new DNA strand can only elongate in the
5’ ->3’ direction This creates a problem at the replication
fork because one parental strand is oriented 3’->5’ into the fork, while the other antiparallel parental strand is oriented 5’->3’ into the fork.
At the replication fork, one parental strand (3’-> 5’ into the fork), the leading strand, can be used by polymerases as a template for a continuous complimentary strand

To elongate the other new strand of DNA, the lagging strand DNA polymerase III must work in the direction
away from the replication fork The lagging strand
Is synthesized as a series of segments called Okazaki fragments, each about 100-200 nucleotides, are then joined together by DNA ligase
Only one primer is needed for synthesis of the leading strand But for synthesis of the lagging strand, each
Okazaki fragment must be primed separately



To summarize, at the replication fork, the leading stand is copied continuously into the fork from a single primer.
The lagging strand is copied away from the fork in short segments, each requiring a new primer

Other Proteins That Assist DNA Replication
Table 16.1

Proofreading and Repairing DNA
DNA polymerases proofread newly made DNA Replacing any incorrect nucleotides The final error rate is only one per billion
nucleotides In mismatch repair of DNA
Repair enzymes correct errors in base pairing Each cell continually monitors and repairs its
genetic material, with over 130 repair enzymes identified in humans

In mismatch repair, special enzymes fix incorrectly paired nucleotides. A hereditary defect in
one of these enzymesis associated with a form of colon cancer.
In nucleotide excision repair, a nuclease cuts out a segment of a damaged strand. The gap is filled in by
DNA polymerase

Replicating the Ends of DNA Molecules
The ends of eukaryotic chromosomal DNA Get shorter with each round of
replication Eukaryotic chromosomal DNA
molecules Have at their ends nucleotide sequences,
called telomeres, that postpone the erosion of genes near the ends of DNA molecules

Figure 16.18
End of parentalDNA strands
Leading strandLagging strand
Last fragment Previous fragment
RNA primer
Lagging strand
Removal of primers andreplacement with DNAwhere a 3 end is available
Primer removed butcannot be replacedwith DNA becauseno 3 end available
for DNA polymerase
Second roundof replication
New leading strand
New lagging strand 5
Further roundsof replication
Shorter and shorterdaughter molecules
5
3
5
3
5
3
5
3
3

Figure 16.19 1 µm

Telomeres and Telomerases
Telomeres protect genes from being eroded through multiple rounds of DNA replication In human telomeres, this sequence is
typically TTAGGG, repeated between 100 and 1,000 times
Eukaryotic cells have evolved a mechanism to restore shortened telomeres.

Telomerase uses a short molecule of RNA as a template to extend the 3’ end of the telomere.
There is now room for primase and DNA polymerase to extend the 5’ end.
It does not repair the 3’-end “overhang,”but it does lengthenthe telomere

Telomeres and Telomerases Telomerase is not present in most cells of
multicellular organisms. DNA of dividing somatic cells and cultured
cells tends to become shorter. Telomere length may be a limiting factor in
the life span of certain tissues and the organism.
Telomerase is present in germ-line cells, ensuring that zygotes have long telomeres.
Active telomerase is also found in cancerous somatic cells. This overcomes the progressive shortening that
would eventually lead to self-destruction of the cancer