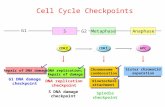
Spindle Assembly Checkpoint And Chromosomal Instability
Transcript of Spindle Assembly Checkpoint And Chromosomal Instability
Spindle Assembly Checkpoint And Chromosomal Instability
Ricardo João Silva e Sousa Mestrado de Bioquímica 2012 Orientador Sara Morais da Silva, PhD
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
ii
Acknowledgments/Agradecimentos
Ao Professor Claudio Sunkel por me ter conferido a oportunidade de desenvolver a
minha tese no seu laboratório e pela preocupação demonstrada a cada passo do seu
desenvolvimento, um muito obrigado.
À Sara Morais da Silva um grande obrigado pela orientação, incentivo e principalmente
por me ter ajudado a desenvolver como investigador dando me os instrumentos para
os próximos passos e como pessoa aumentando o meu sentido de responsabilidade.
Obrigado também pela paciência demonstrada a cada passo menos conseguido por
minha parte. Por fim um obrigado por seres a única pessoa que também gosta de
misturar o português e inglês na mesma frase. Muito obrigado.
Agradeço a todos os restantes membros do GMM pela ajuda prestada, e pela
disponibilidade demonstrada.
Agradeço a quem me aturou diariamente na “sala das moscas”, nomeadamente ao
grupo de Biologia do Desenvolvimento com um especial agradecimento à Sara que
embora não tenha “apostado” em mim (ler com rancor a parte anterior) ainda me ouviu
diversas vezes, quando nem tudo correu pelo melhor, e me deu conselhos valiosos.
Quero também agradecer à Ana Cunha pela falta de “tento na língua” que gerou bons
momentos de diversão durante as pausas de trabalho. Muito obrigado a todos.
Aos amigos e colegas de Mestrado que fizeram esta viagem comigo, um grande
obrigado pelo apoio demonstrado.
Aos meus ex-companheiros de casa Leo e Stéphane que mesmo longe souberam
sempre guiar me no melhor sentido e ouvir me sempre que necessitei.
À Ana Lúcia, Pedro Albuquerque e Sofia Pinho pelas horas intermináveis de apoio e
conselhos e principalmente por terem sido o que eu considero, mentores não só para
a minha carreira científica mas também fora dela. “Miau miau” Um grande obrigado.
Ao André Cunha por constantemente congeminar nos bastidores para eu não me
distrair e escrever (sim eu “topava-te”). Obrigado “sócio”.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
iii
Ao Sérgio Dias, o verdadeiro companheiro de escrita, que fez esta jornada comigo.
Um grande grande obrigado por ouvires e aconselhares e pelo característico humor
seco que tornavam horas de trabalho em.. bem, horas de trabalho na mesma porém
divertidas. “Estão duas gaivotas...” Um obrigado “a tender para o infinito” Sérgio.
Ao Paulo Almeida que poderia não ter apoiado minimamente durante esta tese e
estaria aqui mencionado na mesma. Não teria páginas suficiente para agradecer tudo
o que quereria mas quero apenas mencionar uma mítica frase dita alguns anos atrás
“por ti eu falt…” … “Palo” obrigado por esta amizade ser “para bom entendedor nem
meia palavra é necessária” Um grande grande obrigado.
À Rita Nobre que mais um bocado e teria que lhe dedicar esta tese. Um muito
obrigado não chega para compensar todo o trabalho, tempo, horas de sono perdidas e
paciência que tiveste durante a escrita desta tese. “Rita Nobre tem sempre….” Rita
Nobre…Obrigado… (“B.T.P.M.T.A.T.P.W.”)
À Cátia Mendes que acompanhou desde o início todas as peripécias, problemas, altos
e baixos desta tese. Da tua forma extremamente peculiar tiveste sempre a perícia de
me por no caminho certo. O teu apoio foi “para lá” de fundamental. Podia dissertar por
páginas intermináveis mas basta me agradecer por poder explicar 99% desta amizade
com “é a Cátia”. Cátia Mendes, obrigado…
À minha irmã pela paciência interminável. Sou suspeito eu sei mas dificilmente
conseguiria uma melhor. Já agradeci a paciência interminável? Mana muito obrigado.
À minha família que movem mundos e fundos para que eu seja bem sucedido. Queria
fazer um agradecimento especial aos meus avós. Cada um com seu ensinamento
ajudo-me a tornar a pessoa que sou hoje. Obrigado por me ensinarem como
descontrair e despreocupar quando necessário, o valor da inteligência e da cultura, o
valor da paciência e da diplomacia e o valor da família. Muito, muito obrigado.
Aos meus pais que por mais que eu me proteste é impossível, literalmente, alguém
fazer mais por mim do que vocês fizeram e continuam a fazer. Vocês são a base de
tudo o que eu me tornei e não consigo agradecer o suficiente. Mãe, Pai, “Velhote e
velhota”, obrigado.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
iv
Abstract
The spindle assembly checkpoint (SAC) mechanism works through the
inhibition of APC/C (anaphase promoting complex/cyclosome) in order to delay in the
metaphase-anaphase transition. This mechanism is initiated in the presence of
unattached, unaligned and/or missattached chromosomes and therefore can prevent
abnormal chromosome segregation. Severe chromosome missegregation results in
aneuploidy and when this is very extensive cell death ensues. Given that chromosome
instability and aneuploidy have been closely associated with tumorigenesis, we
decided to analyze what happens to cells that do not have the SAC but at the same
time cannot activate apoptosis. Previous work in the lab using Drosophila had shown
that in these conditions, cells show all the signs of tumorigenesis including
hyperproliferation and chromosome instability. The purpose of this work was to apply
high throughput microarray technology, to determine the transcriptional profile of cells
in which the SAC was abolished and apoptosis was prevented. For this, we used wing
imaginal discs where a SAC gene (Bub3) expression was knocked down through RNA
interference and apoptosis is inhibited by the expression of the baculovirus protein p35.
The transcriptional profile of these cells was then compared to that of cells in which
only bub3 is knocked down, where we only inhibited apoptosis through p35 or in wild
type wing imaginal discs expressing only green fluorescent protein. As a result we
identified more that 200 genes that where specifically up or down regulated and
initiated a RNAi genetic screen using flies where Bub3 was knocked down and the
apoptosis blocked only in the wing imaginal discs. The preliminary results suggest that
the screen should be able to identify candidates that actively contribute for the
tumorigenic phenotype of cells in which the SAC is inactivated and apoptosis is at the
same time inhibited.
Keywords: SAC, apoptosis, tumorigenesis, Bub3, Drosophila, microarrays,
RNAi, screen.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
v
Resumo
Através da inibição do APC/C (complexo promotor da anafase) o mecanismo
do “spindle assembly checkpoint” (SAC) consegue provocar um atraso na transição
metafase-anafase no caso da existência de cromossomas incorretamente ligados a
microtúbulos prevenindo assim a segregação anormal. A ocorrência da separação
incorreta dos cromossomas provoca aneuploidia e consequente morte celular. Uma
vez que tumorigénese se encontra associado com instabilidade cromossómica e
aneuploidia decidimos analisar o que acontece a células que não possuem SAC mas
que ao mesmo tempo não conseguem ativar morte celular. Experiências prévias
demonstraram que em Drosophila as condições mencionadas levam a tumorigénese e
instabilidade cromossómica. O objetivo do nosso trabalho foi aplicar tecnologia de
microarrays para determinar o perfil trancricional de células em que SAC e apoptose
foram inibidos. Para tal, utilizamos discos imaginais da asa onde silenciamos através
de RNA de interferência Bub3 (um gene do SAC) e inibimos apoptose com P35 uma
proteína de um baclovirus. Este perfil trancricional foi comparado com outros perfis de
diferentes genótipos, nomeadamente, quando apenas silenciávamos Bub3, quando
apenas inibíamos apoptose e com o fenótipo selvagem exprimindo apenas green
fluorescente protein. Como resultado identificamos mais de 200 genes que se
encontram sobreexpressos ou subexpressos que utilizamos para efetuar um screen
genético com RNA de interferência usando moscas onde silenciamos a expressão e
inibimos a apoptose em discos imaginais da asa. Resultados preliminares sugerem a
identificação de candidatos que contribuem ativamente para o fenótipo tumorigénico
de células onde SAC se encontra inativado e apoptose inibida.
Palavras-chave: SAC, apoptose, tumorigénese, Bub3, Drosofila, microarrays,
RNAi, screen
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
1
Index
Acknowledgments/Agradecimentos………………………………….……………… ii
Abstract……………………………………………………………………………….…… iv
Resumo……………………………………………………………………………………. v
Index……………………………………………………..………………………………… 1
Abbreviations List…………………………………………………………..…………….. 4
Chapter 1 - Introduction…………………………………………..……………………… 6
1.1. Model Organism……………………………………...……………………… 7
1.2. Tumorigenesis……..………………………………………………………… 8
1.3. Mitosis and Spindle Assembly Checkpoint (SAC) …………..…..………. 11
1.4. Apoptosis……………………...……………………………………………… 16
1.5. Main Technologies………...………………………………………………… 21
1.5.1.Microarrays………………..……………………………………………… 21
1.5.2. RNA Intereference…………….………………………………………… 23
1.6. Aims…………………………………………………………………………… 24
Chapter 2 - Materials and Methods……………………………………....…………….. 25
2.1. Fly stocks used……………………………………..…………………………… 26
2.2. RNA Extraction from Wing Imaginal Discs……………………………….….. 26
2.2.1.Collection of Wing Imaginal ……………………………………………. 26
2.2.3.RNA Extraction…………………………………………………………… 26
2.3. Microarrays……..……………………………………………………………….. 28
Chapter 3 - Results………………………..……………………………………………… 33
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
2
3.1. Introduction………………………………………………………………….…… 34
3.2. Selected Genotypes………………………………………….………………… 35
3.3. RNA Samples for Microarrays…………………………………………………. 37
3.3.1.Qualitative and Quantitative Analysis of the Extracted RNA..………. 37
3.3.1.1.Nanodrop Analysis………………..………………………….…… 37
3.3.1.2.Bioanalyzer Analysis……………………………………………… 38
3.4. Microarray Analysis…………………..………………………………………… 44
3.4.1. Microarray Data………………………………….………………………. 44
3.4.2.Mining the Microarray Data to Achieve Relevant Biological
Information………………………………………………………………………. 48
3.4.3.Validating Venn Diagrams through Bioinformatics Resources…….... 52
3.4.4.Mining the Microarray Data from Raw to Biologically Relevant.... 56
3.5.Screen for Enhancer/Suppression of Tumor Phenotype……………………. 58
Chapter 4 - Discussion……………………………………………...……………………. 65
4.1. Introduction………………………………………………………………………. 66
4.2. Reasons for Phenotype Selection…………………….………………………. 66
4.3. Microarray Comparison Reveals Significant Differences in Expression
Profiles Depending on the Genotypes…..…………………………..…………….. 67
4.4.Bioinformatic Methods Show Enrichment of Relevant Pathways from
Venn Diagram Selection………………………..…………………………………… 68
4.5.Legitimacy of Manual Selection from Venn Diagram………………………... 70
4.6.Bub3 has an Additional Role as a Tumor Suppressor………………………. 71
4.6.1.Genes Upregulated in Bub3rnai background Screen…………....…….. 71
4.6.2.Genes Downregulated in Bub3rnai background Screen……….……… 73
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
3
4.6.3.Genes Upregulated in Bub3rnai DIAP1 background Screen…….…… 75
4.6.4.Genes Upregulated in Bub3rnai DIAP1 background Screen….……… 80
Chapter 5 - References…………………………………..………………………………. 83
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
4
Abbreviations list
AcCoAS - Acetyl Coenzyme A synthase
alphaTub85E - alpha-Tubulin at 85E
Alr - Augmenter of liver regeneration
APC/C - Anaphase promoting complex/ cyclosome
Aplip1 - APP-like protein interacting protein 1
asf1 - anti-silencing factor 1
B-H1 - BarH1
Btl - breathless
Cam - Calmodulin
CanB - Calcineurin B
Damm - Death associated molecule related to Mch2
Ddc - Dopa decarboxylase
Dhc36C - Dynein heavy chain at 36C
Dhc62B - Dynein heavy chain at 62B
Drep-4 - DNA fragmentation factor-related protein 4
f-cup - flyers-cup
fz - frizzled
GFP - Green fluorescence protein
Gyc-89Da - Guanylyl cyclase at 89Da
InR - Insulin-like receptor
Irbp - Inverted repeat-binding protein
Kap3 - Kinesin associated protein 3
Mdr50 - Multi drug resistance 50
mre11 - meiotic recombination 11
mus205 - mutagen-sensitive 205
mus304- mutagen-sensitive 205
ncd - non-claret disjunctional
nimB1 - nimrod B1
nimC1 - nimrod C1
nimC4 - nimrod C4
os - outstretched
Osi6 - Osiris 6
Otu - ovarian tumor
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
5
Phlpp - PH domain leucine-rich repeat protein phosphatase
Phr - photorepair
phr6-4 - (6-4)-photolyase
phyl - phyllopod
Pkc53E - Protein C kinase 53E
PPP4R2r - Protein phosphatase 4 regulatory subunit 2-related protein
Prx5037 - Peroxiredoxin 3
Puc - puckered
Rep - Rab escort protein
RNAi - RNA interference
SAC - Spindle assembly checkpoint
Sns - sticks and stones
Socs36E - Suppressor of cytokine signaling at 36E
Spn100A - Serpin 100A
Spn28Da - Serpin 28Da
Spn28Db - Serpin 28Db
Spn43Aa - Serpin 43Aa
ST6Gal - Sialyltransferase
Top1 - Topoisomerase 1
Traf4 - TNF-receptor-associated factor 4
Tws - twins
UAS - Upstream Activating Sequence
Wnt2 - Wnt oncogene analog 2
Wun - Wunen
wun2 - Wunen - 2
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
7
1.1. MODEL ORGANISM
Drosophila melanogaster has proven to be a very influential model for genetic and
biology studies in human diseases (1). Thanks to genomic sequencing, it is possible to
state that 70% of all known human diseases genes have functional orthologs in
Drosophila and some of the components in major signal transduction pathways that
play essential roles in neural development and cancer, like EGFR/RTK-Ras, PI3K,
Notch, Wnt, Jak-STAT, Hedgehog, and TGF-b are conserved (2-9). Furthermore, many
core components of these pathways were first discovered in Drosophila since it was
one of the first experimental systems to demonstrate that cancer is linked to heritable
genetic mutations as one of the first recessive lethal mutations found in Morgan’s
laboratory, lethal 7, revealed to give rise to transplantable, malignant tumors (10).
Drosophila offers advantages regarding modeling human diseases mainly because
present techniques permit cell type specific manipulation approach of gene expression
even in a single cell in vivo within the CNS and other tissues (11). Also, when
identifying and characterizing new genes, the Drosophila model brings several
advantages through different genetic tools that are able to provide us with information,
such as: the well-annotated and sequenced fly genome, the availability of thousands of
loss-of-function mutants for the majority of genes, the possibility of knocking-out genes
with targeted mutagenesis, insertional mutagenesis, or RNA interference techniques.
Gene overexpression with temporal and spatial precision is also possible using
technology like the binary transgene expression system, Gal4/UAS system, which
allows the expression of multiple transgenes in the same organism (12-14). In this
system, we are able to express the target transgene that is cloned downstream of a
UAS sequence in the same tissue specific pattern as the Gal4 activator. Furthermore,
the ability to introduce the transcriptional activator and the target UAS transgene in
distinct parental lines allows for the viability of parental stocks even in the presence of
toxic transgenes. A transgenic Gal4 Drosophila line can be used as a general resource
due to the fact that, once generated, this line expressing Gal4 in a given spatial pattern
can be crossed to any UAS target line. The opposite is also true as, once generated, a
Drosophila UAS target line can be driven anywhere in the embryo, larvae or adult fly
depending on which Gal4 line was selected to cross (15, 16).
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
8
1.2. TUMORIGENESIS
Cancer is a leading cause of death worldwide and is described as an uncontrolled
malignant growth of cells although its lethality often comes as a result of secondary
tumors, normally spread from the original site through the blood or lymphatic system
becoming highly invasive and aggressive (17, 18). A significant number of cancer are
associated with chromosomal instability, however, whether tumorigensis is the cause
or the consequence of chromosome instability is not known. Extensive work relates
weakened mitotic checkpoint to the transformation process. In mice, Mad2
heterozygotes (19) or hypomorphic alleles of BubR1 (20) produce a weakened
checkpoint and usually result in spontaneous tumors with a frequency of 27% and 6%
respectively. Heterozygous mice for Bub3 (21) or BubR1 (22) exposed to a carcinogen
also showed high percentage of tumorigenesis (70% in Bub3 heterozygotes versus
50% in wild-type and 11% in BubR1 heterzygote versus 0.4% in wildtype). In humans,
Cahill et al. in 1998 reported that 2 out of 19 colorectal cancer cell lines exhibit
heterozygous point mutations in the Bub1 and BubR1 genes (23). This led to an
extensive search for mutations in mitotic checkpoint genes in several tumors and tumor
cell lines. As an example, human patients with a rare recessive disorder mosaic
variegated aneuploidy (MVA) show a raise in aneuploidy rate and childhood cancers
(24). Some MVA patients have mutations in both alleles of the BubR1 gene (25, 26).
On the other hand, not many mitotic checkpoint genes mutations have been identified
in human cancers, however, altered transcription levels of these proteins associated
with hypermethylated promoters are thought to be very frequent (27). Accordingly
human tumor cells and cell lines show a failure in maintaining a mitotic checkpoint
signaling. The underlying concept is that weakened checkpoint signaling yet sufficient
to maintain a viable population of cells leads to a missegregation of chromosomes in
cell division and the consequently aneuploidy and chromosomal instability (28).
Genetic studies in Drosophila have helped to identify many genes that when
mutated or dysregulated lead to tumorigenesis. For instance, some players in the
Hippo pathway, defined as hyperplastic tumor suppressors, are involved in increased
proliferation or survival although not impeding tissue structure or differentiation. On the
other hand, some neoplastic tumor suppressors like Lgl involved in the apical-basal cell
polarity are usually responsible for problems in differentiation and failure to exit cell
cycle (29). An important aspect has been the ability to study these effects in the context
of the whole organism possible through several genetic and cell biology techniques.
For instance, the ability to create clones of mutant tissue aided researchers in their
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
9
quest to understand the role of microenvironment in tumor development and cell
competition mechanisms that offer a malignant cell with growth advantage
characteristics (29). Moreover, adding research using other model organisms, the
knowledge about cancer initiation and progression has greatly increased, even
revealing unknown molecular components and pathways, and helped improve
mammalian model systems for cancer (29). Using Drosophila as a model organism it
has been possible to identify several relevant genes highly studied in humans. Genes
like Notch, Hedgehog (Hh), Wingless, Runt, and Trithorax among others, have been
extensively elucidated in animal models and they seem to play a major role in the
pathophysiology of human cancer (51). Thomas Hunt Morgan described the Notch
mutation almost a century ago when he observed mutant flies with what appeared to
be notches in their wingblades, however only much later molecular analysis revealed
the importance of this highly conserved pathway (30, 31). The phenotype of this
mutation exhibits wings incised at tips and often along edges in heterozygous females
and lethality in males and homozygous females (32). Another interesting feature is the
lethal Notch-like phenotypes shown by Loss-of function mutations in the Drosophila
presenilin gene (33). Notch is related to cancer in humans through NOTCH1, highly
homologous to the Drosophila gene, found in chromosome 9 upon the translocation
t(7;9)(q34;q34.3) in a case of acute T cell lymphoblastic leukemia (34). The Hedgehog,
Wingless and runt genes were discovered by Christiane Nüsslein-Volhard and Eric
Wieschaus through their famous screen looking for embryonic mutants in Drosophila
(35) Phenotypically, hh homozygous embryos show the posterior naked portion of the
ventral surface of each segment deleted being replaced by a mirror image of the
anterior denticle belts. The length of wild-type larvae is greatly diminished and appears
to have a hedgehog-like appearance with a lawn of denticles (32). The association of
this gene to human cancer was first identified through the mutation of patched, the
human homolog of Drosophila, in the Gorlins syndrome, a rare syndrome linked with a
number of skeletal, skin, and neural abnormalities and usually related to the
development of multiple skin basal cell carcinomas, and a considerable increase in risk
for medulloblastoma and rhabdomyosarcoma (36). Wingless was originally identified
through a recessive mutation that confers an absence of one or both wings and/or
halters (37). Observation of embryos with embryonic lethal alleles showed a mirror-
image duplication of the denticle belts at the cost of naked cuticle (32). In human
cancer, wingless is dysregulated in the event of the loss of an APC allele, associated
with most colon carcinomas (38). Drosophila embryonic patterning and segmentation
studies revealed the pair-rule gene Runt. Runt embryos are shorter due to possessing
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
10
half the normal number of denticle belts (35). Mirror image duplications substitute the
deleted regions of what is left of the more anterior pattern elements (32). Runt relation
with human cancer is associated with the discovery of cDNAs of the alpha subunit of
polyomavirus binding protein 2 that is considerably homologous to that of Drosophila
segmentation gene runt (run). In humans RUNX1/AML1 also shows a run homology
region and it is a gene associated with the translocation t(8;21) (39). Interestingly,
AML1 is usually a target of chromosome translocations related with leukemia (40). For
instance, t(3;21) translocation in cases of myelodysplasia and blast transformation of
chronic myelogenous leukemia there is a fusion between either AML1 and EVI1 gene
(zinc fingercontaining transcription factor) or AML1 with EAP or MDS1 which function is
still unkown (41). The identification of the trithorax gene was due to its part of
maintaining expression of Antennapedia and bithorax homeotic gene complexes in
Drosophila melanogaster (42). Analyzing mutants demonstrate transformations of the
first and the third thoracic segments to the second thoracic segment, and the abdomen
showed alterations (32). In humans, acute leukemias are associated with the
MLL/ALL1 gene present in the human chromosome 11 band q23 usually involved in
interstitial deletions or reciprocal translocations between this region and chromosomes
1, 4, 6, 9, 10, or 19. Regarding homology, three regions are considered homologous to
sequences of the Drosophila trithorax gene (43, 44). The existence of translocations
involving MLL in leukemia is usually associated with unique clinical and biologic
features and with poor prognosis. Interestingly, topoisomerase II inhibitors can lead to
this disease when used to treat other malignancies since it causes double strand
breaks in DNA, leading to chromosomal MLL fusions. This feature might indicate an
involvement of the DNA repair pathway although information on such is still scarce
(45). The longstanding question was how such a large amount of different proteins
could give rise to the same disease. At first the suggestion involved the importance of a
dimerization of the amino-terminus of MLL (46). Furthermore, proteins fused with MLL
can be divide in two categories: nuclear and cytomplasmic and only six frequent
partners were responsible for over 85% of the diseases (47, 48). However, with access
to gene array technology, expression profiles between MLL and non-MLL leukemic
samples could be compared and showed a different pattern of gene expression that
discriminate that group from other leukemias (49). Further observation showed that
MLL fusion Proteins loose the H3K4 methyltransferase activity and are able to
transform hematopoietic cells into leukemia stem cells in an efficient way (50).
The knowledge of key pathways and genes discussed previously resulted from
several fly genes with mutant phenotypes consistent with defects in signaling and/or
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
11
gene transcription and it has been increasing due to studies of the vertebrate
counterpart of these genes. Furthermore, a rising number of evidences from several
studies suggest that elements of these pathways, found in human cancer, might
reactivate key cell programs in embryonic development since pathways as Notch,
Hedghog and Wingless show atypical signaling without activation due to mutation. Like
previously mentioned Runx and Trithorax are essential in normal hematopoietic
development and are usually associated with translocations relevant in acute
leukemias. All this information and knowledge of these highly conserved pathways is
expected to be used in a near future in order to contribute with therapies and drugs
targeting these specific development pathways to fight against problems like resistance
to standard chemotherapy and the subsequent recurrence caused by cancer stem
cells. However, the effects of these pathways seems to be context dependent, so
without a clear understanding it will be hard to discover effective drugs and therapies.
Lastly, being able to know what pathway is altered in a particular cancer using
biomarkers would be a great improvement for choosing the correct drug administration
(51).
Defects in mitosis could play a role in initiating tumorigenesis and consequently,
drugs that target this process might represent a successful antitumor target. The use of
drugs capable of arresting mitosis has become a widespread treatment for several
human tumors like breast, ovarian non-small-cell lung cancer among others. Tao et al.
in 2005 showed some of the first evidence regarding the link between mitotic arrest, the
mitotic checkpoint and cell death. However, this approach is not ideal due to secondary
effects in other otherwise normal tissues (28).
1.3. MITOSIS AND SPINDLE ASSEMBLY CHECKPOINT
(SAC)
Cell division involves a series of temporally and spatially regulated events that
must be completed in strict order for a successful outcome, the generation of two
daughter cells with normal genome content. One of the major forces behind this orderly
progression is the periodic rise and fall of cyclin-dependent kinases (Cdk) activity. In
order to achieve this, an important mechanism known as the ubiquitin-proteasome
pathway is responsible for the timely degradation of cyclins, which by themselves
regulate Cdks (52). Multiple cell cycle transitions like metaphase-anaphase transition
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
12
and exit from mitosis are regulated by the anaphase-promoting complex or cyclosome
(APC/C) a multisubunit ubiquitin ligase (52, 53). APC/C relies mainly on two activators
that directly bind to it activating its ubiquitin ligase activity: Cdc20 is the mitotic activator
and Cdh1 executes its function during telophase and G1. The substrates of APC/C,
during mitosis are Securin and the mitotic cyclin (cyclin B). Degradation of securin
allows the activation of separase which then proceeds to cleave one subunit of the
cohesion complexes setting in motion sister-chromatid separation. Cyclin B
degradation also leads to mitotic exit by inhibiting the activity of Cdk1 (54). During S-
phase cells duplicate their chromosomes that are kept together through sister-
chromatid cohesion. During mitosis, the correct attachment of chromosomes to the
mitotic spindle is under the surveillance of the spindle assembly checkpoint ensuring
that anaphase onset does not take place before microtubule attachment and tension
between two opposing kinetochores of all pairs of sister chromatids (54, 55). Previous
experiments showed that even a single unattached kinetochore is enough to arrest cell
in mitosis and removal of this kinetochore was able to undo this metaphase arrest and
allow cells to enter anaphase. These experiments demonstrated the existence of a
conserved mitotic checkpoint (56, 57). These findings suggest that APC/C activation
and the consequence degradation of securing and cyclin B, separase activation and
sister-chromatid separation only occurs when all pairs of sister kinetochores are
correctly attached to spindles microtubules and are under tension (54, 55). Spindle
assembly checkpoint (SAC) proteins inhibit APC/CCdc20 by both stoichiometric
binding to and sequestration of Cdc20 by Mad2 and BubR1, and Cdc20
phosphorylation by Bub1 (58). It is possible to define APC/C as a multisubunit ubiquitin
ligase distantly related to the cullin-RING family of ubiquitin ligases. Ubiquitin-
conjugating enzymes are then activated and recruited by the subcomplexes of APC/C
namely, Apc2 (a cullin-like subunit) and Apc11 (a RING-contaning subunit) (59). Small
peptide motifs, APC/C degrons like the destruction box (D-box with a consensus
RXXLXXXXN), which bind to WD40 domains, and the KEN-box (with a consensus of
KEN), mediate recognition of APC/C substrates. Moreover, another protein called
Apc10 also recognizes the D-box motif (60-62). The inhibition of APC, occurs thorugh a
complex formed by Mad2, BubR1, Bub3, and Cdc20 identified as mitotic checkpoint
complex (MCC) (63, 64). Mad2 was subject of several studies including conformational
and genetic studies that reveal that this protein was in fact a two-state protein which
binding to Cdc20 is closely regulated by the spindle checkpoint and has a vital
importance in maintaining chromosome stability (65, 66). Mad2 appears in two native
folds: a latent open conformer (O-Mad2 or N1-Mad2) and a closed conformer more
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
13
active in Cdc20 binding and APC/C inhibition (C-Mad2 or N2-Mad2) (67, 68). For
activation, Mad2 binds to Mad1 during the cell cycle maintaining a Mad1-C-Mad2 core
complex that when spindle checkpoint is active it is targeted to unattached
kinetochores. Upon this event, Mad1-C-Mad2 core complex recruits cytosolic O-Mad2
through O–C Mad2 dimerization and this precise dimerization is responsible for a
conformational alteration of O-Mad2 turning into a transient conformation I-Mad2. I-
Mad2 can then dissociate from Mad1-Mad2 core turning into a C-Mad2 without a ligand
and being able to bind to Cdc20. However the mechanics behind APC/C inhibition
through Mad2 binding to Cdc20 are not clear yet (69). Another key player in the spindle
assembly checkpoint is BubR1 and this protein seems to induce a higher inhibition of
APC/CCdc20 than Mad2 in vitro (26, 70). Analysis of BubR1 structure revealed a
domain required for kinetochore targeting named N-terminal tetratricopeptide repeat
(TPR), a conserved Bub3 binding motif GLEBS and also a C-terminal kinase domain.
However, the kinase domain of BubR1 is not conserved in yeast and is not necessary
for APC/C inhibition (71). Studies in yeast showed that Mad3 (the yeast homologue of
BubR1) contains KEN-boxes and a D-box motif responsible for Mad3 binding to Cdc20,
competing with APC/C for its subtrates, resulting in the consequent checkpoint
function, highly conserved mechanism higher organisms like human, mouse and flies
(71, 72). BubR1/Mad3 seem to act as a competitive, pseudo-substrate inhibitor of
APC/CCdc20 as it does not suffer APC/CCdc20-dependent degradation. The reason
behind this might be the acetylation of BubR1 at K250 (73). What is clear is that BubR1
and Mad2 are able to inhibit APC/CCdc20 in vitro independently, however, most likely
they collaborate by forming MCC. Studies with unattached kinetochores showed an
enhanced inhibition of APC/C through purified MCC components (Mad2, BubR1, and
Bub3) and an enhanced binding of BubR1 to APC/CCdc20. This enhancement
depends on kinetochore-bound Mad1-Mad2 core complex and Mad2 dimerization (74).
The role of unattached kinetochores in the formation of MCC seems to be the
production of C-Mad2 and generate an initial Mad2–Cdc20 complex that allows further
interaction with BubR1–Bub3 (74). The binding of BubR1–Bub3 and Mad2 to Cdc20
changes how Cdc20 and APC/C interact. Therefore, it appears that the MCC inhibits
the activity of APC/CCdc20 by impeding substrate binding both competitively and non-
competitively (75). Inhibition of APC/C can also be performed by Cdc20
phosphorylation. However, it is unclear how this occurs (76). Through the course of
evolution, most checkpoint proteins remained conserved, however the knowledge
about the role of checkpoint proteins and their importance is still increasing, including
for Bub3. It is possible to find Bub3 in most eukaryotes (77-79). This protein contains
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
14
seven WD40 repeats, which are found in several distinct proteins and it is believed to
be involved in protein-protein interactions (80). Throughout mitosis, Bub3 is visible on
kinetochores during prophase, decreasing its levels by metaphase. If kinetochores
remain unattached the kinetochore-associated Bub3 antigen increases (79). Studies in
mice showed that one functional copy of the Bub3 gene is enough for normal
development since heterozygous Bub3 mice do not show significant abnormalities in
development or fertility (81). However, upon analyzing the progeny of their
heterozygous crosses they found no Bub3−/− pups, suggesting an embryonic lethal
phenotype. Therefore, Bub3 seems to be essential in mouse for normal mitosis and
early development (81). Likewise, depletion of Bub3 in Drosophila also leads to
embryonic lethality at the larval/pupal transition stage and cells from these mutants
larvae fail to arrest in mitosis in response to microtubule-depolymerizing drugs and
show severe mitotic abnormalities like premature chromatid separation, lagging
chromosomes and chromatin bridging (82, 83). It was shown that Bub3 forms two
independent complexes with Bub1 and BubR1 in mitotic mammalian and Xenopus cells
and seems to be required for the localization these proteins to the kinetochore. Yeast
has also provided some evidence of a complex with Mad1 and Bub1. Moreover, this
complex is dependent on Mad2 and seems essential for a checkpoint response (84).
Research in Drosophila S2 Bub3 depleted cells revealed an accelerated mitotic timing
when compared to normal cells (from NEBD to anaphase onset). It was also suggested
that there is a slower progression through early stages of mitosis specifically a delay in
prophase (84). Recently Niikura and co-workers linked Bub3 with cell death defined as
caspase independent mitotic death (CIMD). They demonstrated that when BUB3 is
dissociated from BUB1 it activates CIMD. BUB1 interacts with p73, a structural and
functional homolog of the p53 tumorsuppressor protein, in cells showing CIMD. It
seems that Y99 phosphorylation on p73 by c-Abl tyrosine kinase and S19
phosphorylation on BUB3 by BUB1 kinase are required for CIMD activation. These
observations point to the hypothesis that CIMD protects cells from aneuploidy, which
eventually leads to tumorigenesis (85).
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
15
Figure I.1 - Schematic representation of the stages of mitosis. Adapted from Weaver and Cleveland, 2005 (28).
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
16
1.4. APOPTOSIS
The ability to self-destruct by undergoing apoptosis is present in most animal
cells (86). Apoptosis is a genetically encoded process with characteristic morphological
features that serve to dispose of superfluous or unwanted cells and when this process
fails, it can be associated with many human diseases (87). In order for apoptosis to
occur, it is necessary to activate caspases, a family of cystein proteases that are
present in cells as weakly active zymogens (88). Caspases were initially identified in a
screen for C. elegans mutants unable to undergo developmental programmed cell
death (89). Different stimuli may be responsible for caspase activation such as
developmental signals as well as various forms of persistent cellular stress or injury
(86). Amongst some of these are DNA damage, viral infection, hypoxia, increased
presence of reactive oxygen species, loss of cellular adhesion, accumulation of
unfolded proteins, excitotoxicity, cytoskeletal damage as well as other insults (86).
Healthy cells contain caspase zymogens as inactive precursors (88). Caspase
zymogens possess a small amount of catalytic activity, however they are kept in check
by several regulatory molecules and only after receiving an apoptotic signal they are
able to perform its full activity upon undergoing proteolytic processing and generate two
subunits (9, 90). The cleavage of the zymogen is not always required for caspase
activation. Structural studies predict that the mature caspase is a heterotetramer
formed by two heterodimers originating from two precursor molecules (9, 90).
Moreover, beside the regions that give rise to the two subunits, procaspases include N-
terminal prodomains of different sizes and when analyzing these lengths one can
divide caspases into two groups: those with relatively long prodomain and those with a
shorter prodomain (9, 90). Several specific protein-protein interaction domains of
relevant importance in caspase activation are possible due to the existence of long
prodomains (9, 90). Procaspases are then recruited by these domains to specific death
signaling complexes which in turn lead to their autocatalytic activation by a mechanism
often designated “proximity-induced” activation (9, 90). The caspases recruited to
signaling complexes, therefore being activated, are known as the initiator caspases
due to providing a link between cell signaling and apoptotic execution (88). The main
initiator caspases present in mammals are caspase-2, -8, -9 and -10 and in Drosophila
is the Nedd2-like caspase (DRONC). Caspases -2, -9 and DRONC contain a caspase
recruitment domain (CARD) and caspase -8 and -10 contain a pair of death effector
domains (DEDs) (88). All these domains are able to bind to similar motifs in adaptor
proteins, however caspases lacking a long prodomain are not able to self-activate and
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
17
need to be cleavage by previously activated initiator caspases (9, 90). After being
activated these caspases cleave most of the cellular substrates upon apoptosis,
therefore being designated effector caspases. Some of the most important effector
caspases are caspases-3, -6 and 7 in mammals and drICE in Drosophila (88).
Activation of this caspases must be kept in check. A class of anti-apoptotic proteins
called inhibitor of apoptosis proteins IAPs accomplishes this task (91, 92). IAPs are
able to bind to both types of caspases through their BIR domains (93, 94) and they also
contain a RING motif that enable them to act as E3 ubiquitin ligases promoting
ubiquitination of caspases (95, 96). In Drosophila in order to prevent caspase activation
and apoptosis in virtually all somatic cells one key IAP must be present, DIAP1 (97-99).
DIAP1 exists in an auto-inhibited conformation being unable to inhibit the effector
caspase drICE (100). In order to disabled the auto-inhibition a caspase-mediated
cleavage of DIAP must occur (100). A cleaved DIAP is then able to bind to mature
drICE resulting on its inhibition and apparently targeting drICE for ubiquitylation (100).
If a cell is doomed to die, specific antagonists inactivate IAPs promoting apoptosis (98,
101, 102). In Drosophila it is possible to find three IAP antagonists clustered together in
the genome, Reaper (Rpr), Head involution defective (Hid) and Grim, and proceeding
to deletion of these genes gives rise to severe apoptosis inhibition (103-105). Another
IAP antagonist, Sickle (Skl), similar to Rpr was also found, however, since there is no
mutants and a thorough analysis of its role in apoptosis is still not possible (106). IAP
antagonists bind to IAPs through an evolutionary conserved feature, the N-terminal
IBM (IAP-binding motif) (94, 107), resulting in the displacement of IAP-bound caspases
(108-110). The active caspases generate a proteolytic cascade that proceeds to the
destruction of the cell. Moreover, IAP antagonists also stimulate IAP turnover by
proteasomal degradation (111, 112). Drosophila IAP antagonists expressed in human
cells also showed cell death inducing activity (113, 114). It is also important to mention
that IAP antagonists, Rpr, Hid and Grim may be induced by a several upstream factors
such as the p35 gene (115, 116) or the Jun N-terminal Kinase (JNK) pathway (117).
Apparently, these factors seem to direct developmental or environmental stimuli like
irradiation or cell competition to apoptosis (118-120). The current image of apoptosis in
Drosophila defines DIAP1 as a central brake on apoptosis performing the inhibition of
activated caspases. DIAP is able to bind to caspases resulting in its sequestration
impeding the caspase binding to its target or even in caspase degradation. Apoptosis
occurs when IBM proteins Rpr, Grim, Hid and Skl compete with caspases for binding to
DIAP1. Additionally Rpr and Grim are also able to suppress DIAP1 translation (108,
112). Several other IAP-like proteins in flies, DIAP2, deterin and dBRUCE seem to be
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
18
able to suppress apoptosis although their specific mechanisms are not completely
elucidated yet (121-124). Moreover, developmental and environmental inputs may
influence the apoptotic pathway either by changing the transcription and activity of IBM
proteins, or synthesis of some caspases and even the synthesis of some IAPs (103,
125-127).
Figure I.2 - Schematic representation of the understanding of apoptosis in Drosophila. Adapted from Kornbluth,
2005 (128).
There are two types of distinguishable apoptosis, a developmental regulated
(129) one and a stress induced one (130). The developmental regulated apoptosis is
important in processes like the cleft formed between the maxillary and mandibular
segments in early Drosophila embryos, which is possible through the activation of
Reaper by Deformed, a Hox protein (131). The elimination of the excess of
interomatidial cells during metamorphosis is also a good example of the developmental
apoptosis importance (132). In addition, another morphogenic role of apoptosis was
recently found, the formation of leg joints, resulting from a induced discontinuity of Dpp
signaling levels in the presumptive region providing with the required activation of
Reaper (133). The other type of apoptosis, the stress induced apoptosis is not
programmed however it has become a tool to dispose of cells damaged by insults like
irradiation, heat shocks, hypoxia among other examples (105, 134, 135). In Drosophila,
the wing imaginal disc is a suitable system to study stress-induced apoptosis since
developmental apoptosis is almost none existent therefore the majority of observed
apoptosis is caused by stress treatments (105, 134, 135). To induce cell death in
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
19
imaginal tissue there are several treatments available: irradiation of larvae with
moderate doses of X-Ray (eliminating a number close to 40-60% of wing imaginal cells
(136); heat-shock at 37ºC for 3 hours is also a mean to achieve high apoptotic levels
(135). Which genes are involved in the apoptotic stress response? The fly homolog of
the tumor suppressor gene p53, Dmp53 activates the apoptotic response after
irradiation and mutants or dominant negatives for this protein produce very little
apoptosis response to X-Rays (4, 116, 118, 120). It was shown that IAP antagonist like
Rpr, Hid and Sickle are transcriptional targets of Dmp53 and that X-ray induced
apoptosis in wing disc is highly dependent on Hid (115, 116). Upon γ-Ray irradiation,
the activity of the JNK pathway is altered giving rise to an up-regulation of the JNK
target gene puckered (puc, a phosphatase that negatively regulates the pathway
activity) present in the entire wing disc (117). Recently, the Hippo pathway and
apoptosis showed a connection (137) because γ-Rays are able to induce Hippo Kinase
activity in a Dmp53-dependent manner in cell culture and imaginal discs. Furthermore,
DIAP1 is also a target of the Hippo pathway (138). An important characteristic of the
stress-induced apoptosis is the non-lethality of some treatments even after the
production of large amounts of cell death. Even after more than 50% of the cells in
wings imaginal discs enter apoptosis, the flies recover and are able to restore organs of
normal shape and size (135, 136). Therefore, an additional proliferation by surviving
cells must compensate and originate a phenomenon called compensatory proliferation
(136, 139). Interestingly, Drosophila and its genetic tools allow us to interfere with
apoptosis at quite different stages of its progression: hinder the function of upstream
activators therefore meddling at the activation of the apoptotic program; the activation
of caspases; the function of effector caspases (140).
1) In order to prevent upstream activation it is possible to use p53 null mutants
or dominant negative forms of the gene (4, 116, 118, 120). Inhibition of JNK activity or
overexpression of puc are also effective (141, 142).
2) When the objective is to prevent caspase activation a deficiency in Rpr, Hid
and Grim functions can be used with the H99 deletion which impede DIAP1 inactivation
suppressing apoptosis (103). Another way to achieve prevention in caspase activation
is through the overexpression of DIAP1 using UAS-DIAP1 transgene, although it does
not inhibit apoptosis completely (111). At last, mutants or dominant negative forms of
the initiator caspase DRONC, do not allow the activation of the effector caspase drICE
(143).
3) In order to prevent effector caspase function, the usual method is to utilize
the baculovirus protein P35 (144) which blocks the function of drICE (unaffecting
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
20
DRONC) allowing however, its activation. It seems like P35 binds to drICE inhibiting its
protease activity (145).
Figure I.3 - A schematic view of Drosophila apoptotic machinery. Adapted from Martín, 2009 (140).
In order to prevent upstream activation it is possible to use p53 null mutants or
dominant negative forms of the gene (4, 116, 118, 120). Inhibition of JNK activity or
overexpression of puc are also effective (141, 142). When the objective is to prevent
caspase activation a deficiency in Rpr, Hid and Grim functions can be used with the
H99 deletion which impede DIAP1 inactivation suppressing apoptosis (103). Another
way to achieve prevention in caspase activation is through the overexpression of
DIAP1 using UAS-DIAP1 transgene, although it does not inhibit apoptosis completely
(111). At last, mutants or dominant negative forms of the initiator caspase DRONC, do
not allow the activation of the effector caspase drICE (143). Finally, in order to prevent
effector caspase function, the usual method is to utilize the baculovirus protein P35
(144) which blocks the function of drICE (not affecting DRONC) allowing however, its
activation. It seems that P35 binds to drICE inhibiting its protease activity (145).
The effects of arresting apoptosis at different stages were observed and
compared (140). Either using p53 mutants or overexpressing its the dominant negative,
therefore inhibiting apoptosis, does not seem to have a striking effect on the
development or morphology of the flies, however, there may be uncharacterized
consequences on longevity or fertility. An analogous outcome is observed upon
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
21
overexpressing DIAP1, impeding caspase activation. On the other hand, hinder the
activity of the effector caspase drICE activity using P35 produces great developmental
consequences (135). Without radiation, P35 by itself does not cause major alterations.
However if apoptosis is induced in cells expressing P35 using radiation, a group of
“undead cells” arise. These cells have their apoptosis program active but due to the
blockage of effector caspases they cannot die. These cells also keep their original
identity as they differentiate into wing structures (135) but retain all the molecular
markers of apoptosis such as low levels of DIAP1 and high levels of Hid, DRONC or
drICE proteins and even behave as normal apoptotic cells moving to the basal
membrane of epithelia (140). Genes expressed by undead cells were identified,
namely, decapentaplegic (dpp) and wingless (wg) which are defined by being major
players in patterning and growth in the wing disc (146). This feature allied with the
persistence of undead cells is probably responsible for an abnormal growth of the P35
containing compartments although additional proof is necessary (140).
1.5. MAIN TECHNOLOGIES
1.5.1. MICROARRAYS
Nowadays, aided by several “whole genome sequencing” projects, the
availability of genetic information is outstanding. RNA expression in cell and tissue at a
given moment can now be studied using high-throughput techniques due to the
emergence of microarray-based technologies in the transcriptomic field (147).
Microarray technology is becoming increasingly important for molecular biology mainly
because it analyzes the overall transcription profile of a cell or tissue instead of being
limited to “one gene at a time” (148). Microarray techonology was introduced 17 years
ago in a seminal paper by Schena et al. (149). This technology grants quantitative
monitoring of the expression profile of thousand of genes (150) and relies on base pair
complementary for identification of mRNA highly expressed in a sample of interest
(151). Microarray assays can be divided in five distinct experimental steps: biological
query, sample preparation, biochemical reaction, detection, data visualization and
modeling (152). Upon concluding a microarray assay, a researcher ends up with a
snapshot of genes expressed in a cell or tissue at the time of harvest and RNA
purification, which is a way to compare samples horizontally and obtain an expression
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
22
profile. For instance, in biomedicine the value of expression patterns is immense
yielding valuable insight on biological processes in diseased tissue, disease prediction,
diagnosis and treatment, cellular differentiation, development and drug discovery (153).
Currently DNA microarrays offer a way of exploring the genome in a systematic and
understandable way and they present themselves as oligonucleotide arrays and a
variety of cDNA arrays (154). Oligonucleotide arrays contain short fragments of DNA
spotted or synthesized in situ on solid supports (glass, coated glass, silicon or plastic)
in a way that knowing the location defines which oligonucleotide is present there (155).
They differ from cDNA, firstly because their probes are a set of 20 to 25 short
oligonucleotides, specific for each gene or exon, and are synthesized in situ by
photolithographic deposition. Secondly, the arrays are hybridized to a single
biotinylated amplified RNA sample, and an algorithm computes the result (156). It is
also important to refer to the general superiority of oligonucleotide-based microarrays
in terms of quality and reproducibility although there is a need for a larger capital input
as well as a larger amount of starting biological material (157, 158). cDNA arrays, on
the other hand, contain long fragments of DNA, reaching thousands of base pairs, and
are created using robotically spotting of individual samples of purified cDNA clones in a
solid support like glass or a membrane (159). Some of the most general applications of
cDNA arrays are comparative and functional genome analysis. Recently it was used in
drug resistance tests (159).
Following a microarray experiment, the amount information generated is
enormous and to be able to find the relevant information is still a quite challenge.
Therefore, bioinformatics methods are essential in order to extract pertinent biological
information (160). Over the last few years, these bioinformatic tools have indeed
contributed to several studies and large gene lists analysis from various high
throughput techniques. However, there is no rule or gold standard as far as these tools
go and there is still significant room for improvement (160). On the other hand as these
bioinformatics techniques evolve so does the difficulty to search and choose the right
way to perform, for instance, a microarray analysis (160). The complexity achieved by
the overabundance of tools led to several problems including which tools are most
fitting for a particular type of analysis (160). Because of the complexity of biological
data mining situations, we can define the analysis of large gene lists with the latest
bioinformatics techniques as a process of exploratory data-mining instead of a solution
based on statistics. Clearly, the current solutions depend on the aid of human
knowledge and thought, and this, supported with computing algorithms, integrated
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
23
annotations databases, and P-values derived from statistical methods is what allows
for a comprehensive and as best as possible analysis (160).
1.5.2 RNA interference
RNA interference (RNAi) was first seen by researchers trying to alter flower
colors in petunias (161). However, only when work on RNAi in Caenorhabditis elegans
was published (162) the mechanism became better understood. Later it was
demonstrated that RNAi works efficiently in many cell types including Drosophila and
can be used for large-scale gene silencing applications (163-165). Therefore, large-
scale RNAi-based screening is now a important tool for the identification of gene
products involved in different biological processes and pathways (166, 167). Research
centers like the Vienna Drosophila RNAi Center (VDRC) maintain and distribute to the
research community transgenic RNAi stocks constructed by Dickson and Keleman
groups (168). RNAi mechanism was unraveled through a series of genetic and
biological studies. The model starts with the conversion of the dsRNA into small RNAs
(siRNAs) by an RNase III family nuclease named Dicer (169). Then these small RNAs
join a multicomponent nuclease complex, and become the tool to identify homologous
mRNAs (170). Two of the major keys
players in this system are universally
conserved namely, Dicer and
Argonaute (Ago) gene family
members. Structure wise, dicer
shows a tandem repeat of RNase III
catalytic domains, a carboxylterminal
deRNA binding domain, an animo-
terminal DExH/DEAH RNA helicase
domain, and a PAZ domain (169,
171). Ago proteins, that take part of
the RNA-induced silencing complex
(RISC) include a PAZ domain and a
carboxyl-terminal PIWI domain
(172).
Figure I.4 - Schematic representation of a model of RNA interference in
mammalian cells. Adapted from Paddison, 2002 (172).
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
24
1.6 AIMS
In his project we established a few goals:
Use microarray technologies to determine the transcription profile of cells from
the wing disc of larvae expression an RNAi construct against Bub3 that at the
same time were prevented from undergoing apoptosis and showed clear sings
of transformation.
Apply a bioinformatic approach to identify possible genes or gene networks
associated with the transformation phenotype.
Perform a RNAi screen to validate candidate genes chosen through the
previous bioinformatic approaches
Eventually determine which which are the genes responsible for Bub3-related
tumorigenesis.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
25
CHAPTER 2
MATERIALS AND METHODS
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
26
2.1. FLY STOCKS USED
Below are The Drosophila stocks used during this thesis:
Ap-Gal4>UAS-GFP/CyO;
Ap-Gal4>UAS-P35/CyO;
Ap-Gal4>UAS-GFP UAS-Bub3RNAi/CyO;
Ap-Gal4>UAS-GFP UAS-Bub3RNAi/UAS-P35;
Ap-Gal4>UAS-GFP UAS-Bub3RNAi UAS-DIAP1/CyO.
The RNA interference constructs used are mentioned in annex 1 and are
described in Flybase (http:/flybase.bio.indiana.edu/). Standard procedures were used
for the flies management.
2.2. RNA EXTRACTION FROM WING IMAGINAL DISCS
2.2.1. COLLECTION OF WING IMAGINAL DISCS
To obtain RNA we dissected in Phosphate Buffer Saline (PBS) 70 larvae (140
wing imaginal discs) of each genotype:
Ap-Gal4>UAS-GFP/CyO;
Ap-Gal4>UAS-P35/CyO;
Ap-Gal4>UAS-GFP UAS-Bub3RNAi/CyO;
Ap-Gal4>UAS-GFP UAS-BubRNAi /UAS-P35.
After dissection, the larvae were conserved frozen at -80ºC.
2.2.2. RNA EXTRACTION
Dissected wing discs were incubated in 500 µl TRIzol (ref. 15596-026,
Invitrogen) for 5 minutes (min) at room temperature (RT). Solution was cleared by
centrifugation at 12000g for 10 min at 4ºC.
The supernatant was transferred to a 2 ml microfuge tube and 100 µl of
chloroform (ref. 1.02445.2500, Merck) were added and vortexed for 1 min.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
27
Samples were incubated 5 min at RT, centrifuged for 15 min at 4ºC at max
speed (13,2 rpm) and 250 µl of the aqueous phase were transferred to a 2 ml tube.
Afterwards we used RNeasy Mini Kit (ref. 74104, Qiagen) to proceed with the
extraction. We added 875 µl of RLT buffer and briefly vortex the mixture followed by the
addition of 625 µl absolute Etanol (ref. 1.00983.2511, Merck) and vortex again for a
few seconds.
The mix was loaded onto a RNeasy column and centrifuged for 30 seconds at
RT, max speed (13,2 rpm) and the flow through was discarded. This last procedure
was repeated two more times.
The columns were washed with 500 µl RPE buffer and centrifuged for 30
seconds at RT, max speed (13,2 rpm), and the flow through was discarded. This last
procedure was done three times. The column was spin dry for 2 min and placed in a
1ml RNAse free tube followed by the addition of 10 µl of water to the center of the
column.
After 2 min at RT the colum was centrifuged at max speed (13,2 rpm).
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
28
2.3. MICROARRAYS
Figure II.1 - Required equipment for microarray protocol. Adapted from One-Color Microarray-Based
Gene Expression Analysis Low Input Quick Amp Labeling, Agilent Technologies.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
29
Figure II.2 - Required reagents for microarray protocol. Adapted from One-Color Microarray-Based
Gene Expression Analysis Low Input Quick Amp Labeling, Agilent Technologies.
Started by performing the sample preparation
Step 1. Preparation of the One-Color Spike Mix
- Equilibrated water baths to 37°C, 65°C, 80°C, 40°C and 70°C.
- Vigorously mixed the One-Color Spike Mix stock solution on a vortex mixer.
- Heated at 37°C for 5 minutes, and mixed on a vortex mixer once more.
- Briefly span in a centrifuge to drive contents to the bottom of the tube prior
to opening.
Step 2. Preparation of the labeling reaction (Time required: approximately 5,5 hours)
- Added 10 to 200 ng of total RNA to a 1,5-mL microcentrifuge tube in a final
volume of 1,5 µL.
- Added 2 µL of diluted Spike Mix to each tube.
- We prepared and added T7 Promoter Primer:
o Mixed the T7 Promoter Primer and water to prepare the T7 Promoter
Primer Master Mix
o Added 1.8 µL of T7 Promoter Primer Mix to the tube that contains 3.5 µL
of total RNA and diluted RNA spike-in controls.
o Denatured the primer and the template by incubating the reaction at
65°C in a circulating water bath for 10 minutes.
o Placed the reactions on ice and incubate for 5 minutes
- Prewarmed the 5X first strand buffer at 80°C for 3 to 4 minutes to ensure
adequate resuspensions of the buffer components.
- Prepared and add cDNA Master Mix:
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
30
o Immediately prior to use, we added the components for cDNA Master
Mix (description in original protocol)and kept it at room temperature.
o We then Briefly spin each sample tube in a microcentrifuge to drive
down the contents from the tube walls and the lid.
o Added 4.7 µL of cDNA Master Mix to each sample tube and mix by
pipetting up and down.
o Incubated samples at 40°C in a circulating water bath for 2 hours.
o Moved samples to a 70°C circulating water bath and incubate for 15
minutes.
o Moved samples to ice. Incubated for 5 minutes.
o Span the samples briefly in a microcentrifuge to drive down tube
contents from the tube walls and lid.
- Prepared and added Transcription Master Mix
Step 3. Purification of the labeled/amplified RNA (Time required: approximately 0,5
hours)
- Added 84 µL of nuclease-free water to your cRNA sample, for a total volume of
100 µL.
- Added 350 µL of Buffer RLT and mixed well by pipetting.
- Added 250 µL of ethanol (96% to 100% purity) and mixed thoroughly by
- pipetting.
- Transfered the 700 µL of the cRNA sample to an RNeasy mini column in a 2 mL
collection tube. Centrifuged the sample at 4°C for 30 seconds at 13,000 rpm.
Discarded the flow-through and collection tube.
- Transferred the RNeasy column to a new collection tube and add 500 µL of
buffer RPE (containing ethanol) to the column. Centrifuged the sample at 4°C
for 30 seconds at 13,000 rpm. Discarded the flow-through. Re-use the
collection tube.
- Added another 500 µL of buffer RPE to the column. Centrifuged the sample at
4°C for 60 seconds at 13,000 rpm. Discarded the flow-through and the
collection tube.
- Eluted the cleaned cRNA sample by transferring the RNeasy column to a new
1.5 mL collection tube. Added 30 µL RNase-free water directly onto the RNeasy
filter membrane. Waited 60 seconds, then centrifuged at 4°C for 30 seconds at
13,000 rpm.
- Maintained the cRNA sample-containing flow-through on ice. Discarded the
RNeasy column.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
31
Hybridization
Step 1. Prepared the 10X Blocking Agent
- Added 500 µL of nuclease-free water to the vial containing lyophilized 10X
Blocking Agent supplied with the Agilent Gene Expression Hybridization Kit
- Mixed by gently vortexing.
- Drived down any material adhering to the tube walls or cap by centrifuging for 5
to 10 seconds.
Step 2. Prepared hybridization samples
- Equilibrated water bath to 60°C.
- For each microarray, added each of the components as indicated in the original
protocol
- Mixed well but gently on a vortex mixer.
- Incubated at 60°C for exactly 30 minutes to fragment RNA.
- Immediately cooled on ice for one minute.
- Add 2x GEx Hybridization Buffer HI-RPM to the 1-pack, 2-pack, 4-pack, and 8-
pack microarray formats at the appropriate volume to stop the fragmentation
reaction. Information detailed in the original protocol
- Mixed well by careful pipetting.
- Span for 1 minute at room temperature at 13,000 rpm in a microcentrifuge to
drive the sample off the walls and lid and to aid in bubble reduction.
- Placed sample on ice and loaded onto the array as soon as possible.
Microarray Wash
Step 1. Proceeded to add Triton X-102 to Gene Expression wash buffers
- Opened the cardboard box with the cubitainer of wash buffer and carefully
removed the outer and inner caps from the cubitainer.
- Used a pipette to add 2 mL of the provided 10% Triton X-102 into the wash
buffer in the cubitainer.
- Replaced the original inner and outer caps and mixed the buffer carefully but
thoroughly by inverting the container 5 to 6 times.
- Carefully removed the outer and inner caps and install the spigot provided
with the wash buffer.
Step 2. Prewarmed Gene Expression Wash Buffer 2 to 37°C
- Dispensed 1000 mL of Gene Expression Wash Buffer 2 directly into a sterile
1000-mL bottle.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
32
- Capped the 1000-mL bottle and place in a 37°C water bath the night before
washing arrays.
Step 3. Prepared the equipment
- Added the slide rack and stir bar to the staining dish.
- Transfered the staining dish with the slide rack and stir bar to a magnetic stir
plate.
- Filled the staining dish with 100% acetonitrile.
- Turned on the magnetic stir plate and adjusted the speed to a setting of 4
(medium speed).
- Washed for 5 minutes.
- Discarded the acetonitrile
- Repeated from step 3
- Air-dried the staining dish in the vented fume hood.
- Proceeded to “Milli-Q water wash”
- Washed all dishes, racks, and stir bars with Milli-Q water.
- Ran copious amounts of Milli-Q water through the staining dish.
- Emptied out the water collected in the dish.
- Discarded the Milli-Q water.
Step 4. Washed the microarray slides with GE Wash Buffer 1 and 2
Finally, Scanned the microarray slides and extract the data.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
34
3.1. INTRODUCTION
Previous work in the laboratory addressed the role of checkpoint genes in
tumorigenesis. As homozygous mutants for Bub3 die during the third instar larval stage
(84), in vivo RNAi transgenic technology was used to knockdown Bub3 in the dorsal
region of the wing imaginal disc (173). Upon knockdown, the adult fly thoraxes show
loss of bristles and empty sockets when compared with controls. This phenotype was
the result from increased cellular death in the wing imaginal disc and it is in accordance
with reports for an increase apoptosis when SAC genes are removed (81, 84).
However, if the apoptosis inhibitor DIAP1 (174) is expressed where Bub3 is knockdown
the adult thorax becomes larger and deformed with missing bristles and empty sockets
and the wing disc shows a higher number of cells. However, if in the tissue where Bub3
is knockdown the stronger apoptotic inhibitor p35 is expressed, the larval wing imaginal
disc becomes highly hyperplastic (175). We used microarray and bioinformatics
approach to identify that were specifically misregulated in these hyperplastic cells.
Then we followed with an in vivo screen with RNAi technology of candidate genes.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
35
Figure III.1 - images from the tumorigenesis
model adopted. a) Wing imaginal disc illustration
showing the division of the anterior and posterior
portions with a blue line (A - anterior portion, P -
posterior portion), the division of the ventral and
dorsal side through a red line (D - dorsal portion,
V - ventral portion). The green color represents
the portion that will generate the wing pouch, the
yellow color represents the portion that will
generate the hinge and the blue color represents
the portion that will generate the body wall. b)
Shows a larvae expressing GFP through a GAL4-
UAS system in its wing imaginal discs. c) Shows
a larvae expressing GFP, Bub3RNAi
and p35
through a GAL4-UAS system in its wing imaginal
discs. In both b)and c) the GFP is expressed in
the dorsal compartment of the wing disc under
the control of the apterous promoter (REF see
above). The size of this wing imaginal disc is
considerably bigger than wild type.
3.2. SELECTED
GENOTYPES
To determine the molecular basis for the
link between SAC genes and tumour formation
in the absence of apoptosis we used four
cellular genotypes for the microarray analysis.
The regulatory region of the apterous gene was
used to drive Gal4 expression in the dorsal
compartment of the wing disc (figure III.1) (176).
The non-expression in the ventral portion allows
for an internal control with a pool of cells not
expressing the RNAi of the candidate genes.
The first transgenic fly has Bub3 knocked down
through RNAi specifically on the dorsal
compartment of the wing disc and expresses
simultaneously GFP (Ap-Gal4>UASGFP
UASBub3RNAi/ CyO). The wing imaginal disc
shows high levels of apoptosis and is
significantly smaller than wild type (Figure
III.2.c). The adult fly presents loss of bristles
and empty sockets with altered wings that often
are necrotic (Figure III.2.d). In the second
transgenic fly, the apoptosis presented in the
previous genotype was blocked. We used two
different inhibitors, the previously mentioned
DIAP1 and P35. We crossed knockdown flies
for Bub3, expressing GFP with flies transgenic
for a UASDIAP1 construct and obtained a
recombinant fly expressing Bub3RNAi, GFP and
DIAP1 in the dorsal compartment of the wing
disc (Ap-Gal4>UASGFP UASBub3RNAi
UASDIAP1/ CyO). Wing imaginal discs from
this genotype are bigger with a higher number
of cells (Figure III.2.e). The adult fly presents a
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
36
larger, and generally deformed thorax, with some loss of bristles, empty sockets and
deformed wings showing necrosis (Figure III.2.f). To inhibit apoptosis with p35, flies
with the UASP35 construct (Ap-Gal4>UASP35/ CyO) were crossed with flies
expressing Bub3RNAi, and GFP in the dorsal compartment, and larvae for Bub3
expressing GFP and P35 (Ap-Gal4>UASGFP UASBub3RNAi/ UASP35). The wing
imaginal discs show overgrowth, loosing most of its structure with a significant
increased size (Figure III.2.g). This particular genotype is lethal as there are no viable
adult. As controls, we used flies expressing only GFP (Ap-Gal4>UASGFP/ CyO) as
their wing imaginal disc remains unaltered (Figure III.2.a). The adult fly, shows no sign
of abnormal phenotype revealing a full set of bristles, a normal thorax and wings
(Figure III.2.b). Finally, we used flies expressing only P35 (Ap-Gal4>UASP35/ CyO).
These flies are a control for the effects of inhibiting apoptosis. They show a normal
thorax although it has more bristles and some presented one black dot in each side on
the boundary between notum/scutellum (Figure III.2.h).
Figure III.2 - Selected genotypes. (a) Wing imaginal disc of Drosophila melanogaster expressing GFP through a
GAL4-UAS System in its dorsal region portion drived by the apterous promoter showing a wild type structure. (b) An
adult Drosophila melanogaster expressing GFP especially in the thorax and wings through a GAL4-UAS System driven
by the apterous promoter. It shows an essentially a wild type thorax. (c) Wing imaginal disc of Drosophila melanogaster
expressing GFP and Bub3RNAi
through a GAL4-UAS system in its dorsal region driven by the apterous promoter, (d) An
adult Drosophila melanogaster expressing GFP and Bub3RNAi
through a GAL4-UAS system driven by the apterous
promoter especially in the thorax and dorsal region of wings. It shows a thorax with empty sockets, missing bristles, and
deformed wings. (e) Wing imaginal disc of Drosophila melanogaster expressing GFP, Bub3RNAi
and DIAP1 through a
GAL4-UAS system in its dorsal region driven by the apterous promoter (f) An adult Drosophila melanogaster expressing
GFP, Bub3RNAi
and DIAP1 in the thorax and wings showing a larger and generally deformed thorax with empty sockets,
and deformed necrotic wings. (g) Wing imaginal disc of a Drosophila melanogaster larvae expressing GFP, Bub3RNAi
and p35 through a GAL4-UAS system in its dorsal region driven by the apterous promoter showing a large overgrowth.
(h) An adult Drosophila melanogaster expressing P35 through a GAL4-UAS system driven by the apterous promoter in
the thorax and wings. It shows a thorax resembling a wild type one although seeming hairier and sometimes showing a
black dot in each side on the boundary notum/scutellum.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
37
3.3. RNA SAMPLES FOR MICROARRAYS
3.3.1. QUALITATIVE AND QUANTITATIVE ANALYSIS OF
THE EXTRACTED RNA
3.3.1.1. NANODROP ANALYSIS
In order to perform a microarray experiment we extracted RNA samples from
wing imaginal discs of several genotypes described above. Three separate
experiments were carried out for ach genotype. After extraction, we resorted to a
Nanodrop analysis to assess the quality of our samples for microarray analysis. All
samples provided an RNA concentration higher than 50ng/ul. The first three samples,
A, B and C (Table III.1) were very close in terms of concentration, however sample D
shows higher values due to the increased size, and consequent higher cell number, of
the wing imaginal discs in this genotype. The purity of the RNA shows was assessed
by Nanodrop showing an average ratio of 2,05 (260/280 nm). Further results using a
bioanalyzer helped to assure us of RNA quality despite the lower values of 260/230
ratio.
Table III.1 - Nanodrop quantifications in ng/ul of four different samples with three repetitions each, Absorbance at 260
nm, 280 nm, 230 nm, the ratio between 260 and 280 and the ratio between 260 and 230.
Genotype Sample
ID ng/ul A260 A280 260/280 260/230
Ap-Gal4>UAS-GFP/CyO A1 51,92 1,298 0,766 1,7 0,14
Ap-Gal4>UAS-GFP/CyO A2 97,34 2,433 1,159 2,1 0,39
Ap-Gal4>UAS-GFP/CyO A3 101,8 2,545 1,227 2,07 1,55
Ap-Gal4>UAS-P35/CyO B1 76,26 1,906 1,017 1,87 0,34
Ap-Gal4>UAS-P35/CyO B2 144,7 3,618 1,721 2,1 1,44
Ap-Gal4>UAS-P35/CyO B3 158,63 3,966 1,864 2,13 0,77
Ap-Gal4>UAS-GFP UAS-Bub3RNAi
/CyO C1 50,83 1,271 0,66 1,93 0,16
Ap-Gal4>UAS-GFP UAS-Bub3RNAi
/CyO C2 125,2 3,13 1,492 2,1 2,06
Ap-Gal4>UAS-GFP UAS-Bub3RNAi
/CyO C3 59,31 1,483 0,787 1,88 0,29
Ap-Gal4>UAS-GFP UAS-Bub3RNAi
/UAS-P35 D1 310,31 7,758 3,533 2,2 2,21
Ap-Gal4>UAS-GFP UAS-Bub3RNAi
/UAS-P35 D2 749,58 18,74 8,373 2,24 2,25
Ap-Gal4>UAS-GFP UAS-Bub3RNAi
/UAS-P35 D3 894,69 22,367 9,551 2,34 2,97
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
38
3.3.1.2. BIOANALYZER ANALYSIS
The isolation of RNA from biological samples is a difficult process, hampered by
the action of ribonucleases within the samples. As such, RNA integrity is usually the
main concern upon RNA sample preparation (177). Generally, the certification of RNA
integrity is usually verified through an electrophoresis gel, monitoring the bands of
rRNA species since they include more than 90 % of the mass of total RNA. The
difficulty arises when there is a continued development and application of gene
expression studies based on quantitative, real-time RT-PCR and microarrays, which
require consistent and highly reproducible technologies (Comparing performance of the
Agilent 2100 Bioanalyzer bioanalyzer to traditional RNA analysis techniques, Agilent
Application Note, 2000). The method for these studies is a chip-based nucleic acid
analysis system, the Agilent 2100 bioanalyzer, where Pre-packed kits, standardized
sample preparation and automated analysis lessen manual interference yielding more
accurate and reproducible data requiring only 25 ng of total RNA (Comparing
performance of the Agilent 2100 Bioanalyzer bioanalyzer to traditional RNA analysis
techniques, Agilent Application Note, 2000). However, new technology is also available
that allows for a detection limit around 200 pg, specially designed for laser
microdissection technologies (Quality assurance of RNA derived from laser
microdissected tissue samples obtained by the PALM MicroBeam system using the
RNA 6000 Pico LabChip kit, Agilent Application Note, 2003). The very first indication of
good quality of total RNA samples when performing the evaluation of bioanalizer
electropherogram is the identification of well-defined ribosomal RNAs (rRNAs) peaks
(Comparing performance of the Agilent 2100 Bioanalyzer bioanalyzer to traditional
RNA analysis techniques, Agilent Application Note, 2000). These peaks however, are
species-dependent, therefore, they are prone to variance between them. Another
bioanalyzer feature involves the addition to all samples of a reference marker
compound with a fast migration so the software is able to align all electropherograms
within one LabChip run. Furthermore, the software can also automatically search and
identify peaks for the two most abundant large rRNAs (always labelled as 18S and
28S), and calculate peak area ratios. In Drosophila melanogaster however 28S rRNA is
usually split in two fragments, comigrating with 18S rRNA. At last, aside from rRNA
peaks the intensity of any other signal should be very low (Stringent RNA quality
control using the Agilent 2100 bioanalyzer, Agilent Technologies, 2005).
Regarding our analysis, the software available did not have the ability to recognize
Drosophila melanogaster rRNA peaks so we could only verify the definition of rRNA
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
39
peaks and concentration through the determination of the area under the entire RNA
electropherogram. The ladder supplies a concentration/area ratio that is applied to
convert the sample area into concentration values (Figure III.3). For the RNA extracted
from larvae expressing only GFP, the first sample electropherogram showed an area of
423,691 resulting in a concentration of 243,997 ng/µl, the second sample presented an
area of 466,238 resulting in a concentration of 268,499 ng/µl and the third an area of
20,896 resulting in a concentration of 12,034 ng/µl (Figure III.4). For the RNA extracted
from larvae expressing only p35, the first sample electropherogram demonstrated an
area of 676,874 resulting in a concentration of 389,802 ng/µl, the second showed an
area of 765,336 resulting in a concentration of 440,745 ng/µl and the third an area of
160,088 resulting in a concentration of 92,192 ng/µl (Figure III.5). For the RNA
extracted from larvae knocked down for Bub3 and expressing GFP, the first sample
electropherogram demonstrated an area of 672,095 resulting in a concentration of
387,049 ng/µl, the second showed an area of 193,792 resulting in a concentration of
111,602 ng/µl and the third an area of 116,453 resulting in a concentration of 67,063
ng/µl (Figure III.6). For RNA extracted from larvae knocked down for Bub3 expressing
GFP, and p35, the first repeat electropherogram presented an area of 2726,804
resulting in a concentration of 1570,324 ng/µl, the second an area of 2214,551
resulting in a concentration of 1275,326 ng/µl and the third area of 1119,473 resulting
in a concentration of 644,687 ng/µl (Figure III.7). Through this technology we also have
access to the gel photo where we ran our samples (Figure III.8).
Figure III.3 - Graphic representing the fluorescence peaks of the RNA ladder. [FU] represents fluorescence units
and [s] represents time in seconds.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
40
Figure III.4 - Agilent Bioanalyzer electropherograms
showing rRNA peaks of RNA extract from wing
imaginal discs of 3rd
instar larvae expressing GFP.
[FU] represents fluorescence units and [s] represents
time in seconds. Graph a) represents the bioanalyzer
electropherograms of the first repeat of RNA extract from
wing imaginal discs of 3rd instar larvae expressing GFP. It
shows well-defined rRNA peaks (28s rRNA at
approximately 41,8 seconds and 18s rRNA at
approximately 43,2 seconds) with an area of 423,691
resulting in a concentration of 243,997 ng/µl. Graph b)
represents the bioanalyzer electropherograms of the
second repeat of RNA extract from wing imaginal discs of
3rd instar larvae expressing GFP. It shows well-defined
rRNA peaks (28s rRNA at approximately 41,8 seconds and
18s rRNA at approximately 43,2 seconds) with an area of
466,238 resulting in a concentration of 268,499 ng/µl.
Graph c) represents the bioanalyzer electropherograms of
the third repeat of RNA extract from wing imaginal discs of
3rd instar larvae expressing GFP. It shows well-defined
rRNA peaks (28s rRNA at approximately 41,8 seconds and
18s rRNA at approximately 43,2 seconds) however, with a
smaller area of 20,896 resulting in a lower concentration of
12,034 ng/µl.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
41
Figure III.5 - Agilent Bioanalyzer electropherograms
showing rRNA peaks of RNA extract from wing
imaginal discs of 3rd
instar larvae expressing P35. [FU]
represents fluorescence units and [s] represents time
in seconds. Graph a) represents the bioanalyzer
electropherograms of the first repeat of RNA extract from
wing imaginal discs of 3rd instar larvae expressing P35. It
shows well-defined rRNA peaks (28s rRNA at
approximately 41,8 seconds and 18s rRNA at
approximately 43,2 seconds) with an area of 676,874
resulting in a concentration of 389,802 ng/µl. Graph b)
represents the bioanalyzer electropherograms of the
second repeat of RNA extract from wing imaginal discs of
3rd instar larvae expressing P35. It shows well-defined
rRNA peaks (28s rRNA at approximately 41,8 seconds and
18s rRNA at approximately 43,2 seconds) with an area of
765,336 resulting in a concentration of 440,745 ng/µl.
Graph c) represents the bioanalyzer electropherograms of
the third repeat of RNA extract from wing imaginal discs of
3rd instar larvae expressing P35. It shows well-defined
rRNA peaks (28s rRNA at approximately 41,8 seconds and
18s rRNA at approximately 43,2 seconds) however, with a
smaller area of 160,088 resulting in a lower concentration
of 92,192 ng/µl.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
42
Figure III.6 - Agilent Bioanalyzer electropherograms
showing rRNA peaks of RNA extract from wing
imaginal discs of 3rd
instar larvae expressing GFP and
Bub3RNAi
. [FU] represents fluorescence units and [s]
represents time in seconds. Graph a) represents the
bioanalyzer electropherograms of the first repeat of RNA
extract from wing imaginal discs of 3rd instar larvae
expressing GFP, p35 and Bub3RNAi
. It shows well-defined
rRNA peaks (28s rRNA at approximately 41,8 seconds
and 18s rRNA at approximately 43,2 seconds) with an
area of 672,095 resulting in a concentration of 387,049
ng/µl. Graph b) represents the bioanalyzer
electropherograms of the second repeat of RNA extract
from wing imaginal discs of 3rd instar larvae expressing
GFP, p35 and Bub3RNAi
. It shows well-defined rRNA peaks
(28s rRNA at approximately 41,8 seconds and 18s rRNA
at approximately 43,2 seconds) with an area of 193,792
resulting in a concentration of 111,602 ng/µl. Graph c)
represents the bioanalyzer electropherograms of the third
repeat of RNA extract from wing imaginal discs of 3rd
instar larvae expressing GFP, p35 and Bub3RNAi
. It shows
well-defined rRNA peaks (28s rRNA at approximately 41,8
seconds and 18s rRNA at approximately 43,2 seconds)
with an area of 116,453 resulting in a concentration of
67,063 ng/µl.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
43
Figure III.7 - Agilent Bioanalyzer electropherograms
showing rRNA peaks of RNA extract from wing
imaginal discs of 3rd
instar larvae expressing GFP,
Bub3RNAi
and p35. [FU] represents fluorescence units
and [s] represents time in seconds. Graph a)
represents the bioanalyzer electropherograms of the first
repeat of RNA extract from wing imaginal discs of 3rd
instar larvae expressing GFP and Bub3RNAi
. It shows well-
defined rRNA peaks (28s rRNA at approximately 41,8
seconds and 18s rRNA at approximately 43,2 seconds)
with an area of 2726,804 resulting in a concentration of
1570,324 ng/µl. Graph b) represents the bioanalyzer
electropherograms of the second repeat of RNA extract
from wing imaginal discs of 3rd instar larvae expressing
GFP and Bub3RNAi
. It shows well-defined rRNA peaks
(28s rRNA at approximately 41,8 seconds and 18s rRNA
at approximately 43,2 seconds) with an area of 2214,551
resulting in a concentration of 1275,326 ng/µl. Graph c)
represents the bioanalyzer electropherograms of the third
repeat of RNA extract from wing imaginal discs of 3rd
instar larvae expressing GFP and Bub3RNAi
. It shows well-
defined rRNA peaks (28s rRNA at approximately 41,8
seconds and 18s rRNA at approximately 43,2 seconds)
with an area of 1119,473 resulting in a concentration of
644,687 ng/µl.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
44
Figure III.8 - Electrophoresis gel with each repeat of each sample used in microarrays.
3.4. MICROARRAY ANALYSIS
3.4.1. MICROARRAY DATA
After verifying that the integrity and concentration of the samples, we proceeded to
perform a microarray protocol. Upon completion, validation and normalization (data not
shown) of the different hybridization, we resorted to the software MeV v4.6.1 to analyze
and compare the retrieved data. We did the following comparisons through t-test
(Figure III.9):
1) Expression profiles of flies expressing GFP were compared with flies
expressing GFP and Bub3RNAi. (annex 2)
2) Expression profiles of flies expressing GFP and Bub3RNAi were compared with
flies expressing GFP, Bub3RNAi and p35. (annex 3)
3) Expression profiles of flies expressing GFP were compared with flies
expressing GFP, Bub3RNAi and p35. (annex 4)
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
45
4) Expression profiles of flies expressing p35 were compared with flies expressing
GFP, Bub3RNAi and p35. (annex 5)
5) Expression profiles of flies expressing GFP were compared with flies
expressing p35. (annex 6)
Each comparison provided us a list with a large amount of genes differently expressed
between two genotypes. Therefore, to reduce the number of genes and obtain an
enhanced biological meaning we carried out t-tests choosing a p-value of 0,05,
allowing us to compare the distinct samples in terms of different expression profiles.
Analysis of these lists also required the choice of a cut-off fold in terms of differences in
expression. The selected fold was two-fold, meaning that we considered a gene
whenever the difference in expression in distinct samples was at least two fold, either
up or down-regulated. To illustrate the expression profiles and their cut-offs the
software provided a volcano plot for each comparison. Genes classified with the
desired cut off, 0,05 p-value and fold difference superior to two, are represented by
green dots. Red dots on the other hand represent genes with a p-value of at least 0,01
and a fold difference of less than two. This notation helps to understand the amount of
information lost and how it could be relevant. A closer look to the volcano plots
tendency reveals that flies expressing GFP and Bub3RNAi or flies expressing only p35
do not show a large amount of genes differently expressed when compared to the
control, flies expressing only GFP, with 1005 and 1096 genes respectively (Figure
III.10 and III.11 respectively). However, whenever the comparison involves flies with
the overgrowth phenotype, expressing simultaneously GFP, Bub3RNAi, and p35, the
amount of green dots and therefore differently expressed genes with a p-value of 0,05
and a fold superior to two increases drastically. This can be perceived when the
comparison involves flies expressing only GFP with 4556 genes (Figure III.12), flies
expressing only p35 with 5460 genes (Figure III.13) or flies expressing GFP and
Bub3RNAi with 3909 genes (Figure III.14).
Figure III.9 - Schematic representation of the t-test comparisons (represented by a black arrow) made with the
microarray output.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
46
Figure III.10 - Volcano plot
comparing expression
profiles from microarray
output and selected genes
according to P-value
(represented in the vertical
axis by -log10(p-value) and
fold difference (represented in the horizontal axis). Expression profiles compared were Group B - tissue expressing
GFP and Bub3RNAi
(Ap-Gal4>UAS-GFP UAS-Bub3RNAi
) and Group A - tissue expressing GFP (Ap-Gal4>UAS-GFP).
Green dots represent genes with a fold induction superior equal to 2 and -2 and p-value superior or equal to 0,05. Red
dots represent genes with a fold induction difference lower to 2 and -2 and p-value superior or equal to 0,05.
Figure III.11 - Volcano plot
comparing expression
profiles from microarray
output and selected genes
according to P-value
(represented in the vertical
axis by -log10(p-value) and
fold difference (represented in the horizontal axis). Expression profiles compared were Group B - tissue expressing
P35 (Ap-Gal4>UAS-p35) Group A - tissue expressing GFP (Ap-Gal4>UAS-GFP). Green dots represent genes with a
fold difference higher or equal to 2 and -2 and p-value higher or equal to 0,05. Red represents genes with a fold
induction difference lower to 2 and -2 and p-value higher or equal to 0,05.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
47
Figure III.12 - Volcano plot
comparing expression
profiles from microarray
output and selected genes
according to P-value
(represented in the vertical
axis by -log10(p-value) and
fold difference (represented in the horizontal axis). Expression profiles compared were Group B - tissue expressing
GFP, Bub3RNAi
and P35 (Ap-Gal4>UAS-GFP UAS- Bub3RNAi
UAS-p35) Group A - tissue expressing GFP (Ap-
Gal4>UAS-GFP). Green dots represent genes with a fold difference higher or equal to 2 and -2 and p-value higher or
equal to 0,05. Red represents genes with a fold induction difference lower to 2 and -2 and p-value higher or equal to
0,05.
Figure III.13 - Volcano plot
comparing expression
profiles from microarray
output and selected genes
according to P-value
(represented in the vertical
axis by -log10(p-value) and
fold difference
(represented in the horizontal axis). Expression profiles compared were Group B - tissue expressing GFP, Bub3RNAi
and P35 (Ap-Gal4>UAS-GFP UAS- Bub3RNAi
UAS-p35) Group A - tissue expressing P35 (Ap-Gal4>UAS-p35). Green
dots represent genes with a fold difference higher or equal to 2 and -2 and p-value higher/equal to 0,05. Red dots
represent genes with a fold induction difference lower to 2 and -2 and p-value higher or equal to 0,05.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
48
Figure III.14. Volcano plot
comparing expression
profiles from microarray
output and selected genes
according to P-value
(represented in the vertical
axis by -log10(p-value) and
fold difference (represented in the horizontal axis). Expression profiles compared were Group B - tissue expressing
GFP, Bub3RNAi
and P35 (Ap-Gal4>UAS-GFP UAS- Bub3RNAi
UAS-p35) Group A - tissue expressing GFP and Bub3RNAi
(Ap-Gal4>UAS-GFP UAS- Bub3RNAi
). Green dots represent genes with a fold difference higher or equal to 2 and -2 and
p-value higher or equal to 0,05. Red dots represent genes with a fold induction difference lower to 2 and -2 and p-value
higher or equal to 0,05.
3.4.2. MINING THE MICROARRAY DATA TO ACHIVE
RELEVANT BIOLOGICAL INFORMATION
The t-tests performed helped narrow the search for genes of biological interest,
however, the amount of genes is still too large to be manageable for a manual search
of genes of interest. Therefore, we resorted to another technique to reduce the list of
genes to a convenient yet relevant lower number. To achieve this we resorted to Venn
diagrams. The underlying principle behind the choices made lays in the selection of
genes that respond to a specific component. The focus was to understand what might
be influencing the overgrowth of the wing imaginal discs when the simultaneous
expression of Bub3RNAi and p35 occurs. Therefore, revolving around that genotype we
were able to distinguish three components. Genes either upregulated or downregulated
in the wing imaginal disc overgrowth could be a consequence of either the Bub3
knockdown component or p35 component or even the component revealing the
synergy between the two, i.e. the gene expression is only altered upon the synergistic
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
49
effect of Bub3RNAi and p35. Therefore, we made intersections with the lists using the
venn diagrams (Figures III.15 to III.20). The upregulated Bub3RNAi component (annex 7)
can be obtained using the upregulated group of genes in flies expressing Bub3RNAi, p35
and GFP that are not upregulated in the group of genes of flies expressing only GFP
(Figure III.15 blue circle). Subsequently, intersection with the upregulated gene group
of flies expressing Bub3RNAi, p35 and GFP that are not upregulated in the group of
genes of flies expressing only p35, removed this component (Figure III.15 green circle).
The red circle in Figure III.15 allows for the removal of the upregulated gene group
resulting from the effect of p35 alone by removing the group of genes upregulated in
flies expressing p35 that are not upregulated in flies expressing only GFP (figure III.15
Red circle). The amount of genes representing this upregulated component is 663 and
it is represented in Figure III.15 inside the yellow box. The downregulated component
(annex 8) rationale is exactly the same as for the upregulated one, using the
downregulated genes (Figure III.16). The amount of genes representing this
downregulated component is 351 and it is represented in Figure III.16 inside the yellow
box. The upregulated p35 component (annex 9) can be obtained using the upregulated
group of genes in flies expressing Bub3RNAi, p35 and GFP that are not upregulated in
the group of genes of flies expressing only GFP (Figure III.17 blue circle).
Subsequently, intersect it with the upregulated gene group of flies expressing Bub3RNAi,
p35 and GFP that are not upregulated in the group of genes of flies expressing only
Bub3RNAi, therefore removing this component (Figure III.17 green circle). The red circle
in Figure III.17 allows for the removal of the upregulated gene group resulting from the
effect of p35 alone by removing the group of genes upregulated in flies expressing p35
that are not upregulated in flies expressing only GFP. The amount of genes
representing this upregulated component is 540 and it is represented in Figure III.17
inside a yellow box. The downregulated component (annex 10) rationale is the exact
same as for the upregulated one but using instead downregulated genes (Figure III.18).
The amount of genes representing this downregulated component is 362 and it is
represented in Figure III.18 inside a yellow box.
The upregulated synergy component (annex 11) can be obtained using the
upregulated group of genes in flies expressing Bub3RNAi, p35 and GFP that are not
upregulated in the group of genes of flies expressing only GFP (Figure III.19 blue
circle). Afterwards, this list is intersected with the upregulated gene group of flies
expressing Bub3RNAi, p35 and GFP that are not upregulated in the group of genes of
flies expressing only p35, therefore removing this component (Figure III.19 green
circle). A final intersection with the upregulated gene group of flies expressing
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
50
Bub3RNAi, p35 and GFP that are not upregulated in the group of genes of flies
expressing only Bub3RNAi, removes this component (Figure III.19 red circle). The
remaining of genes are only differently expressed when Bub3RNAi and p35 are both
present. The amount of genes representing this upregulated component is 181 and it is
represented in Figure III.19 inside the yellow box. The downregulated component
(annex 12) rationale is the exactly the same as for the upregulated one using instead
the list for the downregulated genes (Figure III.20). The amount of genes representing
this downregulated component is 267 and it is represented in Figure III.20 inside the
yellow box.
Figure III.15 - Venn diagram. Blue circle represents genes
upregulated in tissue expressing GFP, Bub3RNAi
and P35 (Ap-
Gal4>UAS-GFP UAS-Bub3RNAi
UAS-p35) when compared with tissue
expressing GFP (Ap-Gal4>UAS-GFP); green circle represents genes
upregulated in tissue expressing GFP, Bub3RNAi
and P35 (Ap-
Gal4>UAS-GFP UAS-Bub3RNAi
UAS-p35) when compared with tissue
expressing P35 (Ap-Gal4>UAS-p35); red circle represents genes
upregulated in tissue expressing P35 (Ap-Gal4>UAS-p35) when
compared with tissue expressing GFP (Ap-Gal4>UAS-GFP); inside
the yellow box is the number of genes in common in the upregulated
portion of tissue expressing GFP, Bub3RNAi
and P35 (Ap-Gal4>UAS-
GFP UAS-Bub3RNAi
UAS-p35) when compared with tissue expressing GFP (Ap-Gal4>UAS-GFP) and the upregulated
portion of tissue expressing GFP, Bub3RNAi
and P35 (Ap-Gal4>UAS-GFP UAS-Bub3RNAi
UAS-p35) when compared
with tissue expressing P35 (Ap-Gal4>UAS-p35).
Figure III.16 - Venn diagram. Blue circle represents genes
downregulated in tissue expressing GFP, Bub3RNAi
and P35 (Ap-
Gal4>UAS-GFP UAS-Bub3RNAi
UAS-p35) when compared with
tissue expressing GFP (Ap-Gal4>UAS-GFP); green circle
represents genes downregulated in tissue expressing GFP,
Bub3RNAi
and P35 (Ap-Gal4>UAS-GFP UAS-Bub3RNAi
UAS-p35)
when compared with tissue expressing P35 (Ap-Gal4>UAS-p35);
red circle represents genes downregulated in tissue expressing P35
(Ap-Gal4>UAS-p35) when compared with tissue expressing GFP
(Ap-Gal4>UAS-GFP); inside the yellow box is the number of genes
in common in the downregulated portion of tissue expressing GFP, Bub3RNAi
and P35 (Ap-Gal4>UAS-GFP UAS-
Bub3RNAi
UAS-p35) when compared with tissue expressing GFP (Ap-Gal4>UAS-GFP) and the downregulated portion of
tissue expressing GFP, Bub3RNAi
and P35 (Ap-Gal4>UAS-GFP UAS-Bub3RNAi
UAS-p35) when compared with tissue
expressing P35 (Ap-Gal4>UAS-p35).
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
51
Figure III.17 - Venn diagram. Blue circle represents genes
upregulated in tissue expressing GFP, Bub3RNAi
and P35 (Ap-
Gal4>UAS-GFP UAS-Bub3RNAi
UAS-p35) when compared with in
tissue expressing GFP (Ap-Gal4>UAS-GFP); green circle represents
genes upregulated in tissue expressing GFP, Bub3RNAi
and P35 (Ap-
Gal4>UAS-GFP UAS-Bub3RNAi
UAS-p35) when compared with tissue
expressing GFP and Bub3RNAi
(Ap-Gal4>UAS-GFP UAS-Bub3RNAi
); red
circle represents genes upregulated in tissue expressing P35 (Ap-
Gal4>UAS-p35) when compared with tissue expressing GFP (Ap-
Gal4>UAS-GFP); inside the yellow box is the number of genes in common in the upregulated portion in tissue
expressing GFP, Bub3RNAi
and P35 (Ap-Gal4>UAS-GFP UAS-Bub3RNAi
UAS-p35) when compared with tissue
expressing GFP (Ap-Gal4>UAS-GFP) and the upregulated portion in tissue expressing GFP, Bub3RNAi
and P35 (Ap-
Gal4>UAS-GFP UAS-Bub3RNAi
UAS-p35) when compared with tissue expressing GFP and Bub3RNAi
(Ap-Gal4\>UAS-
GFP UAS-Bub3RNAi
).
Figure III.18 - Venn diagram. Blue circle represents genes
downregulated in tissue expressing GFP, Bub3RNAi
and P35 (Ap-
Gal4>UAS-GFP UAS-Bub3RNAi
UAS-p35) when compared with in
tissue expressing GFP (Ap-Gal4>UAS-GFP); green circle represents
genes downregulated in tissue expressing GFP, Bub3RNAi
and P35 (Ap-
Gal4>UAS-GFP UAS-Bub3RNAi
UAS-p35) when compared with tissue
expressing GFP and Bub3RNAi
(Ap-Gal4>UAS-GFP UAS-Bub3RNAi
); red
circle represents genes downregulated in tissue expressing P35 (Ap-
Gal4>UAS-p35) when compared with tissue expressing GFP (Ap-
Gal4>UAS-GFP); inside the yellow box is the number of genes in
common in the downregulated portion in tissue expressing GFP, Bub3RNAi
and P35 (Ap-Gal4>UAS-GFP UAS-Bub3RNAi
UAS-p35) when compared with tissue expressing GFP (Ap-Gal4>UAS-GFP) and the downregulated portion in tissue
expressing GFP, Bub3RNAi
and P35 (Ap-Gal4>UAS-GFP UAS-Bub3RNAi
UAS-p35) when compared with tissue
expressing GFP and Bub3RNAi
(Ap-Gal4>UAS-GFP UAS-Bub3RNAi
).
Figure III.19 - Venn diagram. Blue circle represents genes upregulated
in in tissue expressing GFP, Bub3RNAi
and P35 (Ap-Gal4>UAS-GFP
UAS-Bub3RNAi
UAS-p35) when compared with tissue expressing GFP
(Ap-Gal4>UAS-GFP); green circle represents genes upregulated in
tissue expressing GFP, Bub3RNAi
and P35 (Ap-Gal4>UAS-GFP UAS-
Bub3RNAi
UAS-p35) when compared with tissue expressing GFP and
Bub3RNAi
(Ap-Gal4>UAS-GFP UAS-Bub3RNAi
); red circle represents
genes upregulated in tissue expressing GFP, Bub3RNAi
and P35 (Ap-
Gal4>UAS-GFP UAS-Bub3RNAi
UAS-p35) when compared with tissue
expressing P35 (Ap-Gal4>UAS-p35); inside the yellow box is the
number of genes of the unregulated portion in tissue expressing GFP,
Bub3RNAi
and P35 (Ap-Gal4>UAS-GFP UAS-Bub3RNAi
UAS-p35) when compared with tissue expressing GFP (Ap-
Gal4>UAS-GFP) however not present in neither the upregulated portion in tissue expressing GFP, Bub3RNAi
and P35
(Ap-Gal4>UAS-GFP UAS-Bub3RNAi
UAS-p35) when compared with tissue expressing GFP and Bub3RNAi
(Ap-
Gal4>UAS-GFP UAS-Bub3RNAi
), nor the upregulated portion in tissue expressing GFP, Bub3RNAi
and P35 (Ap-
Gal4>UAS-GFP UAS-Bub3RNAi
UAS-p35) when compared with in tissue expressing P35 (Ap-Gal4>UAS-p35).
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
52
Figure III.20 - Venn diagram. Blue circle represents genes
downregulated in in tissue expressing GFP, Bub3RNAi
and P35 (Ap-
Gal4>UAS-GFP UAS-Bub3RNAi
UAS-p35) when compared with tissue
expressing GFP (Ap-Gal4>UAS-GFP); green circle represents genes
downregulated in tissue expressing GFP, Bub3RNAi
and P35 (Ap-
Gal4>UAS-GFP UAS-Bub3RNAi
UAS-p35) when compared with tissue
expressing GFP and Bub3RNAi
(Ap-Gal4>UAS-GFP UAS-Bub3RNAi
); red
circle represents genes downregulated in tissue expressing GFP,
Bub3RNAi
and P35 (Ap-Gal4>UAS-GFP UAS-Bub3RNAi
UAS-p35) when
compared with tissue expressing P35 (Ap-Gal4>UAS-p35); inside the
yellow box is the number of genes of the downregulated portion in
tissue expressing GFP, Bub3RNAi
and P35 (Ap-Gal4>UAS-GFP UAS-Bub3RNAi
UAS-p35) when compared with tissue
expressing GFP (Ap-Gal4>UAS-GFP) however not present in neither the downregulated portion in tissue expressing
GFP, Bub3RNAi
and P35 (Ap-Gal4>UAS-GFP UAS-Bub3RNAi
UAS-p35) when compared with tissue expressing GFP and
Bub3RNAi
(Ap-Gal4>UAS-GFP UAS-Bub3RNAi
), nor the downregulated portion in tissue expressing GFP, Bub3RNAi
and
P35 (Ap-Gal4>UAS-GFP UAS-Bub3RNAi
UAS-p35) when compared with in tissue expressing P35 (Ap-Gal4>UAS-p35).
3.4.3. VALIDATING VENN DIAGRAMS THROUGH
BIOINFORMATICS RESOURCES
Resorting to Venn Diagrams was the final bioinformatics approach to attempt to
turn the expression profiles more meaningful and manageable. Therefore, before
moving forward we attempted to understand if we were successful in our selections.
We decided to use DAVID bioinformatics resources 6.7. This resource refers to an
integrated biological knowledgebase and the analytic tools try to systematically extract
biological significance from large gene/protein lists (178). With DAVID, it is possible to
analyze gene lists resultant from high-throughput and integrated data-mining
environment. Initially, the gene list of interest is uploaded, holding any number of
common gene identifiers, and then analyzed with any of the desired pathway-mining
tool such as gene functional classification, functional annotation chart or clustering and
functional annotation table (178). However, due to the redundancy of biological
annotations, tools like the Functional Annotation Chart have the drawback of showing
analogous and significant annotations repeatedly. Such event weakens the focus on
biology. In order to diminish the redundancy, DAVID researchers have developed the
Functional Annotation Clustering tool that provides a clearer biology reading. The
hypothesis sustaining the grouping algorithm implies that closely related annotations
should have similar gene members. The Functional Annotation Clustering tool uses the
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
53
same techniques of Kappa statistics in order to measure the extent of common genes
between two annotations. Then with fuzzy heuristic clustering already used in Gene
Functional Classification Tool, it proceeds to classify the groups of similar annotations
according to the calculated kappa values (160). This technique leads to a higher
probability of groups being clustered together the more common genes annotations
they share. Annotation P-values integrated in each cluster possess the exact same
meaning as a p-value shown in a normal chart report for the same terms. However,
DAVID bioinformatics resources 6.7 uses a system called Group Enrichment Score
where they rank biological relevance according to the geometric mean in logarithm
scale of member’s p-values of the corresponding annotation cluster (160). Upon
uploading our gene lists obtained through Venn diagram, we proceeded by performing
a Functional Annotation Clustering. The obtained information emerged as very long
spreadsheets in form of an excel file showing large clusters of closely related
annotations with a particular enrichment score. The selected cut off for enrichment
score was 1,3 and it means that the geometric mean of the different cluster members is
approximately 0,05 providing us with a confidence interval of 95%. Regarding the gene
lists it is also important to refer that the program can only use the genes with an
annotated function, therefore, many of our choices of unknown CG numbers cannot be
included in this analysis. However, it can still give a good idea from the choices that
were annotated. We uploaded six different gene lists, the upregulated and
downregulated of each component: Bub3 knockdown component, the p35 component
and the synergy component and looked for enriched pathways related with anything
that could influence cell cycle, cell division and apoptosis in general. Regarding the
upregulated Bub3 knockdown component, we found thirteen different clusters above
the chosen enrichment score cut-off (annex 13). From these different clusters there is
one of particular interest with an enrichment score of 1,74. Cluster members are
related to DNA repair, response to DNA damage stimulus, telomere maintenance, and
telomere organization among other annotations (Figure III.21). Searching the
downregulated Bub3 knockdown component, eight distinct clusters (annex 14) were
identified with values of enrichment score higher than the cut-off. Upon analyzing them,
we were not able to extract any relevance to the purpose of our work. Nevertheless,
one of the clusters showing an enrichment score of 1,84 and its members are related to
to cell motility which might be important (Figure III.22). Investigating the upregulated
p35 component, we found twelve different clusters above the chosen enrichment score
cut-off (annex15). However, none seems to be related in any way to cell cycle, cell
division or apoptosis (Figure III.23). Inspecting the downregulated p35 component led
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
54
to 17 clusters whose values of enrichment score are higher than the cut-off (annex 16).
Thorough analysis revealed no significant cluster to the purpose of our work (Figure
III.24). Should put the figures next to the analysis.
Regarding the upregulated synergy component, we found one cluster above the
chosen enrichment score cut-off (annex 17). This particular cluster shows an
enrichment score of 1,30 and its members are closely related to cell cycle process,
mitosis, spindle components among other similar annotations (graph not shown). The
downregulated synergy component presents three clusters with an enrichment score
higher than the imposed cut-off (annex 18). Regarding cell cycle, we could not find any
cluster with members related to this event (Figure III.25).
Figure III.21 - Graphic showing annotation clusters from the upregulated gene list from Bub3 knockdown component
with an enrichment score superior to 1,3. Obtained through a functional annotation clustering analysis using DAVID
bioinformatics resources 6.7.
Figure III.22 - Graphic showing annotation clusters from the downregulated gene list from Bub3 knockdown component
with an enrichment score superior to 1,3. Obtained through a functional annotation clustering analysis using DAVID
bioinformatics resources 6.7.
3,33 2,96
2,23 2,22 2,09 2,09 2,02 2,01 1,83 1,74
1,56 1,49 1,42
0
1
2
3
4
A1 A2 A3 A4 A5 A6 A7 A8 A9 A10 A11 A12 A13
en
rich
me
nt
sco
re
Upregulated Bub3 component
4,28
3,24 3,03
2,62 2,28
1,84 1,49 1,47
0
1
2
3
4
5
B1 B2 B3 B4 B5 B6 B7 B8
en
rich
me
nt
sco
re
Downregulated Bub3 component
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
55
Figure III.23 - Graphic showing annotation clusters from the upregulated gene list from p35 component with an
enrichment score superior to 1,3. Obtained through a functional annotation clustering analysis using DAVID
bioinformatics resources 6.7.
Figure III.24 - Graphic showing annotation clusters from the downregulated gene list from p35 component with an
enrichment score superior to 1,3. Obtained through a functional annotation clustering analysis using DAVID
bioinformatics resources 6.7.
Figure III.25 - Graphic showing annotation clusters from the downregulated gene list from p35 component with an
enrichment score superior to 1,3. Obtained through a functional annotation clustering analysis using DAVID
bioinformatics resources 6.7.
3,22
2,11 2,09 2,03 1,79 1,67 1,61 1,57 1,50 1,47 1,41 1,41
0
0,5
1
1,5
2
2,5
3
3,5
C1 C2 C3 C4 C5 C6 C7 C8 C9 C10 C11 C12
en
rich
me
nt
sco
re
Upregulated p35 component
4,99
4,12 3,67 3,61
2,56 2,45 2,31 2,02 2,01 1,95 1,90
1,60 1,60 1,59 1,58 1,42 1,38
0
1
2
3
4
5
6
D1 D2 D3 D4 D5 D6 D7 D8 D9 D10 D11 D12 D13 D14 D15 D16 D17
en
rich
me
nt
sco
re
Downregulated p35 component
2,39
1,38 1,34
0
0,5
1
1,5
2
2,5
3
F1 F2 F3
en
rich
me
nt
sco
re
Downregulated synergy component
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
56
3.4.4. MINING THE MICROARRAY DATA FROM RAW TO
BIOLOGICALLY RELEVANT.
Resorting to Venn diagrammes helped to slim the list of relevant genes to a
much more manageable amount. However, as previously mentioned, there was still a
great quantity of background noise, resulting from the use of complex larval tissue
where many developmental pathways are highly represented. Therefore, it was
required human input to search for biological pertinence. Such action required the
following set of criteria. Firstly, we elected genes due to their biological relevance
taking particular attention to known function, pathways, protein-protein interactions and
the existence of homologous in humans using for that mostly NCBI and Flybase
databases. Secondly, genes with unknown function were selected if the difference of
expression was superior/equal to ten. Thirdly, an abnormally high fold difference was
also taken into account even if it was not an unknown sequence. As the candidates
seemed promising we decided that it was worth moving forward so we built a final list
comprising 210 genes, 139 upregulated (Table III.2) and 71 downregulated (Table
III.3).
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
57
Table III.2 - Table showing upregulated genes manually searched and selected from the Venn diagram components and their fold difference.
Upregulated genes
fold fold fold fold fold fold
Prx5037 1733,3 CG32625 35,74 CG4250 18,69 CG14355 12,95 CG34430 10,27 Irbp 3,58
Kap3 589,66 CG13403 34,72 CG5866 18,67 CG30485 12,9 CG13954 10,09 Dhc62B 3,55
PGRP-SB2 521,04 CG5646 32,63 CG18193 17,59 CG13116 12,52 Thor 10,08 Cp110 3,47
CG8620 382,58 CG32085 31,72 CG13659 17,35 AcCoAS 12,36 CG14907 10,06 otu 3,33
CG9641 250,14 CG2157 30,23 CG15142 17,23 CG12520 12,15 alphaTub85E 9,97 PNUTS 3,18
asf1 132,27 CG18522 28,9 CG13026 17,21 CG15497 12,11 CG14523 9,78 Socs36E 3,11
mus304 122,55 CG15047 28,86 CG14059 16,71 CG6836 12,02 CG32335 9,73 CG1951 3,09
CG4089 117,8 CG13283 28,68 CG42369 16,12 Ance-5 11,92 CG17124 8,26 CG6333 3,05
CG14695 109,48 CG12699 28,38 Ddc 15,88 CG32681 11,76 CG10859 8,11 mre11 2,94
nimC4 83,96 CG3635 26,3 CG32368 15,86 CG14253 11,44 CG14216 8,04 Ku80 2,93
CG6983 82,26 CG5783 26,17 CG15279 15,46 CG11750 11,34 CG6830 7,06 CG6191 2,88
CG13043 57,1 CG33920 25,44 CG31686 15,4 nimB1 11,26 nimC1 6,93 CG1572 2,78
CG6347 50,99 dpr18 24,77 CG18063 15,01 CG13465 11,21 ST6Gal 6,66 puc 2,73
Osi6 47,19 CG13250 24,74 CG4267 15,01 CG32313 11 phr 6,15 CG18745 2,67
CG31436 43,24 CG31533 24,45 CG3746 14,61 CG34301 10,98 wun2 6,13 Rep 2,54
CG34437 41,67 CG3884 24,07 CG31370 14,22 CG6495 10,9 CG12200 6 Dhc36C 2,51
CG33926 39,72 CG31279 23,11 CG33178 14,14 CG32668 10,86 Traf4 5,56 ncd 2,43
CG32155 39,52 CG31219 23,1 CG8160 14,04 CG6118 10,6 moody 5,36 rad50 2,19
CG13541 39,02 CG12911 22,79 CG4362 13,91 CG14636 10,6 rhea 5,36 Alr 2,15
CG4367 38,05 CG11073 22,52 CG15784 13,25 Neu3 10,53 os 5,18 CG15528 2,14
CG12519 37,77 CG8193 21,27 CG4763 13,18 CG17325 10,49 Aplip1 4,9 CG13559 2,13
CG9517 37,1 CG17108 20,07 CG5399 13,06 CG6579 10,49 zormin 4,67 f-cup 2,1
CG11893 35,75 CG14321 19,01 dpr6 13,04 CG7120 10,45 wun 3,86 mus205 2,04
CG7188 2,03
Table III.3 - Table showing downregulated genes manually searched and selected from the Venn diagram
components and their fold difference.
Downregulated genes
fold fold fold fold fold
Gyc-89Da 569,06 CG15905 18,35 Skeletor 11,1 CG12071 5,9 CG7231 2,65
phr6-4 299,38 CG5157 17,64 CG31875 10,65 CG14915 5,77 CG16947 2,65
CG2663 56,94 CG17140 17,04 CG12105 10,63 Pkc53E 5,61 CG34380 2,5
CG14356 39,98 CG14958 16,91 Mdr50 10,53 CG15544 5,36 Cam 2,41
CG3355 28,51 CG14395 15,87 CG31676 10,53 CG13138 4,37 CG14222 2,35
CG13494 28,32 Spn28Db 15 B-H1 10,35 Damm 4,24 CG3769 2,35
CG13036 26,16 CG7906 14,16 CG34342 9,69 Top1 4,22 tws 2,23
Spn43Aa 23,8 by 13,75 CG16758 8,94 CG2556 4,18 rab3-GEF 2,23
CG8419 22,62 CG9416 12,54 CG5023 8,42 Alk 4,12 Roc1a 2,22
CG10317 20,76 CG13081 12,46 Phlpp 6,99 phyl 3,61 Wnt2 2,2
CG9238 20,3 CG13101 12,09 CG6287 6,97 sns 3,14 Drep-4 2,18
Spn28Da 20,29 CG3823 11,96 CG4586 6,78 Ror 3,03 Sin3A 2,09
CG12912 19,57 CG13408 11,84 C15 6,46 btl 2,93 PPP4R2r 2,06
CG31790 19,25 Spn100A 11,47 CanB 6,01 InR 2,81 Rapgap1 2,05
fz 2
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
58
3.5. THE ENHANCER/SUPPRESSION OF PHENOTYPE
SCREEN
The purpose of using a microarray approach and all the subsequent
bioinformatics steps was to be able to select a manageable list of candidate genes for
a relevant biological role in our tumorigenesis model. Then, using RNAi technology, we
performed a screen with in vivo RNAi for each of the chosen genes. Flies carrying
individual RNAis were crossed with either a fly knockdown for Bub3 (Ap-Gal4>UAS-
GFP UAS-Bub3RNAi/ CyO) or a fly knockdown for Bub3 but also expressing the
apoptosis inhibitor DIAP1 (Ap-Gal4>UAS-GFP UAS-Bub3RNAi UAS-DIAP1/ CyO). We
chose to overexpress DIAP1 using a GAL4-UAS system instead of P35 since the
phenotype could be analyzed in the adult (Figure III.26) (140). In addition, the
“intermediate” phenotype of the DIAP1, a fly presenting a larger, and generally
deformed thorax, with some loss of bristles, empty sockets and deformed wings
showing necrosis, would also allow us to look for suppression, as well as enhancement
of the phenotype. It is important to note that the candidates genes are upregulated or
downregulated when comparing Ap-Gal4>UAS-GFP UAS-Bub3RNAi/ UAS-p35 flies with
Ap-Gal4>UAS-GFP/ CyO regardless of which Venn diagram they were chosen.
Figure III.26 - Schematic representation of a
cross that enables the expression of a gene
of interest through the use of Gal4-UAS
system
We classified the phenotypes found as “unaltered”, “voluptuous thorax”, “double
hunchback thorax” and “lethal”. Unaltered (Ua) phenotype defines flies presenting a
phenotype closely related to the phenotype of the driver fly used in the cross, either
expressing Bub3RNAi or expressing Bub3RNAi together with DIAP1. Voluptuous thorax
(VT) phenotype consists of a tall and swollen thorax when compared with the
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
59
phenotype of the driver fly used in the cross, either expressing Bub3RNAi or expressing
Bub3RNAi together with DIAP1 (Figure III.27 - a); b); c)). When comparing to either
driver phenotypes, Double hunchback thorax (DHT) phenotype shows a tall and
swollen thorax with a division in the middle like two separate hunchbacks (Figure III.27
- d); e); f)). Lethal (Lt) phenotype as the name implies reports to the crosses that do
not originate viable adult flies.
Figure III.27 - Image depicting the different phenotypes we found in the screen. a) and b) represent the phenotype
classified as “Voluptuous thorax” (VT) under visible light. c) represents the phenotype classified as “Voluptuous
thorax” (VT) under GFP excitatory light. d) and e) represent the phenotype classified as “Double hunchback thorax”
(DHT) under visible light. f) represents the phenotype classified as “Double hunchback thorax” (DHT) under GFP
excitatory light.
We classified each phenotype found for the 1) RNAi for upregulated genes
crossed to the Bub3RNAi driver (Table T1), 2) RNAi for downregulated genes crossed to
the Bub3RNAi driver (Table T2), 3) RNAi for upregulated genes crossed to the Bub3RNAi
expressed together with DIAP1 driver (Table T3) and 4) RNAi for upregulated genes
crossed to the Bub3RNAi expressed together with DIAP1 driver (Table T4).
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
60
Table III.4 - Table showing RNAi screen results from upregulated genes crossed with the Bub3
RNAi.
Upregulated genes crossed with Bub3RNAi
Ua VT DHT Lt
CG4267 CG32681 f-cup PGRP-SB2 CG9641 CG6830 Rep
os CG6347 CG5399 CG14321 CG14907 CG18745 rhea
CG13026 CG6191 CG14523 CG8620 CG2157 CG5866 CG32335
CG14636 CG13116 CG14216 CG13659 nimB1 CG4089 CG13465
CG31370 CG15497 CG13954 CG4763 Dhc62B Cp110 CG6579
CG10859 CG17108 CG15784 CG12520 CG34430 CG31686 Neu3
Dhc36C nimC4 CG33178 CG31436 CG11750 CG18193 CG32668
CG7120 CG4250 CG32313 CG13250 CG3746 CG33920 mus205
mre11 CG9517 zormin CG12699 wun2 CG30485 PNUTS
CG32085 CG13043 CG5783 CG6836 nimC1 CG12911 asf1
CG6983 CG13559 CG15142 CG13541 Alr CG31533 ncd
CG42369 CG6118 CG11893 otu Aplip1 CG8193
Osi6 CG15047 CG34301 CG12519 CG1572 CG6333
dpr18 alphaTub85E CG15528 CG3884 Ddc CG31279
AcCoAS CG32625 CG17325 CG13283
ST6Gal CG32368 Prx5037 CG34437
CG18522 rad50 CG32155 CG14355
Thor CG11073 CG8160 CG1951
CG14695 dpr6 CG13403 CG7188
Traf4
Table III.5 - Table showing RNAi screen results from downregulated genes crossed with the Bub3RNAi
.
Downregulated genes crossed with Bub3RNAi
Ua VT DHT Lt
Ror CG9238 fz CG13494 Top1 CG34380 CG14958
InR CG6287 CG10317 phr6-4 CG15905 Spn100A Cam
CG13081 CG5157 Roc1a Skeletor CG4586 CG31676 tws
phyl CG12071 CG5023 Spn28Da Rapgap1 CG14222
CG34342 CG3355 CG7231 sns Pkc53E Gyc-89Da
Spn28Db Wnt2 C15 CG16758 Drep-4
CG17140 CG31875 CG8419 CG3769 CG13408
Mdr50 B-H1 CG13101 CG12912 CG13036
CanB Spn43Aa CG3823 btl Damm
CG14356 PPP4R2r CG16947 Phlpp
CG2556 CG31790
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
61
Table III.6 - Table showing RNAi screen results from upregulated genes crossed with Bub3
RNAi also expressing DIAP1.
Upregulated genes crossed with Bub3RNAi also expressing DIAP1
Ua VT DHT Lt
CG4267 CG15047 CG34301 os nimB1 CG33920 Rep
CG13026 alphaTub85E CG15528 CG31370 CG18745 wun2 rhea
CG14636 CG32625 CG17325 CG10859 CG5866 Prx5037 CG32335
Dhc36C CG32368 CG32155 CG32085 AcCoAS nimC1 CG13465
CG7120 rad50 CG8160 CG14907 ST6Gal CG8620 CG32668
mre11 CG11073 CG13403 CG6830 Thor PNUTS mus205
CG6983 dpr6 CG14321 CG2157 CG4089 CG12911 CG30485
CG9641 f-cup CG13659 CG15497 Cp110 CG4763 ncd
CG42369 CG5399 CG31436 CG11750 Dhc62B CG31533
Osi6 CG14523 CG12699 CG31686 CG6347 CG12520
dpr18 CG14216 CG6836 PGRP-SB2 CG6191 CG13250
CG18522 CG13954 otu asf1 CG13116 CG8193
CG14695 CG15784 CG12519 CG13541 CG6579 CG6333
CG32681 CG33178 CG3884 CG34437 CG34430 Alr
nimC4 CG32313 CG13283 Ddc CG17108 Aplip1
CG9517 zormin CG14355 CG4250 CG1572
CG13043 CG5783 CG1951 Neu3 CG31279
CG13559 CG15142 Traf4 CG3746 CG7188
CG6118 CG11893 CG18193
Table III.7 - Table showing RNAi screen results from downregulated genes crossed with Bub3RNAi
also expressing
DIAP1.
Downregulated genes crossed with Bub3RNAi and DIAP1
Ua VT DHT Lt
Ror CG5157 CG10317 Skeletor InR Top1 CG14958
Pkc53E CG3355 Roc1a Spn28Da CG13081 Spn100A Cam
phyl CG13408 CG5023 sns CG15905 CG6287 tws
CG34380 CG13036 C15 CG16758 CG4586 CG12071 Damm
CG17140 Wnt2 CG8419 CG3769 Rapgap1 CG13101 CG14222
Mdr50 CG31875 CG3823 CG12912 Drep-4 Gyc-89Da
CanB B-H1 CG16947 btl CG34342
CG14356 Spn43Aa CG2556 Phlpp Spn28Db
CG9238 PPP4R2r CG13494 CG31790 CG7231
CG31676 fz phr6-4
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
62
We tested 175 out of 210 candidates. We decided to analyze the data starting
with the phenotype:
Hyperplasia - Could indicate that tumor suppressors are downregulated and
oncogenes upregulated, which means RNAi for candidates will suppress growth if they
are oncogenes, increase growth if they are tumour suppressors. High levels of
apoptosis - could indicate that apoptosis promoters are upregulated, and apoptotic
suppressors downregulated. On the other hand if the thorax was bigger it could point to
apoptosis promoters being downregulated or apoptotic suppressors upregulated.
Therefore, RNAi for candidates that increase apoptosis will reduce apoptosis, the ones
that block apoptosis will have apoptosis increased. The cross-talk between the two
screens is possible when we can identify a gene in the screen with Bub3RNAi that can
“replace the DIAP effect” by being either a pro-apoptic gene RNAi will suppress
apoptosis) and mimic the growth phenotype or by being a suppressor of a suppressor
of apoptosis therefore giving the RNAi the same phenotype. This is also important, as
we can’t assume that the candidate gene is directly downstream of Bub3. Therefore,
when we find an enhanced phenotype with the downregulation of a candidate gene
through RNAi we are expecting it to have one of the following functions (table III.8):
1) It could be an upregulated tumor suppressor that downregulation prevents its
function and leads to cell proliferation;
2) It could be a downregulated tumor suppressor that further downregulation could
make it reach levels of translation that allow for an increased cell proliferation;
3) It could be an upregulated suppressor of an oncogene that downregulation
prevents its function allowing the oncogene to act and promote cell proliferation;
4) It could be a downregulated suppressor of an oncogene that further
downregulation prevents its function even more leading increase of the
oncogene function and consequent of promotion of cell proliferation;
5) It could be an upregulated activator of a tumor suppressor that downregulation
prevents its function impeding the tumor suppressor function and promoting cell
proliferation;
6) It could be a downregulated activator of a tumor suppressor that further
downregulation prevents its function not allowing the tumor suppressor to act
and therefore promoting cell proliferation;
7) It could be an upregulated pro-apoptotic gene that downregulation impedes
apoptosis therefore leading to an enhanced phenotype
8) It could be a downregulated pro-apoptotic gene that further downregulation
blocks apoptosis even more therefore leading to an enhanced phenotype
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
63
9) It could be an upregulated suppressor of an anti-apoptotic gene that
downregulation impedes apoptosis therefore leading to an enhanced phenotype
10) It could be a downregulated suppressor of an anti-apoptotic gene that furher
downregulation blocks apoptosis therefore leading to an enhanced phenotype
11) It could be an upregulated activator of a pro-apoptotic gene that downregulation
impedes apoptosis therefore leading to an enhanced phenotype
12) It could be a downregulated activator of a pro-apoptotic gene that further
downregulation blocks apoptosis even more therefore leading to an enhanced
phenotype
On the other hand, when we find a suppressed phenotype with the
downregulation of a candidate gene through RNAi we are expecting it to have one of
the following functions (table III.8):
1) It could be an upregulated oncogene that downregulation prevents its function
and does not lead to cell proliferation;
2) It could be a downregulated oncogene that further downregulation could make it
reach levels of translation that would not allow for an increased cell
proliferation;
3) It could be an upregulated suppressor of a tumor suppressor that
downregulation prevents its function allowing the tumor suppressor to act and
prevent cell proliferation;
4) It could be a downregulated suppressor of a tumor suppressor that further
downregulation prevents its function even more leading increase of the tumor
suppressor function and consequent of inhibition of cell proliferation;
5) It could be an upregulated activator of an oncogene that downregulation
prevents its function impeding the oncogene function not leading to cell
proliferation;
6) It could be a downregulated activator of an oncogene that further
downregulation prevents its function not allowing the oncogene to act and
therefore not leading to cell proliferation;
7) It could be an upregulated anti-apoptotic gene that downregulation promotes
apoptosis therefore leading to a supressed phenotype;
8) It could be a downregulated anti-apoptotic gene that further downregulation
promotes apoptosis even more therefore leading to a suppressed phenotype;
9) It could be an upregulated suppressor of a pro-apoptotic gene that
downregulation leads to apoptosis therefore leading to an enhanced phenotype
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
64
10) It could be an downregulated suppressor of a pro-apoptotic gene that further
downregulation leads to apoptosis therefore leading to an enhanced phenotype
11) It could be an upregulated activator of an anti-apoptotic gene that
downregulation lead to an increased apoptosis therefore leading to a
suppressed phenotype
12) It could be an downregulated activator of an anti-apoptotic gene that further
downregulation increases apoptosis even more therefore leading to a
suppressed phenotype
Table III.8 - Possible functions of candidate genes.
We did not observe any RNAis that suppressed the phenotypes, therefore we
focused on the enhanced phenotype. We then analyze each hit of the most enhanced
phenotype found, the “Double hunchback thorax”, looking for connections with
tumorigenesis, cell proliferation, cell cycle and related annotations
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
66
4.1. INTRODUCTION
Understanding tumorigenesis requires a systematic approach involving data
coming from different studies, from human cancer and from animal models. Regarding
the link between SAC, mitosis and tumorigenesis there seems to be a lot of valuable
data coming from in vitro studies. However, there is a need for in vivo studies using
animal models. Since little is known about Bub3 protein function and its connection with
tumorigenesis, we decided to investigate the link further by first using a high throughput
technology, microarrays. Resorting to microarrays allowed us to perform a large screen
for genes that might be involved in the hyperplastic growth transformation that is
observed when Bub3 is knockdown together with apoptosis inhibition.
4.2. REASONS FOR PHENOTYPE SELECTION
In order to move forward for a high throughput analysis we needed to establish
which flies we would collect wing imaginal discs from third instar larvae for RNA
extraction. This selection includes deciding which phenotypes are relevant for
understanding the overgrowth process and the required controls. We chose four
different phenotypes expressing the different transgenes in the wing imaginal disc. Our
main sample was the phenotype from Bub3 knockdown expressing GFP and p35 as it
has hyperplastic wing imaginal discs therefore serving as the main focus from which
we trimmed the large amount the genes differentially regulated into a manageable list
for further testing. With this phenotype, we wanted to answer one of the main questions
of the project regarding the consequences of impairing apoptosis using p35 when the
SAC protein Bub3 is depleted. To understand the role of Bub3 protein by itself and to
compare the expression profile it originates with the expression profile from the
overgrowth phenotype, we selected the Bub3 knockdown fly expressing GFP. This
particular genotype allowed us to follow the effects of having a fly with a SAC protein
depleted without interfering with apoptosis. Furthermore, comparing the transcriptional
profiles for these wing imaginal disc genotypes with and without p35, we expected to
decipher the changes in terms of transcription when introducing the apoptosis inhibition
factor. The presence of the heterologous P35 protein has been tested in the wing
imaginal disc and does not cause any significant alteration, except for a shortening of
wing vein 5 (135). The transcriptional profile from these flies allow for the identification
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
67
of what could be the effect due to preventing apoptosis or the effect of the synergy it
seems to have with SAC proteins. As we wanted to observe this, we also needed to
verify the consequences of inhibiting apoptosis alone. In order to accomplish that, we
selected a fly expressing the apoptosis inhibitor P35. We also needed an unaltered fly
in terms of SAC and apoptosis. For that purpose, we selected a fly expressing only
GFP. This fly allowed us to compare every other phenotype expression profile with a
normal condition profile and account for any change due to the expression of GFP
alone.
4.3. MICROARRAY OUTPUT REVEALS SIGNIFICANT
DIFFERENCE IN EXPRESSION PROFILES
DEPENDING ON GENOTYPES.
Data retrieved from microarrays from different genotypes was compared in
terms of p-value through t-test and fold difference using volcano plots. When choosing
a p-value for gene expression we considered and tested a p-value of 0,01, however the
amount of the information lost increased drastically. Therefore, we opted for lower 0,05
p-value granting us a confidence interval of 95% so we could reduce the data. Along
with the p-value, there was a need to define the fold difference between expression
profiles of different samples. The selected fold difference was 2 although we have seen
reports of lower values like 1,5 (160). We went for a fold difference of 2 since we were
not interested in very small changes and were expecting drastic changes to be the key.
However, the opposite might also be true since we are working with such important
cellular events, that a minimum change could influence, in theory, cells fate, and by
overlooking lower folds, we might have lost some valuable information. Also as this
was a first approach lower fold numbers would increase the number of genes
drastically and prevent a proper analysis. Observing the comparison between the
expression profiles of Bub3 knockdown fly with the control fly expressing only GFP we
detected one of the less spread volcano plot. Consequently, the number of genes
differently expressed are low. With this particular result, we can assume that the
depletion of Bub3 does not promote a major change in transcriptional profile when
compared to other genotypes. One possibility could be that RNAi is not uniform across
wing discs cells and therefore we are mostly analyzing those that survive and are not
significantly affected. The observation of a less spread volcano plot is also true when
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
68
comparing the expression profile of flies expressing p35 and control flies expressing
only GFP. Flies expressing p35 have their apoptosis inhibited however, SAC is still
functional, therefore the transcriptional profile is not dramatically altered. Moreover,
apoptosis in wild type discs is a rare phenomenon (approximately 1,4%) therefore one
would expect that prevent it does not alter the overall transcription profile (134). On the
other hand, when comparing the expression profile of Bub3 knockdown fly expressing
GFP and p35 with the other phenotypes transcription profiles we observed a dramatic
change in transcription. This means that when the two events occur together and as
cells undergo transformation, become immortal and tumorigenic, their transcription
profile is dramatically altered. Moreover, another trend is visible in the volcano plots for
this data, specifically the there are more upregulated genes than downregulated genes
in the overgrowth phenotype when comparing to any other phenotype. This might
indicate that gene upregulation has a larger role on the overgrowth phenotype,
validating our original assumtions.
4.4. BIOINFORMATIC METHODS SHOW ENRICHMENT
OF RELEVANT PATHWAYS FROM VENN DIAGRAM
SELECTION.
The microarray analysis goal was to find candidate genes responsible for the
tumorigenesis phenotype. The difficulty was to come up with a manageable list of
genes with biological relevance to the events and use them to perform further testing in
vivo. However, simple t-test comparison did not allow a selection of genes with a
manageable size. Therefore, we needed to resort to other methods to narrow the
search. With the visualization aid of Venn diagrams, we developed a rationale to
reduce the data. We separated our candidate genes in three classes and classified
them as components, the Bub3RNAi component, the p35 component, and the
component of the synergy effect of both components. The Bub3RNAi component is
expected to present genes responding to the lack of a functional SAC, as the p35
component should highlight genes responding to a block of apoptosis, and the synergy
component is expected to bring genes responding to the contribution of a lack of a
functional SAC and the apoptosis inhibition together. Although the same gene could
come from the same component we expected to find more often genes in the Bub3RNAi
component with a possible function as oncogenes or tumor suppressors or even their
activators or suppressors. On the other hand from the p35 component we were
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
69
expecting a list of genes more closely related with apoptosis either pro- or anti-
apoptotic genes or their activators or suppressors. We resorted to DAVID
bioinformatics resources 6.7 to evaluate the underlying principle beneath our selection.
We performed functional annotation clustering of each group of genes obtained from
the Venn diagrams. We chose a cut-off of approximately 1,3 enrichment score which
corresponds to the geometric mean of a p-value equal to 0,05 from all the members of
the particular cluster. This tool enrichment p-value is calculated based on, EASE
Score, a modified Fisher Exact Test (179). For further understanding I designed the
following example: assuming one submits a list of 400 genes and 20 hit the particular
category “cell cycle” and the background list from which the comparison was made has
300 genes in that category out of the total number of known genes for that organism
which could be 20000. With this information, we then ask: is the term “cell cycle”
enriched/over-represented in the list sent comparing with the number of hits that term
has in all the genome? The program then calculates one tail (right/greater tail) Fisher
Exact Test giving us a p-value and therefore the confidence interval of that particular
term being over-represented or not. It is important to notice that we are searching for
particular pathways that in theory could be related to an overgrowth phenotype which
represent a very small amount. Moreover, due to complexity and the unpredictable time
point of the tissue from where the data was extracted, it is likely that these pathways, if
altered, are being masqueraded beneath all the other pathways differently expressed
during development. As mention before we identified thirteen clusters above an
enrichment score of 1,3 in the upregulated Bub3RNAi component. From this thirteen we
classified one as relevant for the purpose of this thesis. This cluster is related to
several biological process GO terms like “DNA repair”, “telomere maintenance”,
“telomere organization” (180). Regarding the downregulated Bub3RNAi component we
did not classify any cluster as relevant however one of the clusters with cell motion as a
biological process GO term is worth referring. We did not define any cluster as
relevant for the p35 component either upregulated or down regulated. On the other
hand, the upregulated synergy component revealed only one cluster above the
specified enrichment value with 1,30 and some of its members are cellular component
GO terms like “spindle” or “microtubule cytoskeleton” and biological process “cell
cycle”. This is an expected result and supports the rationale used as the overgrowth
phenotype appears only in the case of the expression of p35 with Bub3 depletion. The
downregulated synergy component does not present any relevant cluster above the
specified enrichment value. Analysis of the previous results shows again, as referred
before in the volcano plots, the tendency for higher relevance in the upregulated genes.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
70
However, downregulated genes should not be overlooked. Another important feature
from the functional annotation clustering is the loss of information due to some
members of clusters with higher p-values might alter the p-value geometric mean from
which the enrichment score is calculated
4.5. LEGITIMACY OF MANUAL SELECTION FROM VENN
DIAGRAM.
Biological data-mining has become increasingly complex and the present state
of enrichment tools impede a pure statistical solution allowing only for exploratory data-
mining methods. The biological knowledge of a researcher with aid from integrated
annotation databases, computing algorithms and the enrichment P-values is still crucial
for a valuable analytic conclusion (160). Our final goal was to perform an in vivo RNAi
screen. Therefore, we required a manageable list of genes differently expressed and
the Venn diagram approach did not confer such list. Additional selection with the help
of known biological data was the last logical step to produce a list small enough to work
with but that did not lose its biological significance. We began by defining selection
criteria. First, we investigated each candidate gene by collecting information that could
be in any way relevant to the work in progress. We started by looking for fold difference
(prioritizing higher values), annotated functions, pathways and reasons for being
involved in tumorigenesis, cell division, cell cycle, apoptosis, any other closely related
function and the existence of human homologs (in a positive case the priority of the
candidate would be higher). With this first approach, we were able to get rid of many
well-known genes that were known to perform in other non-related functions. In case of
doubt, we also paid attention to protein-protein interactions using NCBI database
looking for evidence of relations between the product of our selected genes and
proteins involved in events like cell cycle, apoptosis, microtubules, or spindle assembly
checkpoint among others. We did not want to neglect genes without an annotated
function since ultimately, we wanted to find key new players in the hyperplastic growth.
We decided to include a threshold of a 10-fold difference, imposed by the available
resources, means and maintaining a manageable list, where we would include any
unknown sequences with a superior or equal level.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
71
4.6. BUB3 HAS AN ADDITIONAL ROLE AS A TUMOR
SUPPRESSOR.
We analyzed what could be each of genes that showed the most enhanced
phenotype in the performed screen. We also expect that whatever function they
perform should be influenced by the downregulation of Bub3 although further
investigation like testing the candidate genes by themselves is required. The
information depicted below is provided from Flybase.
4.6.1. GENES UPREGULATED IN BUB3RNAI
BACKGROUND SCREEN
Regarding the Bub3RNAi background the hits from the most enhanced phenotype
were:
CG6830 upregulated 7,06 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. It has ATPase
activity according to flybase and it interacts with HSP83 (181), which has an unknown
function in the mitotic spindle organization (182). Therefore, the gene CG6830 could
have a function of a suppressor of an oncogene or activator of a tumor suppressor.
CG18745 upregulated 2,67 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. It has no
annotated function according to flybase. However it seems to interact in an unknown
way with several cell cycle regulators like CDK4, CYCG, CYCK, MCM5 among others
(183). This interactions lead to the possible function as a tumor suppressor, suppressor
of an oncogene or an activator of a tumor suppressor.
CG5866 upregulated 18,67 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. This gene has
hardly any information available. So it could have any of the possible functions related
to an enhanced phenotype described in table III.8.
CG4089 upregulated 117,8 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. There are no
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
72
studies related to this gene therefore no annotated function. As such, it could perform
any of the possible functions related to an enhanced phenotype described in table III.8.
However, we can speculate that such a high fold could indicate a major role in the
tumorigenesis process.
Cp110 upregulated 3,47 fold in the Bub3 knockdown disc expressing GFP and
p35 when compared with the control disc expressing only GFP. It is linked with
centriole replication (184) it interacts with the Drosophila P53 and since its
phylogenetically related to the ancestor of the mammalian p53 family of tumor
suppressors (185), it could be an activator of the said tumor suppressor.
CG31686 upregulated 15,4 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. This gene has
hardly any information available. So it could have any of the possible functions related
to an enhanced phenotype described in table III.8.
CG18193 upregulated 17,59 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. This gene has
hardly any information available. It does seem to have a function related to electron
carrier activity inferred through electronic annotation. Therefore, it could have any of
the possible functions related to an enhanced phenotype described in table III.8.
CG33920 upregulated 25,44 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. There are no
studies related to this gene therefore no annotated function. As such, it could perform
any of the possible functions related to an enhanced phenotype described in table III.8.
CG30485 upregulated 12,9 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. This gene doesn’t
have any information available. Therefore, it could have any of the possible functions
related to an enhanced phenotype described in table III.8.
CG12911 upregulated 22,79 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. We found no
studies regarding this gene and as such , it could have any of the possible functions
related to an enhanced phenotype depicted in table III.8.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
73
CG31533 upregulated 24,45 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. This gene has
hardly any information available. So it could have any of the possible functions related
to an enhanced phenotype described in table III.8.
CG8193 upregulated 21,27 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. Through gene
similarity it has been associated with monophenol monooxygenase activity. In terms of
interactions it shows a relevant interaction with P53 and like previously mentioned,
since it is phylogenetically related to the ancestor of the mammalian p53 family of
tumor suppressors (185), it could be an activator of a tumor suppressor.
CG6333 upregulated 3,05 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. It has no studies
related to its function. However it interacts with P53 which could reveal an activator of a
tumor suppressor function (185). It also interacts with tribbles (trbl) a protein with
kinase activity and cells enters mitosis prematurely in tribbles mutant embryos (186),
which could mean the gene CG6333 might have a function as an activator of a tumor
suppressor.
CG31279 upregulated 23,11 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. This gene has
hardly any information available. So it could have any of the possible functions related
to an enhanced phenotype described in table III.8.
4.6.2. GENES DOWNREGULATED IN BUB3RNAI
BACKGROUND SCREEN
Regarding the Bub3RNAi background the hits from the most enhanced phenotype
were:
CG34380, downregulated 2,5 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. This particular
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
74
gene has little information however, through electronic annotation it has been
connected to cell adhesion. It also has an interaction with smt3 (181), a Drosophila
gene involved in mitotic cell cycle event as it is involved in the SUMO pathway and
seems to be required for for the syncytial mitotic cycles, for cell cycle progression in S2
cells and larval tissue and it even seems to be connected with Ras signaling (187).
Although CG34380 does not have a large amount of data available, it has a human
homolog, discoidin domain receptor tyrosine kinase 2 (DDR2) from which some cancer
related studies were done. Steven Wall et al. showed that this cell surface-associated
tyrosine kinase receptor that binds to fibrillar collagen (FC), endow with a new
mechanism responsible for the G0/G1 cell cycle arrest observed when tumor cells are
in contact with FC (188). Downregulation of CG34380 could then interfere with the cell
proliferation process either by influencing signaling pathways like the Ras pathway
through its interaction with SMT3, or even through some similar function to its human
counterpart DDR2. Since its annotated function of cell adhesion is not closely
associated with cell proliferation per se, we expect it to probably have a function of a
suppressor of an oncogene or activator of a tumor suppressor, which correlates with its
interaction with smt3 as this gene seems to be involved in pathways that could lead to
tumorigenesis. According to the data from its human homolog of being responsible for
a cell cycle arrest, we could go further and hypothesize that the gene CG34380 could
be an activator of a tumor suppressor.
Serpin 100A (Spn100A), downregulated 11,47 fold in the Bub3 knockdown disc
expressing GFP and p35 when compared with the control disc expressing only GFP.
There are no studies regarding this gene being only classified with serine-type
endopeptidase inhibitor activity through gene or structural similarity. It could have any
of the possible functions related to an enhanced phenotype described in table III.8.
CG31676 downregulated 10,53 fold in the Bub3 knockdown disc expressing
GFP and p35 when compared with the control disc expressing only GFP. This gene
has hardly any information available. So it could have any of the possible functions
related to an enhanced phenotype described in table III.8.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
75
4.6.3. GENES UPREGULATED IN BUB3RNAI DIAP1
BACKGROUND SCREEN
Regarding the Bub3RNAiDIAP1 background the hits from the most enhanced phenotype
were:
The candidate genes CG18745, CG5866, CG4089, Cp110, CG18193, CG33920,
CG12911, CG31533, CG8193, CG6333 and CG31279 functions and previsions are
already depicted in the Bub3RNAi background screen.
Nimrod B1 (NimB1) found upregulated 11,26 fold in the Bub3 knockdown disc
expressing GFP and p35 when compared with the control disc expressing only GFP.
This gene has hardly any information although it is linked with defense response to
bacterium through gene similarity (189). Therefore it could have any of the possible
functions related to an enhanced phenotype described in table III.8.
Acetyl Coenzyme A synthase (AcCoAS) found upregulated 12,36 fold in the Bub3
knockdown disc expressing GFP and p35 when compared with the control disc
expressing only GFP. This gene is linked with acetate-CoA ligase activity and its
closest relation with tumorigenesis is the interaction with supernumerary limbs
(SLMB) (190), which among other functions has been associated with regulation of
mitosis as loss of Slmb correlates with misregulation of mitotic cycles (191). Either
functions of tumor suppressor, suppressor of an oncogene or activator of a tumor
suppressor are a plausible hypothesis for this candidate gene.
Sialyltransferase (ST6Gal) found upregulated 6,66 fold in the Bub3 knockdown disc
expressing GFP and p35 when compared with the control disc expressing only GFP.
This gene has little information annotated being associated oligosaccharide metabolic
process (192). However as no link to tumorigenesis was found it could have any of the
possible functions related to an enhanced phenotype described in table III.8.
Thor found upregulated 10,08 fold in the Bub3 knockdown disc expressing GFP and
p35 when compared with the control disc expressing only GFP. Its function is
annotated as an eukaryotic initiation factor (193). This gene interacts with Eukaryotic
initiation factor 4E (eIF-4E) (181) that has been associated with mitotic spindle
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
76
organization (182). As such, Thor could also have a function of a suppressor of an
oncogene or activator of tumor suppressor.
Dhc62B found upregulated 3,55 fold in the Bub3 knockdown disc expressing GFP and
p35 when compared with the control disc expressing only GFP. This gene has hardly
any information although it is linked with microtubule movements through gene
similarity (194). Therefore it could have any of the possible functions related to an
enhanced phenotype described in table III.8.
CG6347 found upregulated 50,99 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. There are not
many studies regarding this gene being primarily linked with proteolysis through
electronic annotation. Therefore, it could have any of the possible functions related to
an enhanced phenotype described in table III.8.
CG6191 found upregulated 2,88 fold in the Bub3 knockdown disc expressing GFP and
p35 when compared with the control disc expressing only GFP. Its function was
defined, through electronic annotation, as cyclin-dependent protein kinase regulator
activity. It seems to be closely related with regulation of cell cycle and shows several
interactions with cell cycle regulators like Cyclin-dependent kinase 5 (CDK5) Cyclin-
dependent kinase subunit 85A (CKS85A) or altered disjunction (ald) (183). There are
several articles pointing for Cdk5 and Abl enzyme substrate 1 (Cable5), the human
homolog, as associated with tumorigenesis (195). Therefore, the expected function of
the CG6191 gene could be either suppressing an oncogene, activating a tumor
suppressor or even a tumor suppressor itself.
CG13116 found upregulated 12,52 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. We found no
studies regarding this gene and as such, it could have any of the possible functions
related to an enhanced phenotype depicted in table III.8.
CG6579 found upregulated 10,49 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. There are not
many studies regarding this gene. Therefore, it could have any of the possible
functions related to an enhanced phenotype described in table III.8.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
77
CG34430 found upregulated 10,27 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. This gene has
hardly any information so it could have any of the possible functions related to an
enhanced phenotype described in table III.8.
CG17108 found upregulated 20,07 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. This gene has
hardly any information however, the interaction with supernumerary limbs (slmb) (190),
which among other functions has been associated with regulation of mitosis as loss of
Slmb correlates with misregulation of mitotic cycles (191). Either functions of tumor
suppressor, suppressor of an oncogene or activator of a tumor suppressor are a
plausible hypothesis for this candidate gene.
CG4250 found upregulated 18,69 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. There are not
many studies regarding this gene. Therefore, it could have any of the possible
functions related to an enhanced phenotype described in table III.8.
Neu3 found upregulated 10,53 fold in the Bub3 knockdown disc expressing GFP and
p35 when compared with the control disc expressing only GFP. There are not many
studies regarding this gene being primarily linked with proteolysis through electronic
annotation. Its human homolog ADAM metallopeptidase domain 12 (ADAM12) on the
other hand seems to have some relation with tumorigenesis as its expression is low in
most normal tissues but is significantly increased in numerous human cancers, like
breast carcinomas (196). This lead us to hypothesize that Neu3 might have a function
as a tumor suppressor, suppressor of an oncogene or activator of a tumor suppressor.
CG3746 found upregulated 14,61 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. This gene has
hardly any information so it could have any of the possible functions related to an
enhanced phenotype described in table III.8.
Wun2 found upregulated 6,13 fold in the Bub3 knockdown disc expressing GFP and
p35 when compared with the control disc expressing only GFP. This gene known
functions aren’t closely related to tumorigenesis, therefore, it could have any new
unknown function related to an enhanced phenotype described in table III.8.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
78
Peroxiredoxin 3 (Prx5037) found upregulated 1733,3 fold in the Bub3 knockdown disc
expressing GFP and p35 when compared with the control disc expressing only GFP.
Among other functions this gene seems to be related with negative regulation of
apoptotic process and its human homolog peroxiredoxin 3 (Prx III) has been implicated
in the tumorigenesis of various cancers and data suggests that Prx III has a relevant
role in cell cycle regulation and could be a potential proliferation marker in for example
breast cancer (197). Therefore this gene can influence either apoptosis or cell
proliferation and as such any of the function related to an enhanced phenotype
described in table III.8 are a plausible possibility.
NimC1 found upregulated 6,93 fold in the Bub3 knockdown disc expressing GFP and
p35 when compared with the control disc expressing only GFP and its function is
annotated as a Wnt receptor signaling pathway and as such any of the function related
to an enhanced phenotype described in table III.8 are a plausible possibility.
CG8620 found upregulated 382,58 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. There are not
many studies regarding this gene. Therefore, it could have any of the possible
functions related to an enhanced phenotype described in table III.8.
PNUTS found upregulated 3,18 fold in the Bub3 knockdown disc expressing GFP and
p35 when compared with the control disc expressing only GFP. The main link of this
gene to tumorigenesis is its interaction with Spliceosomal protein on the X (SPX) which
seems to have a function in the mitotic spindle organization (182). With the information
available we can only speculate that it might be a tumor suppressor, suppressor of an
oncogene or activator of a tumor suppressor.
CG4763 found upregulated 13,18 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. This gene has
hardly any information so it could have any of the possible functions related to an
enhanced phenotype described in table III.8.
CG12520 found upregulated 12,15 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. There are not
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
79
many studies regarding this gene. Therefore, it could have any of the possible
functions related to an enhanced phenotype described in table III.8.
CG13250 found upregulated 24,74 fold in the Bub3 knockdown disc expressing GFP
and p35 when compared with the control disc expressing only GFP. There are not
many studies regarding this gene. Therefore, it could have any of the possible
functions related to an enhanced phenotype described in table III.8.
Augmenter of liver regeneration (Alr) found upregulated 2,15 fold in the Bub3
knockdown disc expressing GFP and p35 when compared with the control disc
expressing only GFP. This gene is linked to tissue regeneration (198) from which we
could speculate might confer him a function of either tumor suppressor, suppressor of
an oncogene or activator of a tumor suppressor.
Aplip1 found upregulated 4,9 fold in the Bub3 knockdown disc expressing GFP and
p35 when compared with the control disc expressing only GFP. This gene known
functions aren’t closely related to tumorigenesis, therefore, it could have any new
unknown function related to an enhanced phenotype described in table III.8.
CG1572 found upregulated 2,78 fold in the Bub3 knockdown disc expressing GFP and
p35 when compared with the control disc expressing only GFP. There are not many
studies regarding this gene. Therefore, it could have any of the possible functions
related to an enhanced phenotype described in table III.8.
CG7188 found upregulated 2,03 fold in the Bub3 knockdown disc expressing GFP and
p35 when compared with the control disc expressing only GFP. This gene seems to be
related with apoptosis therefore we speculate that its function could be either of a pro-
apoptotic gene, a suppressor of an anti-apoptotic gene or even an activator of a pro-
apoptotic gene.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
80
4.6.4. GENES DOWNREGULATED IN BUB3RNAI DIAP1
BACKGROUND SCREEN
Regarding the Bub3RNAiDIAP1 background the hits from the most enhanced
phenotype were:
CG6287 found downregulated 6,97 fold in the Bub3 knockdown disc expressing
GFP and p35 when compared with the control disc expressing only GFP. The gene
CG6287 has little information available, however it has phosphoglycerate
dehydrogenase activity annotated as function inferred from gene or structural
similarities and seems to be involved in L-serine biosynthetic process. Considering
interactions, we found two particular interesting ones, Lissencephaly-1 (LIS-1) (181)
and TSC1 (181). LIS-1 interacts with dynein through a WD40 repeat. Siller et al.
showed that LIS1 and dynactin are required for centrosome separation and spindle
assembly during prophase/prometaphase and by allowing an efficient mitotic
checkpoint inactivation it contribute to timely metaphase-to-anaphase transition (199).
TSC1 in junction with TSC2 seems the regulate Cell Cycle via Changes in Cyclin
Levels, TSC1 and TSC2 also seem to regulate Cell Cycle Exit as TSC mutant cells
aren’t able to sustain a developmentally induced G1 arrest (200).
Moreover, CG6287 shows homology in vertebrates. In humans its counterpart
is phosphoglycerate dehydrogenase (PHGDH) and it has been associated with cell
cycle processes through electronic annotation and oncogenesis (201, 202). Regarding
this data we expect this gene to have a function of either a tumor suppressor itself, a
suppressor of an oncogene or even an activator of a tumor suppressor.
Topoisomerase 1 (top1) found downregulated 4,22 fold in the Bub3 knockdown
disc expressing GFP and p35 when compared with the control disc expressing only
GFP. This gene is associated with functions in cell proliferation, chromosome
segregation and condensation (203). TOP1 is a monomeric protein and it works by
producing a transient single-strand break in DNA, allowing passage to another strand,
contributing to reduce DNA supercoiling. Top 1 function suggests that this protein might
act at several critical cellular events serving as a swivel for unwinding and rewinding of
DNA helices. The human homolog, besides having similar functions to the Drosophila
counterpart, it has been implicated in small cell lung cancer (204). A closer look to
TOP1 interactions reveal an association with a protein connected with mitotic cell cycle
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
81
G2/M transition DNA damage checkpoint, Topoisomerase I-interacting protein
(TOPORS) (205). By adding all the evidence it is possible to assume that there could
be a synergy work, implying that BUB3 and TOP1 might interact preventing
uncontrolled cell proliferation and their depletion when apoptosis is inhibited might
cause an increased cell proliferation that could eventually lead to an overgrowth
depending on levels of downregulation. This data also suggests that any of the
functions involving an enhanced phenotype, a tumor suppressing, a suppressing of an
oncogene or activatating of a tumor suppressor are viable possibilities to explain how
this protein works regarding tumorigenesis.
CG12071 found downregulated 5,9 fold in the Bub3 knockdown disc expressing
GFP and p35 when compared with the control disc expressing only GFP. There’s only
a reported role for this protein in phagocytosis (206). Although the downregulation of
expression for this candidate is below the threshold imposed for unknown genes, its
reference in a microarray screen for Notch and Ras signaling pathway led to the choice
of including it (207). With this data we can assume that this gene could have any of the
possible functions related to an enhanced phenotype described in table III.8.
CG13101 found downregulated 12,09 fold in the Bub3 knockdown disc
expressing GFP and p35 when compared with the control disc expressing only GFP.
This is one of the unknown genes chosen due to being above the enforced threshold of
10 fold. Consequently, there is no studies regarding this particular gene. As such,
further research would be necessary to link this gene to any tumorigenesis or cell
proliferation event. As such, this gene could have any of the possible functions related
to an enhanced phenotype described in table III.8.
The hits found in the screen with the additional data from previous work send us to the
direction of Bub3 as possible tumor suppressor. Co-expression of Bub3 with p35, an
apoptosis inhibitor, in wing imaginal discs leads to an overgrowth (175). As such, the
RNAi enhancer/suppressor screen gave some insight on what might be downstream of
this process. However, further studies would have to be conducted in order to find if the
hits found are really linked to the Bub3 possible function as a tumor suppressor. These
studies could include using mutants for the most relevant candidates, test the
candidate genes by themselves or dive into molecular studies like pull down assays to
understand the exact role of each candidate in the hyperplastic phenotype and its
relation with Bub3. The screen itself has some flaws, as there is the possibility of many
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
82
false positives in which we might get the phenotype we are looking for although having
no connection with Bub3 and might simply act by themselves or by influencing another
cell cycle or apoptosis genes. However, a screen like this is one of the best
approaches to trim data and lead into the right direction.
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
84
1. F. Hirth, Drosophila melanogaster in the study of human neurodegeneration. CNS &
neurological disorders drug targets 9, 504 (Aug, 2010). 2. X. Gao, T. P. Neufeld, D. Pan, Drosophila PTEN regulates cell growth and proliferation
through PI3K-dependent and -independent pathways. Developmental biology 221, 404 (May 15, 2000).
3. F. D. Karim et al., A screen for genes that function downstream of Ras1 during Drosophila eye development. Genetics 143, 315 (May, 1996).
4. L. A. Lee, T. L. Orr-Weaver, Regulation of cell cycles in Drosophila development: intrinsic and extrinsic cues. Annual review of genetics 37, 545 (2003).
5. D. Pan, The hippo signaling pathway in development and cancer. Developmental cell 19, 491 (Oct 19, 2010).
6. M. A. Simon, Receptor tyrosine kinases: specific outcomes from general signals. Cell 103, 13 (Sep 29, 2000).
7. M. A. Simon, D. D. Bowtell, G. S. Dodson, T. R. Laverty, G. M. Rubin, Ras1 and a putative guanine nucleotide exchange factor perform crucial steps in signaling by the sevenless protein tyrosine kinase. Cell 67, 701 (Nov 15, 1991).
8. M. G. Voas, I. Rebay, Signal integration during development: insights from the Drosophila eye. Developmental dynamics : an official publication of the American Association of Anatomists 229, 162 (Jan, 2004).
9. C. W. Wilson, P. T. Chuang, Mechanism and evolution of cytosolic Hedgehog signal transduction. Development 137, 2079 (Jul, 2010).
10. M. B. Stark, A Benign Tumor that is Hereditary in Drosophila. Proceedings of the National Academy of Sciences of the United States of America 5, 573 (Dec, 1919).
11. D. St Johnston, The art and design of genetic screens: Drosophila melanogaster. Nature reviews. Genetics 3, 176 (Mar, 2002).
12. H. J. Bellen et al., The BDGP gene disruption project: single transposon insertions associated with 40% of Drosophila genes. Genetics 167, 761 (Jun, 2004).
13. A. H. Brand, N. Perrimon, Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development 118, 401 (Jun, 1993).
14. T. Lee, L. Luo, Mosaic analysis with a repressible cell marker for studies of gene function in neuronal morphogenesis. Neuron 22, 451 (Mar, 1999).
15. G. Toba et al., The gene search system. A method for efficient detection and rapid molecular identification of genes in Drosophila melanogaster. Genetics 151, 725 (Feb, 1999).
16. P. Rorth, A modular misexpression screen in Drosophila detecting tissue-specific phenotypes. Proceedings of the National Academy of Sciences of the United States of America 93, 12418 (Oct 29, 1996).
17. A. F. Chambers, A. C. Groom, I. C. MacDonald, Dissemination and growth of cancer cells in metastatic sites. Nature reviews. Cancer 2, 563 (Aug, 2002).
18. G. P. Gupta, J. Massague, Cancer metastasis: building a framework. Cell 127, 679 (Nov 17, 2006).
19. L. S. Michel et al., MAD2 haplo-insufficiency causes premature anaphase and chromosome instability in mammalian cells. Nature 409, 355 (Jan 18, 2001).
20. D. J. Baker et al., BubR1 insufficiency causes early onset of aging-associated phenotypes and infertility in mice. Nature genetics 36, 744 (Jul, 2004).
21. J. R. Babu et al., Rae1 is an essential mitotic checkpoint regulator that cooperates with Bub3 to prevent chromosome missegregation. The Journal of cell biology 160, 341 (Feb 3, 2003).
22. W. Dai et al., Slippage of mitotic arrest and enhanced tumor development in mice with BubR1 haploinsufficiency. Cancer research 64, 440 (Jan 15, 2004).
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
85
23. D. P. Cahill et al., Mutations of mitotic checkpoint genes in human cancers. Nature
392, 300 (Mar 19, 1998). 24. S. Hanks et al., Constitutional aneuploidy and cancer predisposition caused by biallelic
mutations in BUB1B. Nature genetics 36, 1159 (Nov, 2004). 25. Y. Mao, A. Abrieu, D. W. Cleveland, Activating and silencing the mitotic checkpoint
through CENP-E-dependent activation/inactivation of BubR1. Cell 114, 87 (Jul 11, 2003).
26. Z. Tang, R. Bharadwaj, B. Li, H. Yu, Mad2-Independent inhibition of APCCdc20 by the mitotic checkpoint protein BubR1. Developmental cell 1, 227 (Aug, 2001).
27. X. Wang et al., Correlation of defective mitotic checkpoint with aberrantly reduced expression of MAD2 protein in nasopharyngeal carcinoma cells. Carcinogenesis 21, 2293 (Dec, 2000).
28. B. A. Weaver, D. W. Cleveland, Decoding the links between mitosis, cancer, and chemotherapy: The mitotic checkpoint, adaptation, and cell death. Cancer cell 8, 7 (Jul, 2005).
29. W. O. Miles, N. J. Dyson, J. A. Walker, Modeling tumor invasion and metastasis in Drosophila. Disease models & mechanisms 4, 753 (Nov, 2011).
30. K. A. Wharton, K. M. Johansen, T. Xu, S. Artavanis-Tsakonas, Nucleotide sequence from the neurogenic locus notch implies a gene product that shares homology with proteins containing EGF-like repeats. Cell 43, 567 (Dec, 1985).
31. T. H. Morgan, The Theory of the Gene. 51, 513 (1917). 32. D. L. Lindsley, G. G. Zimm, The Genome of Drosophila Melanogaster. (Academic Press,
Inc., 1992). 33. Y. Ye, N. Lukinova, M. E. Fortini, Neurogenic phenotypes and altered Notch processing
in Drosophila Presenilin mutants. Nature 398, 525 (Apr 8, 1999). 34. L. W. Ellisen et al., TAN-1, the human homolog of the Drosophila notch gene, is broken
by chromosomal translocations in T lymphoblastic neoplasms. Cell 66, 649 (Aug 23, 1991).
35. C. Nusslein-Volhard, E. Wieschaus, Mutations affecting segment number and polarity in Drosophila. Nature 287, 795 (Oct 30, 1980).
36. H. Hahn et al., Mutations of the human homolog of Drosophila patched in the nevoid basal cell carcinoma syndrome. Cell 85, 841 (Jun 14, 1996).
37. R. P. Sharma, V. L. Chopra, Effect of the Wingless (wg1) mutation on wing and haltere development in Drosophila melanogaster. Developmental biology 48, 461 (Feb, 1976).
38. K. W. Kinzler et al., Identification of FAP locus genes from chromosome 5q21. Science 253, 661 (Aug 9, 1991).
39. E. Ogawa et al., PEBP2/PEA2 represents a family of transcription factors homologous to the products of the Drosophila runt gene and the human AML1 gene. Proceedings of the National Academy of Sciences of the United States of America 90, 6859 (Jul 15, 1993).
40. T. Okuda, J. van Deursen, S. W. Hiebert, G. Grosveld, J. R. Downing, AML1, the target of multiple chromosomal translocations in human leukemia, is essential for normal fetal liver hematopoiesis. Cell 84, 321 (Jan 26, 1996).
41. C. H. Pui, M. V. Relling, J. R. Downing, Acute lymphoblastic leukemia. The New England journal of medicine 350, 1535 (Apr 8, 2004).
42. E. B. Lewis, A gene complex controlling segmentation in Drosophila. Nature 276, 565 (Dec 7, 1978).
43. M. Djabali et al., A trithorax-like gene is interrupted by chromosome 11q23 translocations in acute leukaemias. Nature genetics 2, 113 (Oct, 1992).
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
86
44. Y. Gu et al., The t(4;11) chromosome translocation of human acute leukemias fuses the
ALL-1 gene, related to Drosophila trithorax, to the AF-4 gene. Cell 71, 701 (Nov 13, 1992).
45. R. K. Slany, The molecular biology of mixed lineage leukemia. Haematologica 94, 984 (Jul, 2009).
46. M. E. Martin et al., Dimerization of MLL fusion proteins immortalizes hematopoietic cells. Cancer cell 4, 197 (Sep, 2003).
47. T. Burmeister et al., The MLL recombinome of adult CD10-negative B-cell precursor acute lymphoblastic leukemia: results from the GMALL study group. Blood 113, 4011 (Apr 23, 2009).
48. C. Meyer et al., New insights to the MLL recombinome of acute leukemias. Leukemia : official journal of the Leukemia Society of America, Leukemia Research Fund, U.K 23, 1490 (Aug, 2009).
49. S. A. Armstrong et al., MLL translocations specify a distinct gene expression profile that distinguishes a unique leukemia. Nature genetics 30, 41 (Jan, 2002).
50. A. V. Krivtsov, S. A. Armstrong, MLL translocations, histone modifications and leukaemia stem-cell development. Nature reviews. Cancer 7, 823 (Nov, 2007).
51. K. Geissler, O. Zach, Pathways involved in Drosophila and human cancer development: the Notch, Hedgehog, Wingless, Runt, and Trithorax pathway. Annals of hematology 91, 645 (May, 2012).
52. J. M. Peters, The anaphase promoting complex/cyclosome: a machine designed to destroy. Nature reviews. Molecular cell biology 7, 644 (Sep, 2006).
53. B. R. Thornton, D. P. Toczyski, Precise destruction: an emerging picture of the APC. Genes & development 20, 3069 (Nov 15, 2006).
54. R. Bharadwaj, H. Yu, The spindle checkpoint, aneuploidy, and cancer. Oncogene 23, 2016 (Mar 15, 2004).
55. A. Musacchio, E. D. Salmon, The spindle-assembly checkpoint in space and time. Nature reviews. Molecular cell biology 8, 379 (May, 2007).
56. C. L. Rieder, R. W. Cole, A. Khodjakov, G. Sluder, The checkpoint delaying anaphase in response to chromosome monoorientation is mediated by an inhibitory signal produced by unattached kinetochores. The Journal of cell biology 130, 941 (Aug, 1995).
57. C. L. Rieder, A. Schultz, R. Cole, G. Sluder, Anaphase onset in vertebrate somatic cells is controlled by a checkpoint that monitors sister kinetochore attachment to the spindle. The Journal of cell biology 127, 1301 (Dec, 1994).
58. H. Yu, Cdc20: a WD40 activator for a cell cycle degradation machine. Molecular cell 27, 3 (Jul 6, 2007).
59. Z. Tang et al., APC2 Cullin protein and APC11 RING protein comprise the minimal ubiquitin ligase module of the anaphase-promoting complex. Molecular biology of the cell 12, 3839 (Dec, 2001).
60. C. W. Carroll, D. O. Morgan, The Doc1 subunit is a processivity factor for the anaphase-promoting complex. Nature cell biology 4, 880 (Nov, 2002).
61. C. W. Carroll, M. Enquist-Newman, D. O. Morgan, The APC subunit Doc1 promotes recognition of the substrate destruction box. Current biology : CB 15, 11 (Jan 11, 2005).
62. C. Kraft, H. C. Vodermaier, S. Maurer-Stroh, F. Eisenhaber, J. M. Peters, The WD40 propeller domain of Cdh1 functions as a destruction box receptor for APC/C substrates. Molecular cell 18, 543 (May 27, 2005).
63. V. Sudakin, G. K. Chan, T. J. Yen, Checkpoint inhibition of the APC/C in HeLa cells is mediated by a complex of BUBR1, BUB3, CDC20, and MAD2. The Journal of cell biology 154, 925 (Sep 3, 2001).
64. H. Yu, Regulation of APC-Cdc20 by the spindle checkpoint. Current opinion in cell biology 14, 706 (Dec, 2002).
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
87
65. M. Li, X. Fang, Z. Wei, J. P. York, P. Zhang, Loss of spindle assembly checkpoint-
mediated inhibition of Cdc20 promotes tumorigenesis in mice. The Journal of cell biology 185, 983 (Jun 15, 2009).
66. M. Yang et al., Insights into mad2 regulation in the spindle checkpoint revealed by the crystal structure of the symmetric mad2 dimer. PLoS biology 6, e50 (Mar 4, 2008).
67. M. Mapelli, A. Musacchio, MAD contortions: conformational dimerization boosts spindle checkpoint signaling. Current opinion in structural biology 17, 716 (Dec, 2007).
68. X. Luo, H. Yu, Protein metamorphosis: the two-state behavior of Mad2. Structure 16, 1616 (Nov 12, 2008).
69. S. Kim, H. Yu, Mutual regulation between the spindle checkpoint and APC/C. Seminars in cell & developmental biology 22, 551 (Aug, 2011).
70. G. Fang, Checkpoint protein BubR1 acts synergistically with Mad2 to inhibit anaphase-promoting complex. Molecular biology of the cell 13, 755 (Mar, 2002).
71. S. Elowe et al., Uncoupling of the spindle-checkpoint and chromosome-congression functions of BubR1. Journal of cell science 123, 84 (Jan 1, 2010).
72. M. Sczaniecka et al., The spindle checkpoint functions of Mad3 and Mad2 depend on a Mad3 KEN box-mediated interaction with Cdc20-anaphase-promoting complex (APC/C). The Journal of biological chemistry 283, 23039 (Aug 22, 2008).
73. E. Choi et al., BubR1 acetylation at prometaphase is required for modulating APC/C activity and timing of mitosis. The EMBO journal 28, 2077 (Jul 22, 2009).
74. A. Kulukian, J. S. Han, D. W. Cleveland, Unattached kinetochores catalyze production of an anaphase inhibitor that requires a Mad2 template to prime Cdc20 for BubR1 binding. Developmental cell 16, 105 (Jan, 2009).
75. F. Herzog et al., Structure of the anaphase-promoting complex/cyclosome interacting with a mitotic checkpoint complex. Science 323, 1477 (Mar 13, 2009).
76. Z. Tang, H. Shu, D. Oncel, S. Chen, H. Yu, Phosphorylation of Cdc20 by Bub1 provides a catalytic mechanism for APC/C inhibition by the spindle checkpoint. Molecular cell 16, 387 (Nov 5, 2004).
77. V. P. Efimov, N. R. Morris, A screen for dynein synthetic lethals in Aspergillus nidulans identifies spindle assembly checkpoint genes and other genes involved in mitosis. Genetics 149, 101 (May, 1998).
78. S. S. Taylor, E. Ha, F. McKeon, The human homologue of Bub3 is required for kinetochore localization of Bub1 and a Mad3/Bub1-related protein kinase. The Journal of cell biology 142, 1 (Jul 13, 1998).
79. M. J. Martinez-Exposito, K. B. Kaplan, J. Copeland, P. K. Sorger, Retention of the BUB3 checkpoint protein on lagging chromosomes. Proceedings of the National Academy of Sciences of the United States of America 96, 8493 (Jul 20, 1999).
80. E. J. Neer, C. J. Schmidt, R. Nambudripad, T. F. Smith, The ancient regulatory-protein family of WD-repeat proteins. Nature 371, 297 (Sep 22, 1994).
81. P. Kalitsis, E. Earle, K. J. Fowler, K. H. Choo, Bub3 gene disruption in mice reveals essential mitotic spindle checkpoint function during early embryogenesis. Genes & development 14, 2277 (Sep 15, 2000).
82. J. Basu et al., Mutations in the essential spindle checkpoint gene bub1 cause chromosome missegregation and fail to block apoptosis in Drosophila. The Journal of cell biology 146, 13 (Jul 12, 1999).
83. J. Basu et al., Localization of the Drosophila checkpoint control protein Bub3 to the kinetochore requires Bub1 but not Zw10 or Rod. Chromosoma 107, 376 (Dec, 1998).
84. C. S. Lopes, P. Sampaio, B. Williams, M. Goldberg, C. E. Sunkel, The Drosophila Bub3 protein is required for the mitotic checkpoint and for normal accumulation of cyclins during G2 and early stages of mitosis. Journal of cell science 118, 187 (Jan 1, 2005).
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
88
85. Y. Niikura, H. Ogi, K. Kikuchi, K. Kitagawa, BUB3 that dissociates from BUB1 activates
caspase-independent mitotic death (CIMD). Cell death and differentiation 17, 1011 (Jun, 2010).
86. A. Bergmann, H. Steller, Apoptosis, stem cells, and tissue regeneration. Science signaling 3, re8 (2010).
87. M. D. Piper, D. Skorupa, L. Partridge, Diet, metabolism and lifespan in Drosophila. Experimental gerontology 40, 857 (Nov, 2005).
88. S. Kumar, Caspase function in programmed cell death. Cell death and differentiation 14, 32 (Jan, 2007).
89. J. Yuan, S. Shaham, S. Ledoux, H. M. Ellis, H. R. Horvitz, The C. elegans cell death gene ced-3 encodes a protein similar to mammalian interleukin-1 beta-converting enzyme. Cell 75, 641 (Nov 19, 1993).
90. P. Fuentes-Prior, G. S. Salvesen, The protein structures that shape caspase activity, specificity, activation and inhibition. The Biochemical journal 384, 201 (Dec 1, 2004).
91. J. C. Reed, Apoptosis mechanisms: implications for cancer drug discovery. Oncology (Williston Park) 18, 11 (Nov, 2004).
92. E. N. Shiozaki, Y. Shi, Caspases, IAPs and Smac/DIABLO: mechanisms from structural biology. Trends in biochemical sciences 29, 486 (Sep, 2004).
93. A. Bergmann, A. Y. Yang, M. Srivastava, Regulators of IAP function: coming to grips with the grim reaper. Current opinion in cell biology 15, 717 (Dec, 2003).
94. Y. Shi, Mechanisms of caspase activation and inhibition during apoptosis. Molecular cell 9, 459 (Mar, 2002).
95. R. Wilson et al., The DIAP1 RING finger mediates ubiquitination of Dronc and is indispensable for regulating apoptosis. Nature cell biology 4, 445 (Jun, 2002).
96. T. Tenev, A. Zachariou, R. Wilson, M. Ditzel, P. Meier, IAPs are functionally non-equivalent and regulate effector caspases through distinct mechanisms. Nature cell biology 7, 70 (Jan, 2005).
97. S. L. Wang, C. J. Hawkins, S. J. Yoo, H. A. Muller, B. A. Hay, The Drosophila caspase inhibitor DIAP1 is essential for cell survival and is negatively regulated by HID. Cell 98, 453 (Aug 20, 1999).
98. L. Goyal, K. McCall, J. Agapite, E. Hartwieg, H. Steller, Induction of apoptosis by Drosophila reaper, hid and grim through inhibition of IAP function. The EMBO journal 19, 589 (Feb 15, 2000).
99. S. Lisi, I. Mazzon, K. White, Diverse domains of THREAD/DIAP1 are required to inhibit apoptosis induced by REAPER and HID in Drosophila. Genetics 154, 669 (Feb, 2000).
100. X. Li, J. Wang, Y. Shi, Structural mechanisms of DIAP1 auto-inhibition and DIAP1-mediated inhibition of drICE. Nature communications 2, 408 (2011).
101. D. Vucic, W. J. Kaiser, A. J. Harvey, L. K. Miller, Inhibition of reaper-induced apoptosis by interaction with inhibitor of apoptosis proteins (IAPs). Proceedings of the National Academy of Sciences of the United States of America 94, 10183 (Sep 16, 1997).
102. N. Yan et al., Structural, biochemical, and functional analyses of CED-9 recognition by the proapoptotic proteins EGL-1 and CED-4. Mol Cell 15, 999 (Sep 24, 2004).
103. K. White et al., Genetic control of programmed cell death in Drosophila. Science 264, 677 (Apr 29, 1994).
104. M. E. Grether, J. M. Abrams, J. Agapite, K. White, H. Steller, The head involution defective gene of Drosophila melanogaster functions in programmed cell death. Genes & development 9, 1694 (Jul 15, 1995).
105. P. Chen, W. Nordstrom, B. Gish, J. M. Abrams, grim, a novel cell death gene in Drosophila. Genes & development 10, 1773 (Jul 15, 1996).
106. S. M. Srinivasula et al., sickle, a novel Drosophila death gene in the reaper/hid/grim region, encodes an IAP-inhibitory protein. Current biology : CB 12, 125 (Jan 22, 2002).
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
89
107. D. Vucic, W. J. Kaiser, L. K. Miller, Inhibitor of apoptosis proteins physically interact
with and block apoptosis induced by Drosophila proteins HID and GRIM. Molecular and cellular biology 18, 3300 (Jun, 1998).
108. C. L. Holley, M. R. Olson, D. A. Colon-Ramos, S. Kornbluth, Reaper eliminates IAP proteins through stimulated IAP degradation and generalized translational inhibition. Nature cell biology 4, 439 (Jun, 2002).
109. J. Chai et al., Molecular mechanism of Reaper-Grim-Hid-mediated suppression of DIAP1-dependent Dronc ubiquitination. Nature structural biology 10, 892 (Nov, 2003).
110. A. Zachariou et al., IAP-antagonists exhibit non-redundant modes of action through differential DIAP1 binding. The EMBO journal 22, 6642 (Dec 15, 2003).
111. H. D. Ryoo, A. Bergmann, H. Gonen, A. Ciechanover, H. Steller, Regulation of Drosophila IAP1 degradation and apoptosis by reaper and ubcD1. Nature cell biology 4, 432 (Jun, 2002).
112. S. J. Yoo et al., Hid, Rpr and Grim negatively regulate DIAP1 levels through distinct mechanisms. Nature cell biology 4, 416 (Jun, 2002).
113. J. V. McCarthy, V. M. Dixit, Apoptosis induced by Drosophila reaper and grim in a human system. Attenuation by inhibitor of apoptosis proteins (cIAPs). The Journal of biological chemistry 273, 24009 (Sep 11, 1998).
114. W. N. Haining, C. Carboy-Newcomb, C. L. Wei, H. Steller, The proapoptotic function of Drosophila Hid is conserved in mammalian cells. Proceedings of the National Academy of Sciences of the United States of America 96, 4936 (Apr 27, 1999).
115. M. H. Brodsky et al., Drosophila melanogaster MNK/Chk2 and p53 regulate multiple DNA repair and apoptotic pathways following DNA damage. Molecular and cellular biology 24, 1219 (Feb, 2004).
116. N. Sogame, M. Kim, J. M. Abrams, Drosophila p53 preserves genomic stability by regulating cell death. Proceedings of the National Academy of Sciences of the United States of America 100, 4696 (Apr 15, 2003).
117. D. G. McEwen, M. Peifer, Puckered, a Drosophila MAPK phosphatase, ensures cell viability by antagonizing JNK-induced apoptosis. Development 132, 3935 (Sep, 2005).
118. M. H. Brodsky et al., Drosophila p53 binds a damage response element at the reaper locus. Cell 101, 103 (Mar 31, 2000).
119. E. Moreno, K. Basler, G. Morata, Cells compete for decapentaplegic survival factor to prevent apoptosis in Drosophila wing development. Nature 416, 755 (Apr 18, 2002).
120. M. Ollmann et al., Drosophila p53 is a structural and functional homolog of the tumor suppressor p53. Cell 101, 91 (Mar 31, 2000).
121. S. Y. Vernooy et al., Drosophila Bruce can potently suppress Rpr- and Grim-dependent but not Hid-dependent cell death. Current biology : CB 12, 1164 (Jul 9, 2002).
122. E. Arama, J. Agapite, H. Steller, Caspase activity and a specific cytochrome C are required for sperm differentiation in Drosophila. Developmental cell 4, 687 (May, 2003).
123. B. A. Hay, D. A. Wassarman, G. M. Rubin, Drosophila homologs of baculovirus inhibitor of apoptosis proteins function to block cell death. Cell 83, 1253 (Dec 29, 1995).
124. G. Jones, D. Jones, L. Zhou, H. Steller, Y. Chu, Deterin, a new inhibitor of apoptosis from Drosophila melanogaster. The Journal of biological chemistry 275, 22157 (Jul 21, 2000).
125. L. Zhou, Z. Song, J. Tittel, H. Steller, HAC-1, a Drosophila homolog of APAF-1 and CED-4 functions in developmental and radiation-induced apoptosis. Molecular cell 4, 745 (Nov, 1999).
126. C. Jiang, A. F. Lamblin, H. Steller, C. S. Thummel, A steroid-triggered transcriptional hierarchy controls salivary gland cell death during Drosophila metamorphosis. Molecular cell 5, 445 (Mar, 2000).
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
90
127. L. Dorstyn, P. A. Colussi, L. M. Quinn, H. Richardson, S. Kumar, DRONC, an ecdysone-
inducible Drosophila caspase. Proceedings of the National Academy of Sciences of the United States of America 96, 4307 (Apr 13, 1999).
128. S. Kornbluth, K. White, Apoptosis in Drosophila: neither fish nor fowl (nor man, nor worm). Journal of cell science 118, 1779 (May 1, 2005).
129. E. H. Baehrecke, How death shapes life during development. Nature reviews. Molecular cell biology 3, 779 (Oct, 2002).
130. Y. H. Song, Drosophila melanogaster: a model for the study of DNA damage checkpoint response. Molecules and cells 19, 167 (Apr 30, 2005).
131. I. Lohmann, N. McGinnis, M. Bodmer, W. McGinnis, The Drosophila Hox gene deformed sculpts head morphology via direct regulation of the apoptosis activator reaper. Cell 110, 457 (Aug 23, 2002).
132. T. Wolff, D. F. Ready, Cell death in normal and rough eye mutants of Drosophila. Development 113, 825 (Nov, 1991).
133. C. Manjon, E. Sanchez-Herrero, M. Suzanne, Sharp boundaries of Dpp signalling trigger local cell death required for Drosophila leg morphogenesis. Nature cell biology 9, 57 (Jan, 2007).
134. M. Milan, S. Campuzano, A. Garcia-Bellido, Developmental parameters of cell death in the wing disc of Drosophila. Proceedings of the National Academy of Sciences of the United States of America 94, 5691 (May 27, 1997).
135. A. Perez-Garijo, F. A. Martin, G. Morata, Caspase inhibition during apoptosis causes abnormal signalling and developmental aberrations in Drosophila. Development 131, 5591 (Nov, 2004).
136. J. L. Haynie, P. J. Bryant, The effects of X-rays on the proliferation dynamics of Cells in the imagina wing disc of Drosophila melanogaster. Wilhelm Roux’s Archives 183, 85 (1977).
137. J. Colombani, C. Polesello, F. Josue, N. Tapon, Dmp53 activates the Hippo pathway to promote cell death in response to DNA damage. Current biology : CB 16, 1453 (Jul 25, 2006).
138. L. J. Saucedo, B. A. Edgar, Filling out the Hippo pathway. Nature reviews. Molecular cell biology 8, 613 (Aug, 2007).
139. B. J. Graves, G. Schubiger, Cell cycle changes during growth and differentiation of imaginal leg discs in Drosophila melanogaster. Developmental biology 93, 104 (Sep, 1982).
140. F. A. Martin, A. Perez-Garijo, G. Morata, Apoptosis in Drosophila: compensatory proliferation and undead cells. The International journal of developmental biology 53, 1341 (2009).
141. Y. Xia, M. Karin, The control of cell motility and epithelial morphogenesis by Jun kinases. Trends in cell biology 14, 94 (Feb, 2004).
142. E. Martin-Blanco et al., puckered encodes a phosphatase that mediates a feedback loop regulating JNK activity during dorsal closure in Drosophila. Genes & development 12, 557 (Feb 15, 1998).
143. K. Mills, T. Daish, S. Kumar, The function of the Drosophila caspase DRONC in cell death and development. Cell Cycle 4, 744 (Jun, 2005).
144. B. A. Hay, T. Wolff, G. M. Rubin, Expression of baculovirus P35 prevents cell death in Drosophila. Development 120, 2121 (Aug, 1994).
145. B. A. Callus, D. L. Vaux, Caspase inhibitors: viral, cellular and chemical. Cell death and differentiation 14, 73 (Jan, 2007).
146. T. Tabata, Y. Takei, Morphogens, their identification and regulation. Development 131, 703 (Feb, 2004).
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
91
147. M. Gardiner-Garden, T. G. Littlejohn, A comparison of microarray databases. Briefings
in bioinformatics 2, 143 (May, 2001). 148. E. Georgii, L. Richter, U. Ruckert, S. Kramer, Analyzing microarray data using
quantitative association rules. Bioinformatics 21 Suppl 2, ii123 (Sep 1, 2005). 149. M. Schena, D. Shalon, R. W. Davis, P. O. Brown, Quantitative monitoring of gene
expression patterns with a complementary DNA microarray. Science 270, 467 (Oct 20, 1995).
150. C. A. Afshari, Perspective: microarray technology, seeing more than spots. Endocrinology 143, 1983 (Jun, 2002).
151. W. P. Kuo et al., Microarrays and clinical dentistry. J Am Dent Assoc 134, 456 (Apr, 2003).
152. M. Schena et al., Microarrays: biotechnology's discovery platform for functional genomics. Trends in biotechnology 16, 301 (Jul, 1998).
153. S. Michiels, S. Koscielny, C. Hill, Prediction of cancer outcome with microarrays: a multiple random validation strategy. Lancet 365, 488 (Feb 5-11, 2005).
154. P. O. Brown, D. Botstein, Exploring the new world of the genome with DNA microarrays. Nature genetics 21, 33 (Jan, 1999).
155. S. V. Tillib, A. D. Mirzabekov, Advances in the analysis of DNA sequence variations using oligonucleotide microchip technology. Current opinion in biotechnology 12, 53 (Feb, 2001).
156. M. Beier, J. D. Hoheisel, Production by quantitative photolithographic synthesis of individually quality checked DNA microarrays. Nucleic acids research 28, E11 (Feb 15, 2000).
157. W. Ji, W. Zhou, K. Gregg, K. Lindpaintner, S. Davis, A method for gene expression analysis by oligonucleotide arrays from minute biological materials. Analytical biochemistry 331, 329 (Aug 15, 2004).
158. R. Lucito et al., Representational oligonucleotide microarray analysis: a high-resolution method to detect genome copy number variation. Genome research 13, 2291 (Oct, 2003).
159. L. Guo et al., Gene expression profiling of drug-resistant small cell lung cancer cells by combining microRNA and cDNA expression analysis. Eur J Cancer 46, 1692 (Jun, 2010).
160. W. Huang da, B. T. Sherman, R. A. Lempicki, Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic acids research 37, 1 (Jan, 2009).
161. C. Napoli, C. Lemieux, R. Jorgensen, Introduction of a Chimeric Chalcone Synthase Gene into Petunia Results in Reversible Co-Suppression of Homologous Genes in trans. The Plant cell 2, 279 (Apr, 1990).
162. A. Fire et al., Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806 (Feb 19, 1998).
163. S. M. Hammond, E. Bernstein, D. Beach, G. J. Hannon, An RNA-directed nuclease mediates post-transcriptional gene silencing in Drosophila cells. Nature 404, 293 (Mar 16, 2000).
164. J. R. Kennerdell, R. W. Carthew, Use of dsRNA-mediated genetic interference to demonstrate that frizzled and frizzled 2 act in the wingless pathway. Cell 95, 1017 (Dec 23, 1998).
165. M. Ramet, P. Manfruelli, A. Pearson, B. Mathey-Prevot, R. A. Ezekowitz, Functional genomic analysis of phagocytosis and identification of a Drosophila receptor for E. coli. Nature 416, 644 (Apr 11, 2002).
166. S. Cherry et al., Genome-wide RNAi screen reveals a specific sensitivity of IRES-containing RNA viruses to host translation inhibition. Genes & development 19, 445 (Feb 15, 2005).
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
92
167. R. DasGupta, A. Kaykas, R. T. Moon, N. Perrimon, Functional genomic analysis of the
Wnt-wingless signaling pathway. Science 308, 826 (May 6, 2005). 168. G. Dietzl et al., A genome-wide transgenic RNAi library for conditional gene
inactivation in Drosophila. Nature 448, 151 (Jul 12, 2007). 169. E. Bernstein, A. A. Caudy, S. M. Hammond, G. J. Hannon, Role for a bidentate
ribonuclease in the initiation step of RNA interference. Nature 409, 363 (Jan 18, 2001). 170. A. Nykanen, B. Haley, P. D. Zamore, ATP requirements and small interfering RNA
structure in the RNA interference pathway. Cell 107, 309 (Nov 2, 2001). 171. R. H. Nicholson, A. W. Nicholson, Molecular characterization of a mouse cDNA
encoding Dicer, a ribonuclease III ortholog involved in RNA interference. Mammalian genome : official journal of the International Mammalian Genome Society 13, 67 (Feb, 2002).
172. P. J. Paddison, G. J. Hannon, RNA interference: the new somatic cell genetics? Cancer cell 2, 17 (Jul, 2002).
173. V. Hartenstein, Atlas of Drosophila development. (Cold Spring Harbour Laboratory Press, 1993).
174. M. Orme, P. Meier, Inhibitor of apoptosis proteins in Drosophila: gatekeepers of death. Apoptosis : an international journal on programmed cell death 14, 950 (Aug, 2009).
175. S. Morais da Silva, personal communication. 176. M. Calleja, E. Moreno, S. Pelaz, G. Morata, Visualization of gene expression in living
adult Drosophila. Science 274, 252 (Oct 11, 1996). 177. B. Winnepenninckx, Y. Van de Peer, T. Backeljau, R. De Wachter, CARD: a drawing tool
for RNA secondary structure models. BioTechniques 18, 1060 (Jun, 1995). 178. W. Huang da, B. T. Sherman, R. A. Lempicki, Systematic and integrative analysis of
large gene lists using DAVID bioinformatics resources. Nature protocols 4, 44 (2009). 179. D. A. Hosack, G. Dennis, Jr., B. T. Sherman, H. C. Lane, R. A. Lempicki, Identifying
biological themes within lists of genes with EASE. Genome biology 4, R70 (2003). 180. L. Ciapponi, G. Cenci, Telomere capping and cellular checkpoints: clues from fruit flies.
Cytogenetic and genome research 122, 365 (2008). 181. L. Giot et al., A protein interaction map of Drosophila melanogaster. Science 302, 1727
(Dec 5, 2003). 182. G. Goshima et al., Genes required for mitotic spindle assembly in Drosophila S2 cells.
Science 316, 417 (Apr 20, 2007). 183. C. A. Stanyon et al., A Drosophila protein-interaction map centered on cell-cycle
regulators. Genome biology 5, R96 (2004). 184. J. Dobbelaere et al., A genome-wide RNAi screen to dissect centriole duplication and
centrosome maturation in Drosophila. PLoS biology 6, e224 (Sep 16, 2008). 185. A. Lunardi et al., A genome-scale protein interaction profile of Drosophila p53
uncovers additional nodes of the human p53 network. Proceedings of the National Academy of Sciences of the United States of America 107, 6322 (Apr 6, 2010).
186. J. Mata, S. Curado, A. Ephrussi, P. Rorth, Tribbles coordinates mitosis and morphogenesis in Drosophila by regulating string/CDC25 proteolysis. Cell 101, 511 (May 26, 2000).
187. M. Nie, Y. Xie, J. A. Loo, A. J. Courey, Genetic and proteomic evidence for roles of Drosophila SUMO in cell cycle control, Ras signaling, and early pattern formation. PloS one 4, e5905 (2009).
188. S. J. Wall, E. Werner, Z. Werb, Y. A. DeClerck, Discoidin domain receptor 2 mediates tumor cell cycle arrest induced by fibrillar collagen. The Journal of biological chemistry 280, 40187 (Dec 2, 2005).
189. E. Kurucz et al., Nimrod, a putative phagocytosis receptor with EGF repeats in Drosophila plasmatocytes. Current biology : CB 17, 649 (Apr 3, 2007).
FCUP/ICBAS Spindle Assembly Checkpoint And Chromosomal Instability
93
190. E. Formstecher et al., Protein interaction mapping: a Drosophila case study. Genome
research 15, 376 (Mar, 2005). 191. M. Muzzopappa, P. Wappner, Multiple roles of the F-box protein Slimb in Drosophila
egg chamber development. Development 132, 2561 (Jun, 2005). 192. K. Koles, K. D. Irvine, V. M. Panin, Functional characterization of Drosophila
sialyltransferase. The Journal of biological chemistry 279, 4346 (Feb 6, 2004). 193. M. Miron et al., The translational inhibitor 4E-BP is an effector of PI(3)K/Akt signalling
and cell growth in Drosophila. Nature cell biology 3, 596 (Jun, 2001). 194. L. S. Goldstein, S. Gunawardena, Flying through the drosophila cytoskeletal genome.
The Journal of cell biology 150, F63 (Jul 24, 2000). 195. L. R. Zukerberg et al., Loss of cables, a cyclin-dependent kinase regulatory protein, is
associated with the development of endometrial hyperplasia and endometrial cancer. Cancer research 64, 202 (Jan 1, 2004).
196. C. Frohlich et al., ADAM12 produced by tumor cells rather than stromal cells accelerates breast tumor progression. Molecular cancer research : MCR 9, 1449 (Nov, 2011).
197. P. J. Chua et al., Silencing the Peroxiredoxin III gene inhibits cell proliferation in breast cancer. International journal of oncology 36, 359 (Feb, 2010).
198. K. D. McClure, A. Sustar, G. Schubiger, Three genes control the timing, the site and the size of blastema formation in Drosophila. Developmental biology 319, 68 (Jul 1, 2008).
199. K. H. Siller, M. Serr, R. Steward, T. S. Hays, C. Q. Doe, Live imaging of Drosophila brain neuroblasts reveals a role for Lis1/dynactin in spindle assembly and mitotic checkpoint control. Molecular biology of the cell 16, 5127 (Nov, 2005).
200. N. Tapon, N. Ito, B. J. Dickson, J. E. Treisman, I. K. Hariharan, The Drosophila tuberous sclerosis complex gene homologs restrict cell growth and cell proliferation. Cell 105, 345 (May 4, 2001).
201. E. Mullarky, K. R. Mattaini, M. G. Vander Heiden, L. C. Cantley, J. W. Locasale, PHGDH amplification and altered glucose metabolism in human melanoma. Pigment cell & melanoma research 24, 1112 (Dec, 2011).
202. J. W. Locasale et al., Phosphoglycerate dehydrogenase diverts glycolytic flux and contributes to oncogenesis. Nature genetics 43, 869 (Sep, 2011).
203. C. X. Zhang, A. D. Chen, N. J. Gettel, T. S. Hsieh, Essential functions of DNA topoisomerase I in Drosophila melanogaster. Developmental biology 222, 27 (Jun 1, 2000).
204. M. Sereno et al., ERCC1 and topoisomerase I expression in small cell lung cancer: prognostic and predictive implications. International journal of oncology 40, 2104 (Jun, 2012).
205. S. Kondo, N. Perrimon, A genome-wide RNAi screen identifies core components of the G(2)-M DNA damage checkpoint. Science signaling 4, rs1 (2011).
206. S. L. Stroschein-Stevenson, E. Foley, P. H. O'Farrell, A. D. Johnson, Identification of Drosophila gene products required for phagocytosis of Candida albicans. PLoS biology 4, e4 (Jan, 2006).
207. R. Artero, E. E. Furlong, K. Beckett, M. P. Scott, M. Baylies, Notch and Ras signaling pathway effector genes expressed in fusion competent and founder cells during Drosophila myogenesis. Development 130, 6257 (Dec, 2003).