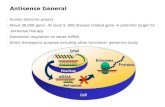
Extensive microRNA-mediated crosstalk between lncRNAs and ...
Knockdown of lncRNAs: exploring RNAi and antisense oligo methods
-
Upload
integrated-dna-technologies -
Category
Science
-
view
2.427 -
download
0
Transcript of Knockdown of lncRNAs: exploring RNAi and antisense oligo methods

Kim Lennox, MScResearch Scientist, Integrated DNA Technologies
Methods to knock down lncRNAs
1

• Lower organisms: mRNA > lncRNA• Higher organisms: lncRNA >>
mRNA• Added regulatory functions of
lncRNAs:– Correlate with an increased
ability of multicellular organisms to differentiate into many different cells types
– Allow an organism to achieve greater diversity from the same number of protein coding genes
Importance of lncRNAs in mammals
2

Target LocationmRNA cytoplasm + nucleussplice nucleus onlymiRNA cytoplasm > nucleus
lncRNAcytoplasm > nucleuscytoplasm = nucleuscytoplasm < nucleus
Does cellular localization matter when choosing a silencing reagent?
3

Rinn study: RNA-FISH on 61 lncRNAs
4
Cabili et al. (2015) Localization and abundance of human lncRNAs at single-cell and single molecule resolution. Genome Biol, 16:20.
Cell lines: HeLa, hlF, hFF

5
RISC
Target RNA
RNA cleavage
siRNA
RNA cleavage
Target RNA
RNA interference (RNAi)
Target RNA
RNase H1
ASO (DNA)
Antisense oligonucleotides (ASOs)
> cytoplasm > nucleus

Study design• lncRNA targets
– 2 nuclear– 3 cytoplasmic– 3 nuclear + cytoplasmic
• Knockdown reagents– Antisense oligonucleotides (ASOs)
• 12 DNA-PS (20mers)• 12 2′OMe-PS 5-10-5 chimeras (same sequence as DNA-PS)• 6 LNA™ longRNA GapmeRs (sites selected by Exiqon)
– siRNA• 12 Dicer-substrate siRNAs (DsiRNAs) (27mer, IDT design)• 12 siRNAs (21mer, Thermo Fisher design)• 4 Silencer® Select siRNAs (21mer, Life Technologies design)
6

The knockdown reagents
7

Methods• Transfection (ASOs and siRNAs):
– HeLa cells (Huh7 for NRON); all data replicated in HCT116 cells – Biological triplicates – 96-well format – Lipofectamine® 2000– Chemistry-matched negative control sequences
• RNA was prepared 24 hours post-transfection• RT-qPCR:
– Two qPCR assays for each target (one towards the 5′ end and one towards the 3′ end)
– Triplicate qPCRs – Quantification standard curves on each 384-well plate – Results normalized against internal control HPRT and SFRS9 gene expression levels
8

Targets
Nuclear Cytoplasmic Nuclear + Cytoplasmic
Target 1: MALAT1 (8 kb)
Target 3: NRON (2.7 kb)
Target 6: TUG1 (7.5 kb)
Target 2: NEAT1 (3.7 kb)
Target 4: DANCR (0.9 kb)
Target 7: CasC7 (9.3 kb)
Target 5: OIP5-AS1 (1.9 kb)
Target 8: HOTAIR (2.3 kb)
9

In situ hybridization: MALAT1 (nuclear)
10

In situ hybridization: NEAT1 (nuclear)
11

In situ hybridization: CasC7 (nuclear + cytoplasmic)
12

MALAT1 knockdown in HeLa cells: nuclear
For mRNA knockdown, we usually expect >80% success rate for the DsiRNA algorithm.
13

NEAT1 knockdown in HeLa cells: nuclear
14

NRON knockdown in Huh-7 cells: cytoplasmic
NRON function involves tight binding of multiple protein species → no knockdown.Therefore, needed to move to a different cytoplasmic target. 15

DANCR knockdown in HeLa cells: cytoplasmic
16

OIP5-AS1 knockdown in HeLa cells: cytoplasmic
17

TUG1 knockdown in HeLa cells: nuclear + cytoplasmic
18

CasC7 knockdown in HeLa cells: nuclear + cytoplasmic
19

HOTAIR knockdown in HeLa cells: nuclear + cytoplasmic
20

Comparison summary
21
2′OMe-PS ASOLNA-PS ASO
LNA-siRNA
DsiRNAsiRNA

Combinatorial approach can have additive effects for lncRNAs localized in both the nucleus and cytoplasm
22
ASO DsiRNA ASO + DsiRNA
ASO DsiRNA ASO + DsiRNA
ASO DsiRNA ASO + DsiRNA
0
20
40
60
80
100
120
Combinatorial Knockdown10 nM5 nM2.5 nM1 nM
% R
emai
ning
RNA
MALAT1Nuclear
OIP5-AS1Cytoplasmic
CasC7Both

Comparison of ASO chemistry/design potency at the same 6 sites in MALAT1
2nd generation “Gapmer” ASOs are more potent than DNA-PS 23

Summary• Overall performance of ASOs vs. siRNAs varies with the
dominant cellular localization of the lncRNAs that were targeted:– ASOs were more often effective at knocking down nuclear
lncRNAs.– siRNAs were more often effective at knocking down cytoplasmic
lncRNAs.– Better knockdown can be achieved by combining RNAi reagents
with ASOs. • Characterizing the localization of a targeted lncRNA and
selecting the appropriate knockdown method is important for improving the success of the knockdown experiment.
• If lncRNA localization is unknown, trying both methods may be prudent.
24

Articles from IDT scientists:
• www.idtdna.com/decoded (Small RNAs/Functional Genomics section)– A New Renaissance for Antisense in the Era of lncRNA– Using Antisense Technologies to Modulate Noncoding RNA
Function• www.idtdna.com (Search for “antisense
oligonucleotide”)– Antisense Oligonucleotides: Strategies and Applications
25

Online tools:
• www.idtdna.com/scitools (Gene Regulation and Knockdown section)– Predesigned DsiRNA Selection ToolSelects DsiRNA Duplexes and
TriFECTa® Screening Kits for your sequence– RNAi Design ToolGenerates duplex siRNA sequences for RNAi
applications
• For additional ASO design assistance, please email [email protected]
26