Expected accuracy sequence alignment
-
Upload
beck-patel -
Category
Documents
-
view
32 -
download
2
description
Transcript of Expected accuracy sequence alignment

Expected accuracy sequence alignment
Usman Roshan

Optimal pairwise alignment
• Sum of pairs (SP) optimization: find the alignment of two sequences that maximizes the similarity score given an arbitrary cost matrix. We can find the optimal alignment in O(mn) time and space using the Needleman-Wunsch algorithm.
• Recursion: Traceback:
where M(i,j) is the score of the optimal alignment of x1..i and y1..j, s(xi,yj) is a substitution scoring matrix, and g is the gap penalty

Affine gap penalties
• Affine gap model allows for long insertions in distant proteins by charging a lower penalty for extension gaps. We define g as the gap open penalty (first gap) and e as the gap extension penalty (additional gaps)
• Alignment:– ACACCCT ACACCCC– AC-CT-T AC--CTT– Score = 0 Score = 0.9
• Trivial cost matrix: match=+1, mismatch=0, gapopen=-2, gapextension=-0.1

Affine penalty recursionM(i,j) denotes alignments of x1..i and y1..j ending witha match/mismatch. E(i,j) denotes alignments of x1..i
and y1..j such that yj is paired with a gap. F(i,j) definedsimilarly. Recursion takes O(mn) time where m and n are lengths of x and y respectively.

Expected accuracy alignment
• The dynamic programming formulation allows us to find the optimal alignment defined by a scoring matrix and gap penalties. This may not necessarily be the most “accurate” or biologically informative.
• We now look at a different formulation of alignment that allows us to compute the most accurate one instead of the optimal one.

Posterior probability of xi aligned to yj
• Let A be the set of all alignments of sequences x and y, and define P(a|x,y) to be the probability that alignment a (of x and y) is the true alignment a*.
• We define the posterior probability of the ith residue of x (xi) aligning to the jth residue of y (yj) in the true alignment (a*) of x and y as
Do et. al., Genome Research, 2005

Expected accuracy of alignment
• We can define the expected accuracy of an alignment a as
• The maximum expected accuracy alignment can be obtained by the same dynamic programming algorithm
Do et. al., Genome Research, 2005

Example for expected accuracy
• True alignment• AC_CG• ACCCA• Expected accuracy=(1+1+0+1+1)/4=1
• Estimated alignment• ACC_G• ACCCA• Expected accuracy=(1+1+0.1+0+1)/4 ~ 0.75

Estimating posterior probabilities• If correct posterior probabilities can be computed
then we can compute the correct alignment. Now it remains to estimate these probabilities from the data
• PROBCONS (Do et. al., Genome Research 2006): estimate probabilities from pairwise HMMs using forward and backward recursions (as defined in Durbin et. al. 1998)
• Probalign (Roshan and Livesay, Bioinformatics 2006): estimate probabilities using partition function dynamic programming matrices

HMM posterior probabilities• Consider the probability of all alignments of sequences X
and Y under a given HMM.• Let M(i,j) be the sum of the probabilities of all alignments of
X1...i and Y1…j that end in match or mismatch.
• Then M(i,j) is given by
• We calculate X(i,j) and Y(i,j) in the same way.• We call these forward probabilities:
– f(i,j) = M(i,j)+X(i,j)+Y(i,j)
(1 2 ) ( 1, 1)
( , ) (or ) (1 ) ( 1, 1)
(1 ) ( 1, 1)m mm
M i j
M i j p p X i j
Y i j

HMM posterior probabilities
• Similarly we can calculate backward probabilties M’(i,j). • Define M’(i,j) as the sum of probabilities of all alignments of
Xi..m and Yj..n such that Xi and and Yj are aligned to each other.
• The indices i and j start from m and n respectively and decrease
• These are also called backward probabilities.– B(i,j)=M’(i,j)+X’(i,j)+Y’(i,j)
(1 2 ) '( 1, 1)
'( , ) (or ) (1 ) '( 1, 1)
(1 ) '( 1, 1)m mm
M i j
M i j p p X i j
Y i j

HMM posterior probabilities
• The posterior probability of xi aligned to yj is given by
( ) ( , ) ( , ) / ( , y)i jP x y f i j b i j P x

Partition function posterior probabilities
• Standard alignment score:
• Probability of alignment (Miyazawa, Prot. Eng. 1995)
• If we knew the alignment partition function then

Partition function posterior probabilities
• Alignment partition function (Miyazawa, Prot. Eng. 1995)
• Subsequently

Partition function posterior probabilities
• More generally the forward partition function matrices are calculated as

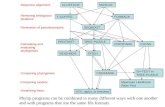
Partition function matrices vs. standard affine recursions

Posterior probability calculation
• If we defined Z’ as the “backward” partition function matrices then

Posterior probabilities using alignment ensembles
• By generating an ensemble A(n,x,y) of n alignments of x and y we can estimate P(xi~yj) by counting the number of times xi is aligned to yj.. Note that this means we are assigning equal weights to all alignments in the ensemble.

Generating ensemble of alignments
• We can use stochastic backtracking (Muckstein et. al., Bioinformatics, 2002) to generate a given number of optimal and suboptimal alignments.
• At every step in the traceback we assign a probability to each of the three possible positions.
• This allows us to “sample” alignments from their partition function probability distribution.
• Posteror probabilities turn out to be the same when calculated using forward and backward partition function matrices.

Probalign1. For each pair of sequences (x,y) in the input set
– a. Compute partition function matrices Z(T)– b. Estimate posterior probability matrix P(xi ~ yj) for (x,y)
by
2. Perform the probabilistic consistency transformation and compute a maximal expected accuracy multiple alignment: align sequence profiles along a guide-tree and follow by iterative refinement (Do et. al.).

Experimental results
• http://bioinformatics.oxfordjournals.org/content/26/16/1958