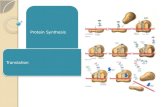
Chapter 17 From Gene to Protein (Protein Synthesis) From Gene to Protein (Protein Synthesis)
AP Details for Protein Synthesis
-
Upload
samantha-mejia -
Category
Documents
-
view
20 -
download
2
description
Transcript of AP Details for Protein Synthesis

AP Details for Protein Synthesis
2014

From gene to protein

AP Biology
Transcriptionfrom
DNA nucleic acid languageto
RNA nucleic acid language

Transcription• Making mRNA– transcribed DNA strand = template strand– untranscribed DNA strand = coding strand
• same sequence as RNA– synthesis of complementary RNA strand
• transcription bubble– Enzyme involved
• RNA polymerase
template strand
rewinding
mRNA RNA polymerase
unwinding
coding strand
DNAC C
C
C
C
C
C
C
C CC
G
GG
G
G G
G G
G
G
GAA
AA A
A
A
A
A
A A
A
AT
T T
T
T
T
T
T
T T
T
T
U U
5
35
3
3
5build RNA 53

RNA polymerases• 3 RNA polymerase enzymes– RNA polymerase 1• only transcribes rRNA genes• makes ribosomes
– RNA polymerase 2• transcribes genes into mRNA
– RNA polymerase 3• only transcribes tRNA genes
– each has a specific promoter sequence it recognizes

Which gene is read?• Promoter region– binding site before beginning of gene – TATA box binding site– binding site for RNA polymerase
& transcription factors
– Enhancer region– binding site for
activators (activate genes)
– Silence region– Binding site for
repressors (turns genesoff)

What are Transcription Factors?• Initiation complex– transcription factors bind to promoter region
• suite of proteins which bind to DNA• turn on or off transcription
– trigger the binding of RNA polymerase to DNA

Matching bases of DNA & RNA• Match RNA bases to DNA
bases on one of the DNA strands
U
A G GGGGGT T A C A C T T T T TC C C CA A
U
UU
U
U
G
G
A
A
A C CRNA polymerase
C
C
C
C
C
G
GG
G
A
A
A
AA
5' 3'

Eukaryotic genes have junk!• Eukaryotic genes are not continuous– exons = the real gene• expressed / coding DNA
– introns = the junk• inbetween sequence
eukaryotic DNA
exon = coding (expressed) sequence
intron = noncoding (inbetween) sequence
intronscome out!

mRNA splicing
eukaryotic DNA
exon = coding (expressed) sequence
intron = noncoding (inbetween) sequence
primary mRNAtranscript
mature mRNAtranscript
pre-mRNA
spliced mRNA
• Post-transcriptional processing – eukaryotic mRNA needs work after transcription– primary transcript = pre-mRNA– mRNA splicing• edit out introns
– make mature mRNA transcript
~10,000 bases
~1,000 bases

Splicing must be accurate• No room for mistakes!– a single base added or lost throws off the reading
frame
AUG|CGG|UCC|GAU|AAG|GGC|CAU
AUGCGGCTATGGGUCCGAUAAGGGCCAUAUGCGGUCCGAUAAGGGCCAU
AUG|CGG|GUC|CGA|UAA|GGG|CCA|U
AUGCGGCTATGGGUCCGAUAAGGGCCAUAUGCGGGUCCGAUAAGGGCCAU
Met|Arg|Ser|Asp|Lys|Gly|His
Met|Arg|Val|Arg|STOP|

RNA splicing enzymes
snRNPs
exonexon intron
snRNA
5' 3'
spliceosome
exonexcisedintron
5'
5'
3'
3'
3'
lariat
exonmature mRNA
5'
No, not smurfs!“snurps”
• snRNPs– small nuclear RNA
• Spliceosome– several snRNPs– recognize splice site
sequence• cut & paste gene

A A AA
A3' poly-A tail
mRNA
5'5' cap
3'
G PPP
50-250 A’s
More post-transcriptional processing
• Need to protect mRNA on its trip from nucleus to cytoplasm– enzymes in cytoplasm attack mRNA
• protect the ends of the molecule• add 5 GTP cap
– Chemically modified molecule of GTP– It facilitates the binding of mRNA to the ribosome and protects the mRNA
from being digested by ribonucleases – enzymes in cytoplasm that break down RNA
• add 3’poly-A tail– longer tail – 100-300 adenine nucleotides– Assists in export of mRNA from nucleus – Important in mRNA stability

Radjewski
Translationfrom
nucleic acid languageto
amino acid language

The code• Code for ALL life!
– strongest support for a common origin for all life
• Code is redundant– several codons for each
amino acid– 3rd base “wobble”
Start codon AUG methionine
Stop codons UGA, UAA, UAG

Transfer RNA structure• “Clover leaf” structure– anticodon on “clover leaf” end– amino acid attached on 3 end

Loading tRNA • Aminoacyl tRNA synthetase – enzyme which bonds amino acid to tRNA– bond requires energy
• ATP AMP• bond is unstable• so it can release amino acid at ribosome easily
activatingenzyme
anticodontRNATrp binds to UGG condon of mRNA
Trp Trp Trp
mRNAA C CU G G
C=OOH
OHH2OO
tRNATrp
tryptophan attached to tRNATrp
C=O
O
C=O

Building a polypeptide• Initiation
– brings together mRNA, ribosome subunits, initiator tRNA
• Elongation– adding amino acids based on codon
sequence
• Termination– end codon 123
Leu
Leu Leu Leu
tRNA
Met MetMet Met
PE AmRNA
5' 5' 5' 5'3' 3' 3' 3'
U UA A AACC
CA U UG G
GUU
A AAAC
CC
A U UG GGU
UA A A
ACC
CA U UG G
GU UA A ACCA U UG G
G A C
Val Ser
Ala Trp
releasefactor
AA A
C CU UG G 3'

Protein targeting • Signal peptide– address label
Destinations: secretion nucleus mitochondria chloroplasts cell membrane cytoplasm etc…

Can you tell the story?
DNA
pre-mRNA
ribosome
tRNA
aminoacids
polypeptide
mature mRNA
5' GTP cap
poly-A tail
large ribosomal subunit
small ribosomal subunit
aminoacyl tRNAsynthetase
E P A
5'
3'
RNA polymerase
exon introntRNA

AP Biology
Protein Synthesis in Prokaryotes
Bacterial chromosome
mRNA
Cell wall
Cellmembrane
Transcription
Psssst…no nucleus!

Prokaryote vs. Eukaryote genes• Prokaryotes– DNA in cytoplasm– circular chromosome– naked DNA
– no introns
• Eukaryotes– DNA in nucleus– linear chromosomes– DNA wound on histone
proteins– introns vs. exons
eukaryoticDNA
exon = coding (expressed) sequence
intron = noncoding (inbetween) sequence
intronscome out!

• Transcription & translation are simultaneous in bacteria – DNA is in
cytoplasm– no mRNA
editing – ribosomes
read mRNA as it is being transcribed
Translation in Prokaryotes

Translation: prokaryotes vs. eukaryotes• Differences between prokaryotes &
eukaryotes– time & physical separation between processes• takes eukaryote ~1 hour
from DNA to protein– no RNA processing

AP Biology
Any Questions??What color would a smurf turnif he held his breath?