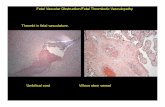
Using HLAMatchMaker: Maternal-Fetal Incompatibility and the Risk of Disease
-
Upload
vielka-cline -
Category
Documents
-
view
23 -
download
2
description
Transcript of Using HLAMatchMaker: Maternal-Fetal Incompatibility and the Risk of Disease

Using HLAMatchMaker:Using HLAMatchMaker: Maternal-Fetal Incompatibility and Maternal-Fetal Incompatibility and
the Risk of Diseasethe Risk of Disease
Miguel Angel GonzalezMiguel Angel Gonzalez
Mentor: Janet Sinsheimer, UCLAMentor: Janet Sinsheimer, UCLA
Hsinju Hsieh, UCLAHsinju Hsieh, UCLA

OutlineOutline
IntroductionIntroduction
HLAMatchMakerHLAMatchMaker
ResultsResults
ConclusionsConclusions

HLA IncompatibilityHLA Incompatibility
Human Leukocyte AntigenHuman Leukocyte AntigenPresence of structures on cells that are sites Presence of structures on cells that are sites
for antibody bindingfor antibody binding
ProblemsProblemsMother and fetus closely matchedMother and fetus closely matched
Fetus does not set up correct immunological Fetus does not set up correct immunological barrierbarrier

Experimental DataExperimental Data
Family StudiesFamily StudiesMother, father, and at least 1 child with Mother, father, and at least 1 child with
schizophreniaschizophreniaHLA-A, B, & DRB1 types were determined HLA-A, B, & DRB1 types were determined Study focuses on mother-fetus relationshipStudy focuses on mother-fetus relationship

Current StudiesCurrent Studies WHOWHO Current method Current method
for HLA comparisonfor HLA comparison
Biological Biological comparison using comparison using serum screeningserum screening
Match = 1Match = 1 Mismatch = 0Mismatch = 0
Chris O’Callaghan – University of Oxfordhttp://neelix.molbiol.ox.ac.uk:8080/userweb/cocallag/

HypothesisHypothesis
Incompatibility between the mother and Incompatibility between the mother and fetus with respect to HLA can be a risk fetus with respect to HLA can be a risk factor for neurological diseases such as factor for neurological diseases such as schizophrenia.schizophrenia.
Also, to quantify the level of mismatching Also, to quantify the level of mismatching and to determine how sensible the WHO and to determine how sensible the WHO method is.method is.

HLAMatchMakerHLAMatchMaker
Written by Rene J. Duquesnoy, Ph.D.Written by Rene J. Duquesnoy, Ph.D.
Computer algorithm for identifying Computer algorithm for identifying acceptable HLA mismatches for patients acceptable HLA mismatches for patients without the need for extensive serum without the need for extensive serum screening screening

HLAMatchMakerHLAMatchMaker
Purpose: to identify the best candidates for Purpose: to identify the best candidates for organ transplantationorgan transplantation
Compares the Compares the
amino acid amino acid
sequences of sequences of
HLA antigens.HLA antigens.

Internship GoalInternship Goal
To adapt HLAMatchMaker to the studies of the To adapt HLAMatchMaker to the studies of the Sinsheimer lab.Sinsheimer lab.
Use HLAMatchMaker to identify differences in Use HLAMatchMaker to identify differences in the amino acid sequences of mother and fetus the amino acid sequences of mother and fetus HLA.HLA.
Compare to the current analysis: Compare to the current analysis: WHO vs HLAMatchMakerWHO vs HLAMatchMaker
To determine mismatch severityTo determine mismatch severity

HLAMatchMaker ResultsHLAMatchMaker Results
HVG Compatibility #mmDonor
Donor types as Trp(9-199) Mismatched Trps in Positions 9-199
A*0201 0 , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , ,
A*2301 9 9S, , , , , , , ,62Ee,66GKv, ,74D,76En,80rIa,82aLr, , , , , , , , , , ,151aRv, , , ,166Dg, , , , , , ,
B*1501 0 , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , ,
B*4901 6 9H, , , ,41T,45Ke, , , , , , ,76En,80rIa,82aLr, , , , , , , , , , , , , , , , , , , , , ,
A*02
A*23
B*15 B*49
A*02
A*02
B*15
B*15
Child Mother

Visual ComparisonVisual Comparison
A*23 9S 62Ee 66GKv 74D 76En 80rIa 82aLr
A*02 9F 62Ge 66rKv 74H 76Vd 80gTl 82lRg
A*02
A*23
166Ew151aHv149aAh144tKh142T107W
166Dg151aRv149aAr144tQr142I107G

HLAMatchMaker WHO ComparisonHLAMatchMaker WHO Comparison
Using an excel macro (efficient), large sets Using an excel macro (efficient), large sets of data can be analyzed.of data can be analyzed.
To make comparable to WHO, To make comparable to WHO,
Match = 1Match = 1
Mismatch = 0Mismatch = 0
Results used to construct a comparison Results used to construct a comparison table (Pivot Table).table (Pivot Table).

Pivot TablePivot Table
Count of mmHLA-ACount of mmHLA-A match_amatch_a (WHO)(WHO)
mmHLA-AmmHLA-A 00 11 Grand TotalGrand Total
00 158158 158158
11 8383 8383
Grand TotalGrand Total 158158 8383 241241
HLAMatchMaker

ConclusionConclusion
Goal AchievedGoal AchievedUsing excel macros it was possible to adapt Using excel macros it was possible to adapt
HLAMatchMaker to the Sinsheimer lab HLAMatchMaker to the Sinsheimer lab studies.studies.
Results showed no difference between Results showed no difference between WHO and HLAMatchMakerWHO and HLAMatchMaker
HLAMatchMaker results can be used to HLAMatchMaker results can be used to support WHO findingssupport WHO findings

Future StudiesFuture Studies
Determine the severity of mismatchingDetermine the severity of mismatchingDifferences in amino acidsDifferences in amino acids

AcknowledgementsAcknowledgements
Dr. Janet Sinsheimer, UCLADr. Janet Sinsheimer, UCLADr. Christina Palmer, UCLADr. Christina Palmer, UCLAHsinju Hsieh, UCLAHsinju Hsieh, UCLA
FundingFundingNIHNIHNIMHNIMHSoCalBSISoCalBSI