University of Groningen Translocation across biological ... the “short” vs. “extended”...
Transcript of University of Groningen Translocation across biological ... the “short” vs. “extended”...

University of Groningen
Translocation across biological membranes: activity, structure and regulation of transportersRuiz, Stephanie
IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite fromit. Please check the document version below.
Document VersionPublisher's PDF, also known as Version of record
Publication date:2017
Link to publication in University of Groningen/UMCG research database
Citation for published version (APA):Ruiz, S. (2017). Translocation across biological membranes: activity, structure and regulation oftransporters [Groningen]: University of Groningen
CopyrightOther than for strictly personal use, it is not permitted to download or to forward/distribute the text or part of it without the consent of theauthor(s) and/or copyright holder(s), unless the work is under an open content license (like Creative Commons).
Take-down policyIf you believe that this document breaches copyright please contact us providing details, and we will remove access to the work immediatelyand investigate your claim.
Downloaded from the University of Groningen/UMCG research database (Pure): http://www.rug.nl/research/portal. For technical reasons thenumber of authors shown on this cover page is limited to 10 maximum.
Download date: 16-05-2018

19
Author contributions: Stephanie J. Ruiz implemented the P. pastoris pPICZ expression system (with initial supervision by Ina L. Urbatsch); introduced the pBGP1 expression system; cloned pBGP1na and the pSR plasmids; performed expression and purification screening for Can1, Lyp1 and Gap1; supervised Masters student Łukasz Niemiec during cloning and screening of the VBA proteins; developed the “short” vs. “extended” theory for Vba2 and Vba3; and wrote the chapter.
Some of the work in this chapter was published in:Bianchi F, Klooster JS, Ruiz SJ, Luck K, Pols T, Urbatsch IL, Poolman B (2016) Asymmetry in inward- and outward-affinity constant of transport explain unidirectional lysine flux in Saccharomyces cerevisiae. Sci Rep 6: 31443.
Chapter 2
Expression of Saccharomyces cerevisiae amino acid transporters in Pichia pastoris


21
Chapter 2
2.1 Pichia pastoris as an expression hostThe methylotrophic yeast Pichia pastoris has been used for several decades as a
heterologous host for protein expression and purification (reviewed in [149-154]). Although the most commonly used laboratory strains have since been reclassified as Komagataella phaffii [155], the name Pichia pastoris is still used.
Successful expression of eukaryotic proteins, and in particular membrane proteins, most often occurs in other eukaryotic hosts due to the need for co- and post-translational modifications (e.g. glycosylation and disulfide bond formation), as well as specific lipid requirements. Based on the availability of 3D structures, Pichia pastoris is the second-most successful expression host for the production of eukaryotic membrane proteins (http://blanco.biomol.uci.edu/mpstruc/, [154,156]). These structures include several high-profile mammalian proteins such G-protein coupled receptors and P-glycoproteins. The most successful expression system, baculovirus-infected insect cells, is slower and more expensive compared to P. pastoris [156]. One of the specific advantages of P. pastoris is that it can be grown to extremely high cell densities (OD600 of ~500 in bioreactors). Increased biomass is particularly important in the production of membrane proteins, which are normally expressed at low yields.
After attempts to express S. cerevisiae amino acid transporters at high levels endogenously were unsuccessful (Frans Bianchi, personal communication), we decided to screen expression constructs in P. pastoris.
2.2 Screening amino acid transporter constructsS. cerevisiae proteins were heterogeneously expressed in P. pastoris using two
different systems. The pPICZ system from Invitrogen (now part of Thermo Fisher Scientific, Waltham, MA, USA) results in methanol-induced expression from chromosomally integrated expression cassettes. The other system, based on the pBGP1 plasmid developed by Lee et al. [157], involves constitutive expression from an episomal plasmid. The pPICZ system is designed to generate stable expression strains that produce the highest possible levels of recombinant protein. It is not suitable for more high-throughput screening, and it is for this reason that I also developed a plasmid-based expression system.

22
Expressing S. cerevisiae transporters in P. pastoris
2.2.1 Inducible expression based on pPICZIn the pPICZ vectors (Fig. 2.1), the gene of interest is cloned in-frame behind
a 5’ AOX1 fragment, which contains the promoter sequence for the tightly-regulated, methanol-inducible alcohol oxidase gene AOX1. Glucose or glycerol repress the expression of alcohol oxidase I, which catalyzes the first step in methanol utilization, while methanol induces high-level expression (≥ 30% of total protein) [158]. The pPICZ vectors do not contain a yeast origin of replication, and instead are designed to integrate into the chromosome via homologous recombination. Linearizing the plasmid by restriction digest within the 5’ AOX1 sequence promotes recombination within the P. pastoris chromosome at this site. The AOX1 sequence is maintained after recombination, meaning that multiple insertion events can occur within a single cell. Transformant strains can thus contain multiple genomic copies of both the expression and resistance cassettes. Screening on increasing amounts of zeocin allows for selection of strains with higher zeocin resistance and thus (potentially) higher expression levels.2.2.2 Constitutive expression based on pBGP1
The P. pastoris episomal expression vector pBGP1 was designed for high-throughput screening of mutant gene libraries [157]. Target proteins are expressed from the strong constitutive GAP (glyceraldehyde-3-phosphate dehydrogenase) promoter. Inclusion of the P. pastoris autonomous replication sequence PARS1 means that the vector is stably maintained as a single-copy plasmid. The AmpR
CYC
1 TT ZeoR PTEF1 A
OX1 TT
p
UC ori
5’ AOX1
pPICZ A, B, C3.3 kbp
PEM7
pUC ori CYC1 TT ZeoR
PTE
F1
Am
pR
PGAP AOX1 TT
pBGP1na4.3 kbp
PARS1
PEM
7
Figure 2.1 Plasmid vectors used for expression in P. pastoris. The pPICZ plasmids allow high-level, methanol-inducible expression from the promoter for AOX1 (alcohol oxidase 1). pBGP1na allows for constitutive expression from the promoter for GAP (glyceraldehyde-3-phosphate dehydrogenase). Promoter and terminator sequences are white, genes coding for resistance proteins are grey, and replication origins are black. The pUC origin of replication allows the plasmid to be maintained in multiple copies in E. coli, while the PARS1 (Pichia autonomous replication sequence 1) allows for the plasmid to be stably maintained in P. pastoris. The promoters PEM7 and PTEF1 allow for expression of the zeocin resistance gene in both E. coli and P. pastoris, respectively.

23
Chapter 2
resistance gene allows for selection in E. coli using ampicillin, which is more stable and economical than zeocin. Because I did not require secretion of the recombinant protein into the growth media, I removed the a-factor signal sequence to produce pBGP1na (Fig. 2.1).
The introduction of this constitutive expression system was the catalyst for the discovery of the amino acid sensitivity phenotype described in Chapter 3. I first tested the system using several constructs that I had already successfully expressed in P. pastoris. I found that cells expressing Can1 grew poorly, and discovered that this was due to the switch from minimal media to YPD. After a few simple growth experiments, I determined that the growth defect was specific to the presence of arginine and lysine.
Plasmids based on pBGP1 were used in Chapter 4 to show that the novel properties of C-terminal peptides from S. cerevisiae amino acid transporters also occur in P. pastoris.2.2.3 Screening S. cerevisiae amino acid transporters for expression in P. pastoris
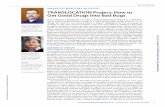
Three plasma membrane amino acid transporters (Gap1, Can1, Lyp1) as well as five reported or putative vacuolar amino acid transporters (Vba1, -2, -3, -4, and -5) were cloned into pPICZ A and transformed into P. pastoris KM71H. Each protein was C-terminally tagged with GFP (Green Fluorescent Protein) in order to allow fluorescence-based optimization of expression and purification as described in Drew et al. [159]. Three colonies each from selection plates with low, medium, or high zeocin concentrations were tested in small-scale expression cultures. The cells were imaged using confocal fluorescence microscopy to determine whether the constructs were successfully expressed and localized (Fig. 2.2).
Vba2 and Vba3 constructs resulted in extremely low fluorescence levels, with no clear localization pattern (further discussed in Section 2.2.4). In Can1- and Lyp1-expressing cells, the fluorescence signal was localized to the cell periphery as well as one or more circular internal structures with indistinct edges (Fig. 2.2). We judged that this represents plasma membrane localization of full-length protein, as well as persistence of free GFP in the vacuolar lumen after protein degradation. Whole cell lysine uptake assays confirmed that the recombinant Can1 and Lyp1 are functional in vivo (Fig. 2.3).
The fluorescence signal from Vba5-expressing cells was indistinguishable from that of Can1 or Lyp1 cells. At the time this was surprising, but it has since been reported that Vba5 is in fact a plasma membrane protein involved in amino acid uptake and drug sensitivity [56]. Expression of our Vba5 construct did not increase the lysine uptake above that of the KM71H control (Fig. 2.3), but the substrate concentrations used here were 4- to 20-fold lower than the 0.4 mM used in Shimazu et al. [56].
The fluorescence signal in Gap1 and Vba4 cells showed similar localization patterns: the vacuolar lumen, as well as a single internal ring structure and a patchy

24
Expressing S. cerevisiae transporters in P. pastoris
stain at the cell periphery. We judge that this represents localization to internal membranes, in particular the perinuclear and peripheral endoplasmic reticulum. The presence of Gap1 in internal membranes is not unusual. Gap1 activity is regulated by trafficking of the full-length protein between the plasma membrane and internal compartments (see references [143-146] and Section 1.4). Vacuolar membrane localization of fluorescently-tagged Vba4 in S. cerevisiae has been
0 1 2 30.0
0.2
0.4
0.6
0.8
nmol
.OD−1
Time (min)0 2 4 6 8 10
0.0
0.5
1.0
1.5
2.0
2.5
Time (min)
20 μM lysine 100 μM lysine
KM71H
Can1
Lyp1
KM71H
Can1
Lyp1
Vba5
Figure 2.3 Uptake of lysine by P. pastoris KM71H (diamonds) or strains expressing Can1 (grey circles), Lyp1 (black circles), or Vba5 (white circles). In most cases the data points of the KM71H control obscure the ones for Vba5.
Can1 Lyp1 Vba5
Vba1Gap1 Vba4
Figure 2.2 Confocal fluorescence microscopy images of KM71H cells expressing GFP-tagged constructs. Cells shown are from a single, representative culture. Solid black lines in the Vba1 panel represent the cell outline (in all other cases the fluorescence signal outlines the periphery of the cell). Scale bars are 2 µm.

25
Chapter 2
recently reported [58]. In all the strains we tested, the fluorescence signal from Vba4 was very low, and it is possible this is due to problems with either heterogeneous expression in P. pastoris, or the specific GFP tag used.
Vba1 cells displayed fluorescence signal at the vacuolar membrane as well as in the lumen. Although the membrane signal is not as clear as that observed for the plasma membrane in other strains, the appearance of the vacuoles in Vba1 cells is distinct from the fuzzy-edged appearance of the vacuoles in, for example, Can1 cells.
Fluorescent species corresponding to full-length Can1 (95.5 kDa), Lyp1 (97.7 kDa), Vba1 (92.4 kDa) and Vba5 (93.0 kDa) were clearly visible in crude membrane fractions extracted from expression cultures (Fig. 2.4A). In all cases the recombinant proteins migrated 10–20 kDa lower than would be expected based on their predicted molecular weights; this is likely due to the GFP tag remaining folded even under denaturing conditions [160]. There was also a greater variation between proteins than expected, possibly due to post-translational modifications. Numerous lower-molecular weight bands were visible, particularly for Vba1. These likely represent the products of proteolytic breakdown, and were greatly reduced when proteins were expressed in the protease knockout strain SMD1163 (Fig. 2.4B). 2.2.4 Do the S. cerevisiae VBA2 and VBA3 genes produce functional proteins?
While designing expression constructs, I noticed that VBA2 and VBA3 are N-terminally truncated compared to the other members of the VBA family (Fig. 2.5). Based on sequence comparison, the S. cerevisiae VBA proteins belong to the drug:H+ antiporter-2 (DHA2) family of the Major Facilitator Superfamily (MFS) [99,161]. Members of the DHA2 family have a 14-transmembrane segment (TMS) topology, which matches the TOPCONS [162] predictions for Vba1, Vba4
B
KM71H
*
A
15 s exposure 8 s exposure
Vba5
Vba1
Lyp1
Can1
KM71H
Vba5
Vba1
Lyp1
Can1
KM71H
SMD1163
Vba1kDakDa * * * *
Figure 2.4 Crude membrane fractions from small-scale expression cultures were separated by SDS-PAGE and imaged using in-gel fluorescence. (A) Can1, Lyp1, Vba1 and Vba5 expressed in KM71H. The left and right images were taken using longer and shorter exposure times, respectively. (B) Vba1 expressed in either KM71H or SMD1163. Stars indicate full-length protein.

26
Expressing S. cerevisiae transporters in P. pastoris
and Vba5. Vba2 and Vba3 are predicted to only contain 12 and 11 TMS, respectively. This means that they lack TM1, 2, and/or 3 relative to the other VBAs and related MFS proteins. This is surprising given that in many other MFS proteins TM1 is implicated in gating, proton-coupling, and substrate binding [98].
After our Vba2 and Vba3 constructs failed to produce membrane-localized protein in P. pastoris, I considered the possibility that VBA2 and VBA3, as annotated in the S. cerevisiae S228c genome, do not produce functional proteins. Vba1, Vba2 and Vba3 were all reported to transport basic amino acids over the vacuolar membrane [57]. This conclusion was largely based on a series of individual or combined gene deletions, which were shown to reduce ATP-dependent lysine uptake by vacuolar membrane vesicles. Only for Vba1 was the activity and localization of the gene product confirmed by gene complementation and fluorescence imaging of a GFP-tagged variant [57]. It is possible that the VBA2 and VBA3 gene deletions cause indirect effects, and that the observed phenotypes were not due to an absence of the predicted Vba2 and Vba3 proteins.
Vba3 is a paralog of Vba5 which arose from a genome duplication event [163]. Analysis of the S288c genome sequence shows that Vba3 is shorter than Vba5 only because of a single base pair change (TTA to TGA, leucine -82 to STOP). Without this, the VBA3 ORF would extend another 372 base pairs upstream. The translated product (Vba3ext) is 99% identical to Vba5 and is also predicted to contain 14 TMS (Fig. 2.5). Similarly, without a single TGA stop codon the VBA2 gene could be extended by 261 base pairs upstream to produce a 14 TM protein (Vba2ext). The NCBI Genome database (http://www.ncbi.nlm.nih.gov/genome/, accessed September 2015) contains 84 S. cerevisiae strains that code for Vba3ext, and 86 that code for Vba2ext. Three Vba2 homologs from Schizosaccharomyces pombe, which localize to the vacuolar membrane and are involved in amino acid transport, are N-terminally extended and predicted to have 14 TMS [164-166].
To test these theories, the annotated and extended ORFs were expressed in S. cerevisiae with C-terminal YPet fusions (Fig. 2.6). The “short” versions resulted in intracellular aggregates and no clear membrane localization. Vba2ext-YPet localized exclusively to the vacuolar membrane, while Vba3ext-YPet was observed at the periphery of the cell and internal membranes.2.2.5 Fluorescence based screening for high-expressing transformants
The original screening procedure recommended by Invitrogen essentially involves taking a random selection of 6–10 transformant strains directly to small-scale expression cultures. I used the GFP signal from our expression strains
Figure 2.5 Multiple sequence alignment of Vba1, Vba2, Vba3, Vba4 and Vba5. The first 200 residues of Vba4 are excluded. Grey shading indicates transmembrane segments as predicted by TOPCONS [162]. The translated upstream regions of Vba2 and Vba3 from S. cerevisiae S288C (isogenic to BY4742 for these genes) is included and indicated by black outlines. The upstream stop codons, which were replaced by a Leu codon to generate Vba2- and Vba3ext, are shown in red.

27
Chapter 2
Vba4
Vba3
Vba5
Vba1
Vba2
201 1 1 1 1
280
68 68 67 85
--
--
--
--
-E
VP
VS
GD
EI
TS
YG
YG
SI
PQ
SI
GD
VE
NG
LN
PP
YV
EN
TS
SD
EL
--
--
-V
HD
LT
RR
RI
F-
--
SS
CM
CT
YL
FF
IA
MD
SS
II
LV
IA
SK
IA
SE
FF
SS
CM
CT
YL
FF
IA
MD
SS
II
LV
--
--
--
--
--
--
-ME
--
--
ET
KY
SS
Q-
--
--
--
QE
IE
EA
C-
--
--
GS
DA
SL
NA
RG
SN
DS
PM
GL
SL
YL
CL
AS
LT
LV
LF
IT
AL
DI
LI
VG
TI
ID
VV
AE
QF
LY
LC
LA
SL
TL
VL
FI
TA
LD
IL
IL
--
--
--
--
--
--
-ME
--
--
ET
KY
SS
Q-
--
--
--
QE
IE
GA
C-
--
--
GS
DA
SL
NA
RG
SN
DS
PM
GL
SL
YL
CL
AS
LT
LV
LF
IT
AL
DI
LI
VG
TI
ID
VV
AE
QF
LY
LC
LA
SL
TL
VL
FI
TA
LD
IL
IM
QT
LD
ET
SN
LL
PP
P-
--
--
--
--
--
--
--
--
--
EE
AE
AP
PL
E-
--
--
-Q
K-
FH
EY
NL
AL
PK
FP
IL
--
--
FS
LW
LG
SF
LS
SL
DS
TI
VA
NIM
NR
VA
EE
FI
LF
SL
WL
GS
FL
SS
LD
ST
IV
AN
ML
-K
SS
KH
KV
LP
KS
K-
--
-D
GN
YG
SI
--
--
--
-E
ET
EI
PS
VD
YS
ED
LD
ED
LD
KG
ED
IA
LS
RI
PN
LW
II
EA
TL
FS
NV
FL
SG
FD
GT
VT
AS
TY
QT
IG
NE
FW
II
EA
TL
FS
NV
FL
SG
FD
GT
VL
Vba4
Vba3
Vba5
Vba1
Vba2
281 69 69 68 86
377
165
165
164
182
HE
LW
RL
SL
VI
SA
YL
LS
NA
IG
QL
VF
LK
LS
LI
SS
VK
LL
LC
IA
QF
SF
IL
GG
YL
SW
SS
AH
FW
TF
IF
AR
CV
TG
FG
GG
SL
IA
LK
ST
IMN
RF
SQ
KN
DS
RY
SL
SA
RL
SL
VI
SA
YL
LS
NA
IG
QL
VF
LL
LL
CI
AQ
FS
FI
LG
GY
LS
WS
SA
IF
AR
CV
TG
FG
GG
SL
IA
LK
ST
IL
SA
GN
YS
KT
GW
LV
TG
YS
LP
NA
IL
SL
IWG
RF
AS
II
GF
QH
SL
IL
AI
LI
FE
AG
SL
IA
AL
AS
SM
NM
LI
VG
RV
VA
SV
GG
SG
LQ
TL
CF
VI
GC
TM
VG
ER
SR
PL
VI
SI
KT
GW
LV
TG
YS
LP
NA
IL
SL
IWG
HS
LI
LA
IL
IF
EA
GS
LI
AA
LA
SV
GR
VV
AS
VG
GS
GL
QT
LC
FV
IG
IS
IG
NY
SK
TG
WL
VT
GY
SL
PN
AI
LS
LIW
GR
FA
SI
IG
FQ
HS
LI
LA
IL
IF
EA
GS
LI
AA
LA
SS
MN
ML
IF
GR
VV
AG
VG
GS
GL
QT
LC
FV
IG
CT
MV
GE
RS
RP
LV
IS
IK
TG
WL
VT
GY
SL
PN
AI
LS
LIW
GH
SL
IL
AI
LI
FE
AG
SL
IA
AL
AS
FG
RV
VA
GV
GG
SG
LQ
TL
CF
VI
GI
SI
SE
SS
KK
QW
IA
TS
FL
LT
NT
AF
QP
LY
GK
LS
DI
TG
RK
SA
LL
TA
QF
FF
GL
GC
LL
TC
FA
RN
VT
EF
SI
AR
AI
CG
IG
AG
GL
NA
IS
SI
AV
SD
IC
TA
RE
RG
VY
QG
YK
KQ
WI
AT
SF
LL
TN
TA
FQ
PL
YG
AL
LT
AQ
FF
FG
LG
CL
LT
CF
AR
NF
SI
AR
AI
CG
IG
AG
GL
NA
IS
SI
GY
NQ
MS
IS
NW
IT
TA
YL
IT
ST
SF
QP
LY
GS
FS
DA
LG
RR
NC
LF
FA
NG
AF
TI
GC
LA
CG
FS
KN
IY
ML
SF
MR
AL
TG
IG
GG
GL
IT
LS
TI
VN
SD
VI
PS
SK
RG
IF
QA
FI
SN
WI
TT
AY
LI
TS
TS
FQ
PL
YG
CL
FF
AN
GA
FT
IG
CL
AC
GF
SK
NL
SF
MR
AL
TG
IG
GG
GL
IT
LS
TI
AF
Vba4
Vba3
Vba5
Vba1
Vba2
378
166
166
165
183
456
259
259
237
257
SM
IT
FA
MG
VV
IG
PF
MM
NL
FD
SS
HG
SG
WR
NA
FL
IP
VP
FC
LV
NA
SIM
LA
DM
YS
VK
--
--
--
--
ST
LY
GR
PT
PT
LW
KR
FK
N-
--
--
--
--
-T
LL
SP
DL
YE
SM
IT
FA
MG
VV
IG
PF
MM
NL
RN
AF
LI
PV
PF
CL
VN
AS
IML
AD
PD
LY
EL
SC
AF
AV
AA
IV
GP
II
GG
AF
TT
H-
-V
TW
RW
CF
YI
NL
PI
GG
LA
IIM
FL
LT
-Y
KA
EN
KG
IL
IK
DA
IG
TI
SS
FT
FS
KF
RH
QV
NF
KR
LM
NG
II
FK
FD
FF
GL
SC
AF
AV
AA
IV
GP
II
GG
AW
CF
YI
NL
PI
GG
LA
IIM
FL
LT
YD
FF
GL
SC
AF
AV
AA
IV
GP
II
GG
AF
TT
H-
-V
TW
RW
CF
YI
NL
PI
GG
LA
IIM
FL
LT
-Y
KA
EN
KG
IL
IK
DA
IG
TI
SS
FT
FS
KF
RH
QV
NF
KR
LM
NG
II
FK
FD
FF
GL
SC
AF
AV
AA
IV
GP
II
GG
AW
CF
YI
NL
PI
GG
LA
IIM
FL
LT
YD
FF
GA
NI
VF
GF
GQ
LL
GA
PL
GG
VF
IE
T-
-I
GW
RA
LF
GI
QV
PV
IML
CS
VL
AI
KN
-I
NI
KL
FH
V-
PP
MK
ER
YT
L-
--
--
--
--
--
--
--
--
--
-K
NL
SR
ID
IF
GA
NI
VF
GF
GQ
LL
GA
PL
GG
VF
AL
FG
IQ
VP
VIM
LC
SV
LA
IK
NI
ID
IF
GQ
NL
LL
GF
GA
IC
GA
SF
GG
TI
AS
S-
-I
GW
RW
CF
LI
QV
PI
SV
IS
SI
LM
NY
--
--
--
--
YV
-P
N-
QK
EY
NR
--
--
--
--
--
QN
SS
IF
QN
PG
KI
LR
DI
DV
MG
QN
LL
LG
FG
AI
CG
AS
FG
GT
IR
WC
FL
IQ
VP
IS
VI
SS
IL
MN
YY
ID
VM
G
Vba4
Vba3
Vba5
Vba1
Vba2
457
260
260
238
258
538
341
341
299
330
I-
--
--
-L
TL
TL
FL
L-
--
CF
VQ
VT
SL
DL
TG
LK
NN
TM
IQ
AL
LF
SV
II
VC
GI
LF
--
--
--
FL
IE
TS
DT
YM
NS
VI
SM
SL
QG
DK
RL
IWT
MI
GI
SF
CF
AA
LM
IL
TL
TL
FL
LC
FV
QV
TS
MI
QA
LL
FS
VI
IV
CG
IL
FF
LI
ET
MI
GI
SF
CF
AA
LM
FA
LC
SA
GL
--
VL
FL
LG
LT
FG
GN
KY
SW
--
--
--
--
NS
GQ
VI
AY
LV
LG
VL
LF
IF
SL
VY
DF
FL
FD
KF
NP
EP
DN
--
--
-I
SY
RP
LL
LR
RL
VA
KP
AI
II
IN
MF
AL
CS
AG
LV
LF
LL
GL
TF
GQ
VI
AY
LV
LG
VL
LF
IF
SL
VY
DI
II
NM
FA
LC
SA
GL
--
VL
FL
LG
LT
FG
GN
KY
SW
--
--
--
--
NS
GQ
VI
TY
LV
LG
VL
LF
IF
SL
VY
DF
FL
FD
KF
NP
EP
DN
--
--
-I
SY
RP
LL
LR
RL
VA
KP
AI
II
VN
MF
AL
CS
AG
LV
LF
LL
GL
TF
GQ
VI
TY
LV
LG
VL
LF
IF
SL
VY
DI
IV
NM
SL
SL
VA
TI
SG
VL
F-
--
-L
CS
SQ
L-
--
--
--
--
--
NK
LY
LA
LF
TI
G-
--
SF
IV
F-
--
--
IL
VE
RY
YA
TE
--
--
--
--
--
-K
IL
PF
EL
LT
RS
FC
LS
-S
AS
LS
LV
AT
IS
GV
LF
LC
SL
YL
AL
FT
IG
SF
IV
FI
LV
ER
YY
LS
SA
SI
LI
IT
GL
TL
QL
LY
LS
LG
CS
TS
KL
SW
--
--
--
--
TS
PS
VL
LL
LV
GS
VI
IL
LL
F-
--
--
IL
HE
RK
TS
AR
--
--
--
--
--
-A
II
PM
EL
VN
SS
YS
VV
VL
SS
IL
II
TG
LT
LQ
LL
YL
SS
PS
VL
LL
LV
GS
VI
IL
LL
FI
LH
VV
LS
Vba4
Vba3
Vba5
Vba1
Vba2
539
342
342
300
331
612
429
429
396
416
--
--
-C
II
PF
--
--
-G
TT
YF
II
VL
NL
ST
LQ
LA
ER
LS
PF
FF
SI
VL
GY
FS
VS
YF
WK
SK
GQ
--
-N
FL
LK
FV
LS
GA
TL
LL
--
--
--
YV
AL
MG
VS
LN
L-
--
-C
II
PF
GT
TE
RL
SP
FF
FS
IV
FL
LK
FV
LS
GA
TL
LL
YV
AL
MG
VV
TF
LL
CT
GY
NG
QM
IY
SV
QF
FQ
LI
FA
SS
AW
KA
GL
HL
IP
IV
IT
NV
IA
AI
AS
GV
IT
KK
L-
--
-G
LV
KP
LL
IF
GG
V-
LG
VI
GA
GL
MT
LM
TN
-T
ST
KS
T-
--
VT
FL
LC
TG
YN
GQ
MI
YS
GL
HL
IP
IV
IT
NV
IA
AI
AS
GV
IK
PL
LI
FG
GV
LG
VI
GA
GL
MT
LM
VT
FL
LC
TG
YN
GQ
MI
YS
VQ
FF
QL
IF
AS
SA
WK
AG
LH
LI
PI
VI
TN
VI
AA
IA
SG
VI
TK
KL
--
--
GL
VK
PL
LI
FG
GV
-L
GV
IG
AG
LM
TL
MT
N-
TS
TK
ST
--
-V
TF
LL
CT
GY
NG
QM
IY
SG
LH
LI
PI
VI
TN
VI
AA
IA
SG
VI
KP
LL
IF
GG
VL
GV
IG
AG
LM
TL
MV
TV
IS
SF
VV
FG
EI
FR
SP
IY
LQ
LL
QN
IS
VT
KT
GL
FL
IF
PS
IS
VA
VG
SL
VT
GW
VL
RN
TK
IN
LA
HC
AY
QI
IF
GG
MIM
QL
LG
LG
LG
YF
LL
SH
LN
PD
YT
IY
DV
TV
IS
SF
VV
FG
EI
FR
SP
LF
LI
FP
SI
SV
AV
GS
LV
TG
WV
LA
YQ
II
FG
GM
IMQ
LL
GL
GL
GY
FI
SI
LV
GF
AS
YA
YL
FT
LP
LF
FQ
IV
LG
DS
TA
KA
GL
RL
TI
PS
LF
TP
VG
SL
IT
GF
SM
SK
Y-
--
--
NC
LR
LL
LY
IG
IS
LM
FL
GN
FL
-F
LF
IE
KT
SP
N-
--
--
IS
IL
VG
FA
SY
AY
LF
TL
PL
RL
TI
PS
LF
TP
VG
SL
IT
GF
SM
RL
LL
YI
GI
SL
MF
LG
NF
LF
LF
I
Vba4
Vba3
Vba5
Vba1
Vba2
613
430
430
397
417
689
509
509
487
492
--
--
--
--
--
PV
WK
QY
IC
LS
--
-L
PF
LG
--
SS
MI
LT
LL
SN
LY
HE
YH
EQ
RK
SP
--
--
IS
GS
IV
YC
FG
AV
GG
TV
GI
SL
GG
YV
FH
KT
LI
KL
MH
EK
VM
-P
FY
IC
LS
LP
FL
GS
SM
IL
TL
LS
NL
IV
YC
FG
AV
GG
TV
GI
SL
GG
YV
F-
--
--
--
--
--
--
QI
GV
LL
L-
--
-P
GF
SL
GF
AL
QA
SL
MS
AQ
LQ
IT
KD
RP
EA
AM
DF
IE
VT
AF
NT
FM
KS
LG
TT
LG
GV
LS
TT
VF
SA
SF
HN
KV
SR
AH
LE
PY
QI
GV
LL
LP
GF
SL
GF
AL
QA
SL
MF
NT
FM
KS
LG
TT
LG
GV
LS
TT
VF
--
--
--
--
--
--
-Q
IG
VL
LL
--
--
PG
FS
LG
FA
LQ
AS
LM
SA
QL
QI
TK
DR
PE
AA
MD
FI
EV
TA
FN
TF
MK
SL
GT
TL
GG
VL
ST
TV
FS
AS
FH
NK
VS
RA
HL
EP
YQ
IG
VL
LL
PG
FS
LG
FA
LQ
AS
LM
FN
TF
MK
SL
GT
TL
GG
VL
ST
TV
FM
LE
SI
TF
RS
NS
IWW
KL
IY
VF
AS
VL
VS
FG
YA
CL
LV
AT
LV
SI
VF
TV
EK
SQ
QG
T-
--
--
-MT
GV
FY
LW
RS
IG
NV
LG
AS
LT
LV
SY
EN
SL
SS
ML
WN
YM
FK
TK
SV
LV
SF
GY
AC
LL
VA
TL
VS
IV
FY
LW
RS
IG
NV
LG
AS
LT
LV
SY
EN
--
--
--
--
--
--W
LI
GL
FL
I-
--
PA
NL
GQ
GI
TF
PT
TL
FT
FI
FM
FS
KS
DQ
AT
--
--
--
AT
ST
LY
LF
RS
IG
SV
WG
VA
IS
AG
VI
QL
SF
AG
LL
RS
NL
KG
LL
IG
LF
LI
PA
NL
GQ
GI
TF
PT
TL
FY
LF
RS
IG
SV
WG
VA
IS
AG
VI
QL
Vba4
Vba3
Vba5
Vba1
Vba2
690
510
510
488
493
768
582
582
562
561
SK
QG
--
YL
KK
DL
LK
II
KH
AT
ES
SD
WV
HE
SA
PK
FV
-F
QT
LI
EC
YL
QA
CR
NV
FK
LS
TL
--
--
FF
TI
TV
V-
AI
FI
FN
R-
-I
HC
RS
QN
CL
--
--
SL
S-
-R
NV
FK
LS
TL
FF
TI
TV
VA
IF
IF
EG
KT
--
VD
DM
IL
YR
LQ
N-
--
--
--
YD
--
-G
SH
ST
IG
NI
--
--
LS
DS
IK
NV
FW
MD
--
--
LG
FY
AL
GF
L-
-F
CS
FS
SN
KK
LI
IP
KK
DE
TP
ED
NL
ED
KK
NV
FW
MD
LG
FY
AL
GF
LF
CS
FS
EG
KT
--
VD
DM
IL
YR
LQ
N-
--
--
--
YD
--
-G
SH
ST
IG
NI
--
--
LS
DS
IK
NV
FW
MD
--
--
LG
FY
AL
GF
L-
-F
CS
FS
SN
KK
LI
IP
KK
DD
TP
ED
NL
ED
KK
NV
FW
MD
LG
FY
AL
GF
LF
CS
FS
RD
DE
YH
FT
KK
QY
YS
LI
N-
--
-D
SS
YL
R-
-G
PN
FP
--
TD
IF
VR
IL
DV
YK
KA
FL
IS
YI
PN
IA
LA
AV
GI
VL
SL
YL
VK
H-
-T
YK
RS
SS
S-
--
--
--
--
-F
LI
SY
IP
NI
AL
AA
VG
IV
LS
LY
DE
NK
--
-I
KK
LI
VQ
LS
A-
--
-N
SS
YI
G-
-S
LH
GE
VK
NT
VI
KS
FD
EA
TK
RA
HL
MS
TL
--
--
LS
SL
AL
I-
-L
CI
LK
D-
-N
LA
KP
KT
RR
--
--
--
--
-T
KR
AH
LM
ST
LL
SS
LA
LI
LC
IL

28
Expressing S. cerevisiae transporters in P. pastoris
to introduce a high-throughput procedure for screening transformants before advancing to expression cultures. Although this procedure requires approximately a week to complete, it increased the number of transformant strains we routinely screened by 10-fold.
In this protocol, colonies are grown on nitrocellulose membranes placed directly onto agar plates. The membranes are first placed on minimal glycerol medium, and inoculated from YPD precultures using a 96-pin replicator tool. After 2–3 days growth, when the colonies are at a reasonable size, the membrane is transferred to minimal methanol medium. After 24 h of induction, the membrane can be transferred to a black plate and a fluorescence image taken. My adaptation was to use colony fluorescence imaging; in the original protocol the colonies were lysed onto the nitrocellulose membrane and analyzed via immunoblotting. A similar protocol has since been published, with the authors validating the correlation between expression levels in plate and liquid cultures [167].
Figure 2.7 shows the results of using this method to screen SMD1163/Lyp1 strains. Colonies with the strongest fluorescent signal (Fig. 2.7A) were further
Vba2 Vba2ext
Vba1
Vba3 Vba3ext
Figure 2.6 Localization of Vba1, Vba2, Vba2ext, Vba3 and Vba3ext fused to YPet and expressed in S. cerevisiae BY4742. Panels on the left show the signal from YPet (green) and from the vacuolar membrane (red, stained with FM4-64). Panels on the right show a bright-field image of the same cells. Scale bars are 2 µm.

29
Chapter 2
screened using small-scale expression tests (Fig. 2.7B). Strain M10 was used for the purification of Lyp1 described in Section 2.2.7.
2.2.6 Testing alternative carbon sourcesSorbitol, mannitol and alanine can be used as sole carbon sources by P. pastoris
and do not repress the AOX1 promoter [168]. It has been reported that co-feeding P. pastoris methanol induction cultures with sorbitol allows faster growth while maintaining, or even increasing, recombinant protein production [169,170]. I tested the use of these alternative C-sources in small-scale expression cultures of KM71H/Vba5 (Fig. 2.8). The strains were precultured in minimal glycerol medium (MGY), and then transferred into minimal methanol medium (MM) with or without the addition of 1% (w/v) sorbitol, mannitol, or alanine. Cells were cultured for 72 hours, with 0.5% (v/v) methanol added at 24 h intervals.
100 500 1000 Zeocin (μg/mL)
SMD1163
*
* *
*
*
*
A
B
70
25
kDa M2 M10 M12 M20 M28 M29
SMD1163
Lyp1
Figure 2.7 Screening SMD1163/Lyp1 expression strains. (A) Fluorescence image of methanol-induced colonies grown on a nitrocellulose membrane. Zeocin concentrations are those used in selective transformation plates. Stars indicate strains used in further testing. (B) Crude membrane fractions from small-scale expressions in liquid culture were separated by SDS-PAGE and imaged by in-gel fluorescence.

30
Expressing S. cerevisiae transporters in P. pastoris
All the additional carbon sources increased growth relative to the methanol-only control (Fig. 2.8A). The cell density of alanine cultures decreased after 24 h, likely because they used all the available carbon source. Growth of the methanol-only culture plateaued after 24 hours and the amount of Vba5 in the membrane fraction decreased between 48 and 72 h (Fig. 2.8A,B). This is likely because the carbon source became limiting. Sorbitol and mannitol gave similar results to each other, with expression levels comparable to the methanol-only culture and faster growth rates. The culture containing alanine produced less Vba5 but grew much faster. Fluorescence images (taken at 24 h) show that the addition of sorbitol, mannitol or alanine had no discernable effect on the localization of Vba5 (Fig. 2.8C).
In conclusion, the use of an additional carbon source increased the cell density without significantly decreasing the expression of Vba5. In the future, expressions
0 24 48 720
2
4
6
8
10
OD
600
Time (h)
24 h 72 h48 hMSMe A Me AMS Me AMS
70
25
kDa
*
methanol
alanine
mannitol
sorbitol
A
B
C
methanol
sorbitol
alaninemannitol
Figure 2.8 The effect of additional carbon sources on the methanol-induced expression of Vba5 in P. pastoris KM71H. (A) Cell density over time as measured by OD600. At T = 0 h, cultures were diluted from minimal media with glycerol to minimal media with methanol (control) with or without the addition of 1% (w/v) sorbitol, S; mannitol, M; or alanine, A. (B) SDS-PAGE gel of crude membrane fractions, imaged by in-gel fluorescence. (C) Confocal fluorescence microscopy images. Scale bars are 2 µm.

31
Chapter 2
should be tested with either sorbitol or mannitol to see if they improve the yield of recombinant protein. Alanine could also be considered if a non-fermentative carbon source might be useful. It should be noted that the two P. pastoris strains used in this study grow differently on methanol as a sole carbon source. KM71H does not have an intact AOX1 gene, and thus relies on the less efficient AOX2. This results in a slow growth phenotype commonly referred to as MutS, or methanol utilization slow. SMD1163, on the other hand, does have an intact AOX1 gene and is thus wildtype for growth on methanol (Mut+). Variations in substrate carbon consumption between MutS and Mut+ strains have been observed previously [169].2.2.7 Purification of Can1 and Lyp1
Can1 and Lyp1 were both purified from SMD1163 expression strains using sequential Ni-affinity and size-exclusion chromatography (SEC). Lyp1 expression and purification was optimized by Frans Bianchi and Joury van ‘t Klooster, who were able the reconstitute the purified protein into lipid vesicles and perform in vitro characterization (Fig. 2.9D,E) [105]. The results of the single Can1 purification were promising but require optimization due to a low yield (Fig. 2.9). A large proportion of the protein was either not solubilized from the membrane or did not bind the Ni-affinity column, suggesting that the conditions used are not optimal for stability of the protein. This is supported by the appearance of a higher-molecular weight (HMW) fluorescent species in the elution fractions from the Ni-affinity column. This species, which likely represents some sort of aggregation, was efficiently removed by SEC. The elution fractions from this column, however, were enriched in a non-fluorescent HMW species which likely represents Can1 with an unfolded GFP tag [160]. The final samples contained a 1:1 ratio of fluorescent and non-fluorescent Can1, which is undesirable for downstream applications.
2.3 Materials and methods2.3.1 Strains and culture conditions
All amino acid transporters were amplified from Saccharomyces cerevisiae BY4742 (MATa his3Δ1 leu2Δ0 lys2Δ0 ura3Δ0) [171]. Escherichia coli strain MC1061 was used for cloning and plasmid storage. Pichia pastoris strains used were KM71H (arg4 aox1::ARG4, MutS) and SMD1163 (his4 pep4 prb1, Mut+), which is lacking two vacuolar proteases.
All yeast cultures were grown under aerobic conditions at 30°C, with shaking (180–200 rpm) for liquid cultures. For plate cultures, agar was added to 15 g/L for E. coli or 20 g/L for yeast.
E. coli were grown in LB (1% tryptone, 0.5% yeast extract, 1% NaCl) with 100 µg/mL ampicillin or 50 µg/mL zeocin as required. When zeocin was used, the LB was modified by lowering the salt concentration (0.5% NaCl) and adjusting the pH to 7.5 with HCl/NaOH before autoclaving. The following media were used to culture P. pastoris strains. YPD: 1% yeast extract, 2% peptone, 2% D-glucose.

32
Expressing S. cerevisiae transporters in P. pastoris
mAU
Elution volume (mL)
A B DC f c
A B C D A B C DkDa
*
Bcf
kDa
S PM FWash1 2 53 41 2
ElutionS PM F
Wash1 2 53 41 2
Elution
*
A
kDapurifiedprotein
proteo-liposomes
f fc c
mAu
Elution volume (mL)
D
C
E
Figure 2.9 Purification of Can1 (A,B,C) and Lyp1 (D,E) from P. pastoris SMD1163. (A) Samples from membrane solubilization and Ni-affinity purification of Can1 separated by SDS-PAGE. M, solubilization mixture; S, soluble fraction; P, pellet; F, flowthrough from Ni-affinity column. (B) Elution fraction 2 from (A) was further purified by size-exclusion chromatography (SEC). (C) Elution fractions from (B) were separated by SDS-PAGE. (D) SEC purification of Lyp1 following Ni-affinity column. (E) Samples of purified Lyp1 and proteoliposomes separated by SDS-PAGE. All gels were imaged by both in-gel fluorescence (f ) and Coomassie staining (c). In chromatograms, lines represent absorbance at 280 nm (solid), 260 nm (dotted) and 488 nm (dashed).

33
Chapter 2
YPDS: YPD with 1 M sorbitol. Minimal media: 1.34% YNB (Yeast Nitrogen Base with ammonium sulfate and without amino acids, Formedium, UK), 0.00004% biotin with either 1% glycerol (MGY) or 0.5–1% methanol (MM) as carbon source. For SMD1163, 0.004% L-histidine was added to minimal media.
S. cerevisiae BY4742 strains were grown in YPD or in synthetic complete media containing 0.67% (w/v) YNB, 1.926 g/L Kaiser amino acid drop-out supplement minus uracil (Formedium, UK), and 2% (w/v) D-glucose or raffinose. For expression of VBA proteins from the GAL1 promoter, BY4742 strains were grown to mid-log phase (OD600 0.2–0.6) in raffinose medium and induced by the addition of 0.2% (w/v) galactose. Fluorescence imaging was performed after 2 h of induction.2.3.2 Plasmid and strain construction
Plasmids and oligonucleotide primers used in this study are listed in Tables 2.1 and 2.2, respectively. Isolation of S. cerevisiae BY4742 chromosomal DNA was carried out according to Sherman et al. [172]. All plasmids were generated using uracil excision-based cloning [173]. The amplification of DNA using uracil-containing primers was performed using the polymerase PfuX7 [174]. Crude PCR products were treated with USER enzyme (New England Biolabs, USA) as per the manufacturer’s instructions and transformed into E. coli MC1061 for in vivo assembly. All plasmids were isolated from the E. coli host and verified by DNA sequencing of the fusion genes.
Can1, Gap1, Lyp1, Vba1, Vba2, Vba3 and Vba4 were previously cloned into pBADcLIC-GFP [160] to test for expression in E. coli (Frans Bianchi, unpublished result). These plasmids contain the gene of interest followed by sequences coding for the tobacco etch virus (TEV) protease cleavage site, green fluorescent protein (GFP) and a ten residue His-tag. For expression in P. pastoris, the entire open reading frames from these plasmids were sub-cloned into pPICZ A. For Vba5, separate fragments were PCR amplified coding for the gene of interest, using S. cerevisiae BY4742 genomic DNA as template, and the GFP-tag from pBADcLIC-GFP. Vba2ext and Vba3ext were cloned in a similar fashion, except that overlap extension PCR was first used to create template DNA fragments identical to the S. cerevisiae BY4742 genome except for changing the upstream STOP codons to Leu (TGA → TTA). In all cases, primers were designed to change the His10-tag for Arg-Gly-Ser-His10, as well as to introduce a yeast Kozak consensus sequence ((G/A)NNATGG where ATG is the start codon). To maintain this sequence, glutamate (GAG) was introduced as the second residue for Can1, Gap1, Vba1, Vba2 and Vba3. For the full sequence of the C-terminal tag on expressed proteins, see the Supplementary (Section 2.4).
For homologous integration into the chromosome of P. pastoris, the plasmid was linearized by restriction digest with PmeI (New England Biolabs, USA) and transformed into competent P. pastoris cells using electroporation. DNA preparation and transformation was as described in the EasySelect Manual (Invitrogen), except

34
Expressing S. cerevisiae transporters in P. pastoris
for the following changes. After electroporation, cells were immediately transferred to a culture tube using a Pasteur pipette and then incubated at 30°C without shaking. After 1 h, 1 mL of YPD was added to each tube and the cells incubated at 30°C with shaking. After a further 2–3 h, 100 µL was plated on YPDS agar with 100 µg/mL zeocin. The remaining cell culture was centrifuged briefly and the cell pellet resuspended in 100 µL of YPD. Approximately one third of this cell suspension was plated on YPDS with 500 µg/mL zeocin and the remainder plated on YPDS with 1000 µg/mL zeocin.
To examine localization in S. cerevisiae, the annotated and extended VBA proteins were expressed from a galactose-inducible promoter with a C-terminal YPet tag. Plasmids pFB011, pFB012, pFB013, pFB014 and pFB015 were constructed by a four-way PCR fragment ligation. The backbone of the pDDGFP-2 vector was amplified with primer pairs 4158/3630 and 3631/4171. The resulting two fragments were combined with a third and fourth fragment, coding for YPet and the gene of interest, respectively. The fragment coding for the YPet gene was amplified from a synthetically generated coding sequence ordered from GeneArt (Regensburg, Germany), using primer pair 4159/4160. The gene of interest was amplified from either S. cerevisiae BY4742 chromosomal DNA (VBA1, VBA2, and VBA3), pLN027 (VBA2ext), or pSR030 (VBA3ext), using the primers given in Table 2.2. Ligation of the four PCR amplified fragments using USER enzyme resulted in a C-terminal YPet fusion, separated by a sequence coding for the tobacco etch virus (TEV) protease cleavage site and followed by an eight-residue His tag (e.g. Vba1-TEV-YPet-His8). For the full sequence of the C-terminal tag, see the Supplementary (Section 2.4).2.3.3 Fluorescence imaging
Fluorescence live cell imaging was performed on a LSM 710 commercial laser scanning confocal microscope (Zeiss, Germany), equipped with a C-Apochromat 40x/1.2 NA objective, a blue argon ion laser (488 nm) and a red He-Ne laser (633 nm). Cells were immobilized between a glass slide and coverslip. Images were obtained with the focal plane positioned at the mid-section of the cells.2.3.4 Small-scale expression testing
For small-scale expression testing, 10 mL cultures were grown in 50 mL CELLreactorTM filter top tubes (Greiner Bio-One). Strains were inoculated into MGY cultures from agar plates and grown overnight to an OD600 of 2–6. The tube was then centrifuged (3,000 x g for 10 min at room temperature), the supernatant removed, and the pellet resuspended in 10 mL of mM medium with 1% methanol. After a further 16–24 h of incubation, the expression was checked by fluorescence confocal microscopy and a crude membrane extraction. 2.3.5 Rapid membrane preparation
Preparation of crude membranes was based on protocols communicated by Ina Urbatsch [177,178]. Cells from 10 mL of expression culture were washed with

35
Chapter 2
500 µL of 0.33 M sucrose, 0.3 M Tris.HCl (pH 7.4) and resuspended in 500 µL of ice-cold lysis buffer (0.33 M sucrose, 0.3 M Tris.HCl pH 7.4, 1 mM ethylene glycol-bis(2-aminoethylether)-N,N,N′,N′-tetraacetic acid (EGTA), 1 mM ethylenediaminetetraacetic acid (EDTA), 100 mM ε-aminocaproic acid (EACA), 1 mM phenylmethylsulfonyl fluoride (PMSF), 10 µg/mL leupeptin, 1 µg/mL pepstatin A, 6 µg/mL chymostatin, 2.5 µg/mL E64, 5 mM dithiothreitol (DTT)). Samples were kept on ice from this point forwards. Samples were transferred to
Table 2.1 Plasmids used in this study.
Name Description1 SourcepPICZ A Expression vector designed for chromosomal integration in
P. pastoris; AOX1, ZeoRInvitrogen
pBGP1 P. pastoris/E. coli shuttle vector with GAP, AmpR, ZeoR [157]pBGP1na pGBP1 with the a-factor signal sequence removed This studypBADcLIC-GFP E. coli expression vector, araBAD, allows C-terminal
TEV-GFP-His10 tag[175]
pDDGFP-2 aka pRS426GAL1-GFP; URA3, 2µ, allows C-terminal TEV-GFP-His8 tag
[176]
pSR013 pPICZ derivative with CAN1-TEV-GFP-RGSHis10 This studypSR014 pPICZ derivative with LYP1-TEV-GFP-RGSHis10 This studypSR018 pPICZ derivative with GAP1-TEV-GFP-RGSHis10 This studypLN020 pPICZ derivative with VBA1-TEV-GFP-RGSHis10 This studypLN021 pPICZ derivative with VBA2-TEV-GFP-RGSHis10 This studypLN022 pPICZ derivative with VBA3-TEV-GFP-RGSHis10 This studypLN023 pPICZ derivative with VBA4-TEV-GFP-RGSHis10 This studypLN024 pPICZ derivative with VBA5-TEV-GFP-RGSHis10 This studypLN027 pPICZ derivative with VBA2ext-TEV-GFP-RGSHis10
contains 261 bp 5’ of annotated Vba2 gene, with STOP → Leu mutation (TGA → TTA)
This study
pSR030 pPICZ derivative with VBA3ex-TEV-GFP-RGSHis10contains 372 bp 5’ of annotated Vba3 gene, with STOP → Leu mutation (TGA → TTA)
This study
pFB004 pDDGFP-2 derivative with VBA1-TEV-YPet-His8 This studypFB012 pDDGFP-2 derivative with VBA2-TEV-YPet-His8 This studypFB013 pDDGFP-2 derivative with VBA2ext-TEV-YPet-His8 This studypFB014 pDDGFP-2 derivative with VBA3-TEV-YPet-His8 This studypFB015 pDDGFP-2 derivative with VBA3ext-TEV-YPet-His8 This study
1 2m, multicopy plasmid in yeast; AmpR, selection via ampicillin resistance in E. coli; AOX1, methanol-inducible expression in P. pastoris; araBAD, arabinose-inducible expression in E. coli, CEN/ARS, single copy plasmid in yeast; GAL1, galactose-inducible expression in S. cerevisiae; GAP, P. pastoris constitutive promoter; URA3, selection via uracil prototrophy; ZeoR, selection via zeocin resistance in E. coli and P. pastoris.

36
Expressing S. cerevisiae transporters in P. pastoris
Tabl
e 2.2
Olig
onuc
leotid
e prim
ers u
sed
in th
is stu
dy.
Nam
eSe
quen
ce (5
’ to
3’)
Des
crip
tion1
3334
AC
C A
CC
AC
C A
UC
AT
C A
TC
AT
T A
AG
TT
T T
AG
CC
T T
AG
(F) a
mpl
ifica
tion
of p
PIC
Z A
for U
SER
clon
ing
3335
AG
G T
AC
CG
A U
CC
GA
G A
CG
GC
(R) a
mpl
ifica
tion
of p
PIC
Z A
for U
SER
clon
ing
3345
ATG
GT
G G
TG
GU
G A
TG
AT
G A
TG
AG
A A
CC
AC
G A
CT
AG
T T
TT
GTA
G
AG
CT
C A
TC
CAT
GC
(R) a
mpl
ifica
tion
from
pBA
DcL
IC-G
FP fo
r USE
R cl
onin
g in
to
pPIC
Z (b
inds
at 3
’ end
of G
FP, a
dds R
GSH
is-ta
g)
3828
AC
T T
CC
AA
G G
UC
AAT
TC
A G
CA
AA
G G
(F) a
mpl
ifica
tion
of G
FP fr
om p
BAD
cLIC
-GFP
for c
loni
ng in
to
pPIC
Z af
ter g
ene o
f int
eres
t
3337
ATC
GG
T A
CC
UA
A A
AT G
GA
GA
C A
AA
TT
C A
AA
AG
A A
GA
CG
C C
GA
C
AT A
G(F
) am
plifi
catio
n of
CAN
1 fo
r USE
R cl
onin
g in
to p
PIC
Z
3338
ATC
GG
T A
CC
UA
A A
AT G
GA
GA
G T
AA
TA
C T
TC
TT
C G
TA C
GA
GA
A
GA
A T
AA
TC
C(F
) am
plifi
catio
n of
GAP
1 fo
r USE
R cl
onin
g in
to p
PIC
Z
3339
ATC
GG
T A
CC
UA
A A
AT G
GG
CA
G G
TT
TA
G T
AA
CAT
AAT
AA
C G
TC
C(F
) am
plifi
catio
n of
LYP
1 fo
r USE
R cl
onin
g in
to p
PIC
Z
3340
ATC
GG
T A
CC
UA
A A
AT G
GA
GC
A A
AC
AC
T A
GA
CG
A G
AC
TT
C A
AA
T
CT
AC
(F) a
mpl
ifica
tion
of V
BA1
for U
SER
clon
ing
into
pPI
CZ
3829
ATC
GG
T A
CC
UA
A A
AT G
GA
GA
G T
AT T
TC
AA
A T
TG
GAT
CA
C C
AC
T
GC
G(F
) am
plifi
catio
n of
VBA
2 fo
r USE
R cl
onin
g in
to p
PIC
Z
4088
ATC
GG
T A
CC
UA
A A
AT G
GA
GC
T T
AA
AT
C T
AG
TA
A A
CA
CA
A A
GT
A
CT
AC
C G
AA
AA
G C
(F) a
mpl
ifica
tion
of V
BA2e
xt fo
r USE
R cl
onin
g in
to p
PIC
Z
3830
ATC
GG
T A
CC
UA
A A
AT G
GA
GA
A T
AT G
CT
CAT
TG
T C
GG
TA
G A
GT
T
GT
TG
(F) a
mpl
ifica
tion
of V
BA3
for U
SER
clon
ing
into
pPI
CZ
3831
ATC
GG
T A
CC
UA
A A
AT G
GG
GA
A G
AA
GG
A T
AG
AC
A G
CG
C(F
) am
plifi
catio
n of
VBA
4 fo
r USE
R cl
onin
g in
to p
PIC
Z
3832
ATC
GG
T A
CC
UA
A A
AT G
GA
GG
A A
AC
TA
A G
TA C
TC
TT
C G
C(F
) am
plifi
catio
n of
VBA
5 or
VBA
3ext
for U
SER
clon
ing
into
pPI
CZ
4089
AC
C T
TG
GA
A G
UA
TA
A A
TT
TT
C T
CT
TC
T T
GT
TT
T A
GG
TT
T C
GC
C
AG
AT
T G
TC
(R) a
mpl
ifica
tion
of V
BA2e
xt fo
r USE
R cl
onin
g in
to p
PIC
Z
3835
AC
C T
TG
GA
A G
UA
TA
A A
TT
TT
C C
TT
GT
C T
TC
TA
A A
TT
AT
C T
TC
TG
G
TG
T A
TC
(R) a
mpl
ifica
tion
of V
BA3e
xt fo
r USE
R cl
onin
g in
to p
PIC
Z

37
Chapter 2
4090
CT
T T
CA
AG
A A
TC
CC
A A
AT T
TA T
GG
AT
T A
TT
GA
A G
CT
AC
(F) u
sed
in o
verla
p ex
tens
ion
PCR
to g
ener
ate V
BA2e
xt (b
inds
ove
r ST
OP
codo
n, ch
ange
s to
Leu)
4091
GTA
GC
T T
CA
ATA
AT
C C
AT A
AA
TT
T G
GG
AT
T C
TT
GA
A A
G(R
) use
d in
ove
rlap
exte
nsio
n PC
R to
gen
erat
e VBA
2ext
(bin
ds o
ver
STO
P co
don,
chan
ges t
o Le
u)
4538
GC
C T
GG
CT
T C
GT
TA
A C
TC
TT
G T
AC
(F) u
sed
in o
verla
p ex
tens
ion
PCR
to g
ener
ate V
BA3e
xt (b
inds
ove
r ST
OP
codo
n, ch
ange
s to
Leu)
4539
GTA
CA
A G
AG
TTA
AC
G A
AG
CC
A G
GC
(R) u
sed
in o
verla
p ex
tens
ion
PCR
to g
ener
ate V
BA3e
xt (b
inds
ove
r ST
OP
codo
n, ch
ange
s to
Leu)
4158
AC
C A
CC
AC
C A
UC
AT
C A
TC
AT
C A
TT
AA
C T
GC
AG
G A
AT T
C(F
) am
plifi
catio
n of
pD
DG
FP-2
frag
men
t 1
3630
AG
G G
TA G
TG
CU
G A
AG
GA
A G
CA
TA
C G
AT A
CC
C(R
) am
plifi
catio
n of
pD
DG
FP-2
frag
men
t 1
3631
AG
C A
CT
AC
C C
UT
TA
G C
TG
TT
C T
AT A
TG
CT
G C
C(F
) am
plifi
catio
n of
pD
DG
FP-2
frag
men
t 2
4171
ATT
TT
G G
GA
UC
C A
CT
AG
T T
CT
AG
A A
TC
CG
G G
G(R
) am
plifi
catio
n of
pD
DG
FP-2
frag
men
t 2
4159
AG
G G
GA
AA
A U
TT
ATA
TT
T T
CA
AG
G T
TC
TA
A A
GG
TG
A A
GA
AT
T
ATT
CA
C T
GG
(F) a
mpl
ifica
tion
of Y
Pet f
or U
SER
clon
ing
into
pD
DG
FP-2
4160
AT
G G
TG
GT
G G
UG
GA
G C
TC
TT
T G
TA C
AA
TT
C A
TT
CAT
AC
C(R
) am
plifi
catio
n of
YPe
t for
USE
R cl
onin
g in
to p
DD
GFP
-2
4675
ATC
CC
A A
AA
UG
G G
AC
AA
A C
AC
TA
G A
CG
AG
A C
TT
CA
A A
TC
TA
C(F
) am
plifi
catio
n of
VBA
1 fo
r USE
R cl
onin
g in
to p
DD
GFP
-2
4676
AT
T T
TC
CC
C U
CC
AG
A A
CT
TG
A A
CT
AC
G T
TT
GTA
AG
T A
TG
TT
T C
(R) a
mpl
ifica
tion
of V
BA1
for U
SER
clon
ing
into
pD
DG
FP-2
6287
AT
C C
CA
AA
A U
GG
AG
A G
TA T
TT
CA
A A
TT
GG
A T
CA
CC
A C
TG
(F) a
mpl
ifica
tion
of V
BA2
for U
SER
clon
ing
into
pD
DG
FP-2
6289
AT
T T
TC
CC
C U
CC
TC
T T
CT
TG
T T
TT
AG
G T
TT
CG
C C
AG
AT
T G
TC
(R) a
mpl
ifica
tion
of V
BA2
for U
SER
clon
ing
into
pD
DG
FP-2
6288
AT
C C
CA
AA
A U
GG
AG
C T
TA A
AT C
TA G
TA A
AC
AC
A A
AG
TA
C T
AC
CG
(F) a
mpl
ifica
tion
of V
BA2e
xt fo
r USE
R cl
onin
g in
to p
DD
GFP
-2
6290
AT
C C
CA
AA
A U
GG
AG
A A
TA T
GC
TC
A T
TG
TC
G G
TA G
AG
(F) a
mpl
ifica
tion
of V
BA3
for U
SER
clon
ing
into
pD
DG
FP-2
6292
AT
T T
TC
CC
C U
CC
CT
T G
TC
TT
C T
AA
AT
T A
TC
TT
C T
GG
TG
T C
TC
GT
C(R
) am
plifi
catio
n of
VBA
3 fo
r USE
R cl
onin
g in
to p
DD
GFP
-2
6291
AT
C C
CA
AA
A U
GG
AG
G A
AA
CTA
AG
T A
CT
CT
T C
GC
AG
C(F
) am
plifi
catio
n of
VBA
3ext
for U
SER
clon
ing
into
pD
DG
FP-2
1 F, f
orwa
rd p
rimer
; R, r
ever
se p
rimer

38
Expressing S. cerevisiae transporters in P. pastoris
2 mL microcentrifuge tubes containing ~500 µL of 0.5 mM acid-washed glass beads. Cells were lysed using a TissueLyser (5 min at 50 Hz) in a pre-cooled tube holder. Samples were centrifuged (11700 x g, 5 min, 4°C) and 200 µL of supernatant transferred to a clean 1.6 mL microcentrifuge tube. 200 µL of lysis buffer was added to the already lysed cells and the TissueLyser, centrifugation, and supernatant removal steps repeated. The resulting 400 µL of supernatant was centrifuged at maximum speed in a benchtop centrifuge (20817 x g in the Eppendorf 5417R) for 1 h at 4°C. The resulting pellet (yellowish-brown and translucent in appearance) was resuspended in 40 µL of lysis buffer to give a crude membrane fraction. In our hands, the pellet was loose enough to be resuspended by pipetting up and down several times.
The total protein concentration was estimated by a Bradford assay in a microplate format. After adding 1/5 volume of 5X Laemmli buffer, the samples were briefly vortexed, incubated at room temperature for 15–30 min, vortexed again, and then 20 µg of total protein loaded on a 10% SDS-PAGE gel.2.3.6 Lysine transport assays
Cells from small-scale expression cultures were washed and resuspended in 100 mM potassium phosphate (pH 6.0), and kept on ice before being added to transport assay reactions. Each assay contained cells at a final OD600 of 0.5. The buffer used was 100 mM potassium phosphate (pH 6.0), 10 mM glucose. The radioactive substrate used was L-[14C(U)]-lysine (PerkinElmer, USA). All assays were performed at 30˚C with continuous mixing by magnetic stirring.
At given time intervals 50 µL or 100 µL samples were taken and quenched in 2 mL of ice-cold buffer. Samples were passed over a 0.45 µm pore size nitrocellulose filter (GE-Healthcare, UK) by filtration and washed with another 2 mL of ice-cold buffer. Filters were dissolved in 2 mL of scintillation solution (Emulsifierplus, PerkinElmer). Radioactivity was determined by liquid scintillation counting (Tri-Carb 2800TR liquid scintillation analyzer, PerkinElmer). 2.3.7 Large-scale expression
For large-scale expression of Can1 and Lyp1 from P. pastoris SMD1163, 0.5–1 L cultures were grown in 5 L baffled Erlenmeyer flasks. For Lyp1, cultures were grown to an OD600 of 0.6–0.8 in MGY with 0.004% L-histidine and then diluted to an OD600 of 0.05 in the same medium with 1% (w/v) mannitol instead of glycerol. When the cultures reached an OD600 of 1–2, expression of Lyp1 was induced by adding methanol to a final concentration of 0.5% (v/v). Cultures were incubated for a further 24 h, after which they were cooled to 4°C, harvested by centrifugation (7500 x g, 15 min), washed once with 50 mM Tris-HCL (pH 6.7), 1 mM EDTA, 0.6 M sorbitol, and suspended to a final OD600 of ~100. For Can1, 0.5 L cultures were grown to an OD600 of 5.5, then centrifuged and resuspended in 1 L of minimal media with 0.5% (w/v) mannitol, 1% (v/v) methanol, and 0.004% L-histidine. After a further 24 h of incubation, cells were harvested by

39
Chapter 2
centrifugation (3500 x g, 15 min, 4°C), washed with ice-cold demi water, and resuspended to a final OD600 of ~150 in 0.33 M sucrose, 0.3 M Tris.HCl (pH 7.4), 1 mM EDTA, 1 mM EGTA, 100 mM EACA, 5 mM DTT, 2 µg/mL pepstatin A, and one protease inhibitor tablet (cOmplete Mini EDTA-free, Roche) per 50 mL. 2.3.8 Membrane preparation and purification
Cells were disrupted by three sequential passes through the T series cell disrupter (Constant Systems, UK) at 39 kpsi. After the last passage, PMSF was added to the cell lysate to a final concentration of 1 mM. Intact cells and large debris were removed by centrifugation at 18,000 x g for 30 min at 4°C. Crude membranes were isolated from the supernatant by ultracentrifugation at 186,000 x g for 2 h. Membranes were suspended to homogeneity using a potter Elvehjem tissue grinder in 20 mM Tris-HCl pH 7.5, 300 mM sucrose, 0.1 mM CaCl2, 1 mM PMSF, 1 mM pepstatin A plus one tablet of protease inhibitor per 50 mL. Aliquots of 1 mL were snap frozen in liquid nitrogen and stored at -80°C.
For purification of Lyp1, crude membranes were diluted into 50 mM ammonium acetate (pH 7.5), 50 mM NaCl, 10% (v/v) glycerol, 15 mM imidazole, 2 mM PMSF and 1% (w/w) β-D-dodecylmaltoside (DDM) to a concentration of 5 mg/mL total protein. This solubilization mix was incubated for 30 min at 4°C with slow agitation. The insoluble pellet was removed by ultracentrifugation at 444,000 x g for 20 min. The supernatant was incubated with 0.5 mL of Ni-Sepharose resin under slow agitation at 4°C for 1 h. The resin was collected into a column and sequentially washed with 10 column volumes of buffer P (50 mM ammonium acetate pH 7.5 (at 24˚C), 50 mM NaCl, 10% v/v glycerol and 0.020% w/v DDM) containing 30 mM imidazole pH 7.5 and with buffer P containing 50 mM L-histidine. The protein was eluted with 2.5 mL of buffer P containing 235 mM L-histidine, applied in steps of 0.5 mL with 10 min intervals. Na-EDTA (stock concentration 500 mM) was added to each elution fraction to give a final concentration of 5 mM.
Elution fractions were pooled and concentrated to a volume of 0.5 mL on a VivaSpin column (Sartorius Stedim, Vivaspin 500, 100 kDa cut-off ) by centrifugation at 18,000 x g and 4°C. The protein sample was loaded onto a Superdex200 10/300 GL column (GE-Healthcare) attached to an AKTA purifier (GE-Healthcare) equilibrated with buffer P containing 100 mM L-histidine. The sample was analyzed by SDS-PAGE and the protein concentration determined at 280 nm absorption on a NanoDrop ND-1000 spectrophotometer (Isogen Lifescience), using an extinction coefficient of 126,110 M-1cm-1 and molecular mass of 97.7 kDa as determined by ProtParam (http://web.expasy.org/protparam).
Can1 was purified in a similar fashion to Lyp1, with the following changes. The solubilization mix was 3 mg/mL total protein in 20 mM Tris.HCl (pH 7.5), 50 mM NaCl, 10% (v/v) glycerol, 15 mM imidazole, 2mM PMSF, 2 mM DTT, and 0.5% (w/w) DDM. The Ni-Sepharose column was sequentially washed with 10 column volumes of buffer Q (50 mM Tris.HCl pH 7.5, 50 mM NaCl, 10%

40
Expressing S. cerevisiae transporters in P. pastoris
v/v glycerol, 2 mM DTT and 0.05% w/w DDM) containing 60 mM imidazole and with buffer Q containing 50 mM L-histidine. The protein was eluted with buffer Q containing 235 mM of L-histidine. A single 0.5 mL elution fraction was passed over a Superdex200 10/300 GL column in buffer Q with 20 mM Tris.HCl.
Samples were dissolved in Laemmli buffer and resolved on 10% or 12% polyacrylamide gels by SDS-PAGE. In-gel fluorescence images were collected prior to Coomassie staining and both were imaged using a Fujifilm LAS-3000 (Fujifilm, Tokyo, Japan).2.3.9 Liposome preparation and membrane reconstitution of Lyp1
For the reconstitution of Lyp1 into lipid vesicles, liposomes were prepared from total lipid extracts of S. cerevisiae and E. coli (Avanti Polar Lipids, USA). The lipids (20 mg/mL, in chloroform) were mixed in at a 1:2 ratio (total lipid extracts of S. cerevisiae:E. coli). Chloroform was removed by evaporation using a rotary vaporizer (Rotavapor R-3, BÜCHI, Flawil, Switzerland). Lipids were suspended in diethylether, evaporated, and then dissolved in 50 mM ammonium acetate (pH 7.5 at 24°C), 50 mM NaCl to a concentration of 10 mg/mL. The lipid solution was homogenized by tip sonication with a Vibra-Cell sonicator (Sonics & Materials, USA) at 4°C for 1 min, with 5 sec pulses and a 10 sec pause between every pulse. The amplitude was set to 60%.
Thawed liposomes were pelleted by ultracentrifugation at 444,000 x g for 20 min and re-dissolved in reconstitution buffer: 50 mM ammonium acetate (pH 7.5), 50 mM NaCl. In order to completely exchange buffers, liposomes were snap-frozen and thawed (at room temperature) three times. Liposomes were homogenized by repeated extrusion through a 400 nm polycarbonate filter, using the LiposoFast LF-50 (Avestin, Mannheim, Deutschland) with nitrogen flow. For reconstitution of the vesicles with protein, the liposomes were destabilized using Triton X-100 and titrated to a point beyond the saturation point (Rsat) as described by [179,180]; the final turbidity at 540 nm was at approximately 60% of Rsat.
Purified Lyp1 (w/w protein:lipid ratio of 1:1000) was mixed with detergent-destabilized liposomes and incubated for 15 min under slow agitation at 4°C. Bio-Beads SM-200 (Bio-Rad, Hercules, CA, USA) were sequentially added after 15, 30 and 60 min (10 mg/mg lipids). The final mixture was left to incubate overnight, after which a final batch of Bio-Beads was added and incubation continued for another two hours. Protein-containing liposomes (proteoliposomes) were separated from the Bio-Beads by filtration on a column followed by ultracentrifugation (444,000 x g at 4°C for 35 min). Proteoliposomes were suspended in 5 mM potassium phosphate (pH 7.0) at a concentration of 10 µg protein/mL, snap frozen and stored in liquid nitrogen. In order to exchange the inside buffers, centrifugation of the proteoliposomes at 444,000 x g at 4°C for 35 min was followed by suspension in the desired buffer, two freeze-thaw cycles, and eleven cycles of extrusion through a 400 nm polycarbonate filter.

41
Chapter 2
2.4 SupplementaryThe C-terminal GFP tag used in pPICZ vectors, whose full amino acid
sequence is shown below, contains a TEV protease cleavage site (ENLYFQ↓G, underlined) followed by GFP (in bold) and an Arg-Gly-Ser-His10 epitope (in italics). The residue pairs “QF” and “TS” are cloning artifacts.
ENLYFQGQFS KGEELFTGVV PILVELDGDV NGHKFSVSGE GEGDATYGKLTLKFICTTGK LPVPWPTLVT TLTYGVQCFS RYPDHMKRHD FFKSAMPEGYVQERTISFKD DGNYKTRAEV KFEGDTLVNR IELKGIDFKE DGNILGHKLEYNYNSHNVYI TADKQKNGIK ANFKIRHNIE DGSVQLADHY QQNTPIGDGPVLLPDNHYLS TQSALSKDPN EKRDHMVLLE FVTAAGITHG MDELYKTSRGSHHHHHHHHH H
The C-terminal YPet tag, whose full amino acid sequence is shown below, contains a TEV protease cleavage site (ENLYFQ↓G, underlined) followed by the fluorescent protein YPet (in bold) and an eight-residue His epitope (in italics). The residue pairs “GG” and “EL” are a short linker and a cloning artifact, respectively.
GGENLYFQGS KGEELFTGVV PILVELDGDV NGHKFSVSGE GEGDATYGKL TLKLLCTTGK LPVPWPTLVT TLGYGVQCFA RYPDHMKQHD FFKSAMPEGY VQERTIFFKD DGNYKTRAEV KFEGDTLVNR IELKGIDFKE DGNILGHKLE YNYNSHNVYI TADKQKNGIK ANFKIRHNIE DGGVQLADHY QQNTPIGDGP VLLPDNHYLS YQSALFKDPN EKRDHMVLLE FLTAAGITEG MNELYKELHH HHHHHH