The tRNA i Met is needed to start synthesis of a polypeptide chain
description
Transcript of The tRNA i Met is needed to start synthesis of a polypeptide chain

McGillMcGill BIOL200 - Fall 2008BIOL200 - Fall 2008
© R. Roy, 2008© R. Roy, 2008
77

McGillMcGill BIOL200 - Fall 2008BIOL200 - Fall 2008
© R. Roy, 2008© R. Roy, 2008
The tRNAThe tRNAiiMet
is needed to start is needed to start synthesis of a polypeptide chainsynthesis of a polypeptide chain
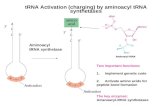
two types of tRNAs exist for the
methionine codon
both are charged by
the same aminoacyl
tRNA synthetase
tRNAMet is exclusively used for elongation of a polypeptide chain
tRNAiMet is exclusively
used for initiation of a polypeptide chain
the methionine in bacterial tRNAi
is modified by addition of a formyl group

McGillMcGill BIOL200 - Fall 2008BIOL200 - Fall 2008
© R. Roy, 2008© R. Roy, 2008
Ribosomal subunits have to be Ribosomal subunits have to be kept apart before translation kept apart before translation
initiationinitiation
the eukaryotic Initiation Factors (eIFs) eIF3 and eIF6 are required to keep the ribosomal subunits apart after they have finished the synthesis of a protein

McGillMcGill BIOL200 - Fall 2008BIOL200 - Fall 2008
© R. Roy, 2008© R. Roy, 2008
a ternary complex made up of eIF2, GTP and the tRNAiMet then binds the small ribosomal subunit together with the stabilizing factor eIF1a to form the 43S pre-initiation complex
This event can be regulated through phosphorylation of eIF2 after which it can no longer exchange GDP for the active GTP.
ternary complex
tRNAitRNAiMetMet is required for the is required for the formation of a pre-initiation formation of a pre-initiation
complexcomplex

McGillMcGill BIOL200 - Fall 2008BIOL200 - Fall 2008
© R. Roy, 2008© R. Roy, 2008
eIF4 is instrumental in recruiting eIF4 is instrumental in recruiting an mRNA to the preinitiation an mRNA to the preinitiation
complexcomplex next, a protein complex
named eIF4 binds the pre-initiation complex as well as the cap of an mRNA.
An associated helicase activity removes any secondary structure in the 5’ region of the transcript...

McGillMcGill BIOL200 - Fall 2008BIOL200 - Fall 2008
© R. Roy, 2008© R. Roy, 2008
The initiation complex scans the The initiation complex scans the mRNA for the start codonmRNA for the start codon
...secondary structure unwinding enables unhindered movement of the initiation complex along the mRNA to find the start codon, which is often embedded in a Kozak consensus sequence: ACCAUGG
eventually, the tRNAi
Met is positioned correctly at the translation start site the initiation factors are then released following GTP hydrolysis
the scanning requires energy from ATP hydrolysis

McGillMcGill BIOL200 - Fall 2008BIOL200 - Fall 2008
© R. Roy, 2008© R. Roy, 2008
Bring on the big one!Bring on the big one!
Finally,with the aid of eIF5 and energy from GTP hydrolysis…
the large 60 S ribosomal subunit is recruited,
the tRNAiMet is correctly
positioned at the P site,
The eIFs are released and translation begins

McGillMcGill BIOL200 - Fall 2008BIOL200 - Fall 2008
© R. Roy, 2008© R. Roy, 2008
the P site of the ribosome is the site where the initiator tRNA (tRNAi
Met) is bound
only the tRNAi can bind to this site during ribosome assembly
this is why the tRNAi
Met is so special...
The tRNAThe tRNAii is the only tRNA that can bind the is the only tRNA that can bind the
"P" position of the ribosome"P" position of the ribosome

McGillMcGill BIOL200 - Fall 2008BIOL200 - Fall 2008
© R. Roy, 2008© R. Roy, 2008
Entry of the next amino acid-loaded Entry of the next amino acid-loaded tRNA into the A site starts the tRNA into the A site starts the
elongation cycleelongation cycle a ternary complex of the elongation factor (EF) 1 alpha, GTP and a tRNAaa whose anticodon matches the next mRNA codon is allowed to enter the ribosomal A site

McGillMcGill BIOL200 - Fall 2008BIOL200 - Fall 2008
© R. Roy, 2008© R. Roy, 2008
Energy from GTP hydrolysis promotes Energy from GTP hydrolysis promotes a confirmational change in the a confirmational change in the
ribosomeribosome
upon binding of the ternary complex in the A site, EF1 alpha is released and concurrent GTP hydrolysis leads to aconformational change in the ribosome.
This change brings the amino acids linked to the tRNAs in the P and A site into close proximity

McGillMcGill BIOL200 - Fall 2008BIOL200 - Fall 2008
© R. Roy, 2008© R. Roy, 2008
the 23S rRNA catalyzes the peptide bond formation between the two amino acids without any help from proteins
the equivalent applies to the 28S rRNA in eukaryotic ribosomes
The ribosome is a ribozymeThe ribosome is a ribozyme ribozyme -> an enzymatically active RNA molecule
the peptidyltransferase activity, which results in peptide bond formation between the amino acids of the neighbouring aminoacyl-tRNAs, depends on the 23S rRNA in the large ribosomal subunit of prokaryotic ribosomes

McGillMcGill BIOL200 - Fall 2008BIOL200 - Fall 2008
© R. Roy, 2008© R. Roy, 2008
EF2 and energy from GTP hydrolysis EF2 and energy from GTP hydrolysis promotes translocation of the tRNAs promotes translocation of the tRNAs
into the E and P site respectivelyinto the E and P site respectively
upon peptide bond formation, EF2-bound GTP is recruited and GTP hydrolysis leads to a conformational change in the ribosome, resulting in the translocation of the tRNA molecules into the E and P site respectively
a new elongation cycle can begin

McGillMcGill BIOL200 - Fall 2008BIOL200 - Fall 2008
© R. Roy, 2008© R. Roy, 2008
In the next elongation cycle...In the next elongation cycle...
the tRNA in the E site, now rid of its amino acid, is kicked out of the ribosome during the conformational change associated with EF1 alpha release and GTP hydrolysis
the growing polypeptide chain is always covalently linked to one of the tRNAs in the P or A site
gets recycled
21
3
321

McGillMcGill BIOL200 - Fall 2008BIOL200 - Fall 2008
© R. Roy, 2008© R. Roy, 2008
RF1
RF2
UAG
UAA
UGA
Translation termination involves Translation termination involves termination factors that recognize termination factors that recognize
the stop codonsthe stop codons once the ribosome encounters a
STOP codon in the A site, this codon is recognized by a termination (release) factor, eRF1
eRF1 is thought to structurally mimic aminoacyl-tRNAs
another factor, GTP-bound eRF3, then associates with eRF1
in prokaryotes, two release factors for different STOP codons exist:

McGillMcGill BIOL200 - Fall 2008BIOL200 - Fall 2008
© R. Roy, 2008© R. Roy, 2008
hydrolysis of the GTP bound by eRF3 is necessary for the cleavage of the last amino acid in the P site from its tRNA
this results in the release of the polypeptide chain and the disassembly of the ribosome
eRF3 terminates translation by eRF3 terminates translation by releasing the polypeptide chainreleasing the polypeptide chain

McGillMcGill BIOL200 - Fall 2008BIOL200 - Fall 2008
© R. Roy, 2008© R. Roy, 2008
Interactions between the 5’ and 3’ Interactions between the 5’ and 3’ end of the translated mRNA favour end of the translated mRNA favour
reinitiationreinitiation • Protein:protein interactions
indicate that the complexes associated with the 3’ end of the mRNA interact with protein complexes associated with translational initiation.
• Maintaining the 3’ terminal region of the mRNA in close proximity to the site of translational initiation could favour reinitiation.
• Efficient translation often is associated with stabilisation