Supplementary Information 1) Supplementary Methods fileStep qRT-PCR kit from Invitrogen (Carlsbad,...
Transcript of Supplementary Information 1) Supplementary Methods fileStep qRT-PCR kit from Invitrogen (Carlsbad,...

Carlson et al.
Supplementary Information
1) Supplementary Methods
Mice: C57BL/6 (B6), B6.SJL (CD45.1 congenic B6), Rag-2-/- mice were
purchased from Jackson Labs and Taconic Farms. Mice carrying the KLF2
knockout allele(1) were backcrossed to C57BL/6 (N>9). All experimental
procedures were carried under IACUC guidelines at the University of Minnesota.
Fetal Liver Chimeras: Following timed mating of KLF2+/- breeders, pups were
isolated at day 12.5 of gestation and genotyped by PCR. Fetal livers from the
pups were transferred into irradiated (440 rads) Rag-/- hosts. After time for
hematopoetic development (>8 weeks), the fetal liver chimeras (FLC) were
sacrificed and tissues harvested for flow cytometry or adoptive transfers.
Adoptive Transfers: Thymocytes were harvested from FLC and transferred
into unmanipulated B6.SJL hosts. Typically, 2-3 x 106 donor CD4 SP were
transferred per recipient animal. Following the indicated time, the hosts were
sacrificed and the presence of donor cells monitored in the blood, spleen and
pooled lymph nodes via flow cytometry, gating on CD45.2+ve cells.
Intrathymic Injections: In order to assess thymic emigration in a situation
where KLF2-/- and wt cells were present in the same animal, we created
secondary mixed chimeras as follows. Bone marrow from KLF2-/- or +/- FLC
was harvested and mixed with B6.SJL bone marrow, and introduced into
lethally irradiated (900 rads) B6.SJL hosts. Variable levels of chimerism were
observed in such animals whether the FLC donor was KFL2-/- or FKL2+/-,

presumably due to variable chimerism in the hematopoietic stem cells of the
FLC donors. After >8 weeks, animals were given an intrathymic injection of
10ul of a 5mg/ml solution of biotin-NHS, as described. 36 hours later thymus,
LN, spleen, and blood were stained with streptavidin PE, and other markers,
and analyzed by flow cytometry. Use of this in vivo biotinylation approach was
based on the studies of Rybak et al. (2).
Flow cytometry: Cells were stained with antibodies purchased from BD or
eBioscience. CCL19-Fc fusion protein (a generous gift of Jason Cyster, UCSF)
was used to detect CCR7 expression. Cells were analyzed on Becton Dickinson
FACScalibur and LSR II instruments and the data was processed using Diva and
FlowJo software.
In vitro cell culture: FLC thymocytes were harvested and placed in culture
with RP10 (RPMI media, antibiotics, 2-ME and 10% fetal calf serum), with or
without 10ng/ml recombinant mouse IL-7 (R&D Systems, Minneapolis, MN).
Recovery of thymocytes was determined on successive days by flow cytometry,
excluding dead cells by staining for Annexin V (Becton Dickinson).
Cell Sorting and Real-time RT-PCR: Fluorescence-activated cell sorting
was used to purify CD4+ single positive and CD4+CD8+ double positive
thymocytes from either KLF2-/- or KLF2+/- animals. Each group was sorted in at
least three experiments. PE-conjugated anti-CD4 and APC-conjugated anti-CD8
antibodies used for sorting were from BD PharMingen (San Diego, CA) and
eBioscience (San Diego, CA), respectively. Sorting was performed on a
FACSVantage (Becton Dickinson) and was reliably >90% of target population.
RNA was isolated from sorted populations using the Qiagen (Valencia, CA)
RNeasy kit and cDNA was produced using the SuperScriptIII Platinum Two-

Step qRT-PCR kit from Invitrogen (Carlsbad, CA). One µl of cDNA generated
from 8-24ng of total RNA was then used with the QuantiTect SYBR Green PCR
kit from Qiagen. Reactions were quantitated using the SmartCycler real-time
PCR machine from Cepheid (Sunnyvale, CA). Real-time RT-PCR primers for β-
Catenin were used to normalize samples for Figure 3b, and similar trends were
obtained using primers for HPRT and GAPDH (the latter was used for the data in
figure S3). Primers were as follows; β-Catenin: 5'-TCCACGCAGCGGTGTC-3' & 5'-
GAGCCGTCAGTGCAGGAG-3', S1P1: 5’-GTGTAGACCCAGAGTCCTGCG-3’ & 5’-
AGCTTTTCCTTGGCTGGAGAG-3’, CCR7: 5’-CAGCCTTCCTGTGTGATTTCTACA-3’
& 5’-ACCACCAGCACGTTTTTCCT-3’, CD62L: 5’-GTGGAGCATCTGGAAACTGG-3’ &
5’-CGGCTACAGGAATGAAGAGG-3’, β7-Integrin: 5’-GGACGACTTGGAACGTGTG-3’
& 5’-CGTTTTGTCCACGAAGGAG-3’, CCR4: 5’-AGCAGCCACTCTCCCATTC-3’ & 5’-
GCACTTCCTCTCTTCCCATTT-3’, CCR9: 5’-GTCACCTTGGGGTTTTTCCT-3’ & 5’-
TGGATGACTTCTTGGCCTGT-3’, Tdt (S): 5’-GAAGCCACAGAGGATGAAGAGC-3’ &
5’-GAAACACCCTCTTAGTCCTG-3’, Tdt (L): 5’-GAAGCCACAGAGGATGAAGAGC-3’
& 5’-CCATCCAAAGGTGAAATCGTGAC-3’, Rag1: 5’-TCTCTGTGGCATCGAGTGTT-
3’ & 5’-AAGGAGGCAGCCATGTTG-3’. Primers for Bcl-2 (3) and others (4) were
as previously described.
Chromatin Immunoprecipitation assay: ChIP assays were performed by
modification of previously reported procedures (5-7). Briefly, J/KLF2 cells were
treated with Dox and without Dox for 48 h, respectively. Native protein-DNA
complexes were cross-linked by 1% formaldehyde for 10 min. Equal aliquots of
isolated chromatin DNA were subjected to immunoprecipitation with a rabbit
anti-HA Ab or rabbit IgG control. The DNA fragments associated with specific
immunoprecipitates or with the control IgG were purified and used as
templates for the PCR to amplify 150 bp of the S1P1 promoter sequence
starting141 bp before the potential transcription start site, which includes a

CACCC consensus KLF-family binding site (see Fig S4). The primer set (set 1)
was (5'-primer) GATCTTTCCTGGACAGTGCG and (3‘-primer)
CTGCTACGCGAAGTCACCCAG for this fragment (indicated on Fig S4). Another
primer set (set 2) amplifies a 106 bp fragment starting 474 bp upstream of the
proposed transcript start site, which includes another CACCC site. This
fragment served as a negative control to confirm KLF2 binding to the more
proximal promoter region. Primer set 2 was as follows: (5'-primer)
CACAAGCTCAGCACACCGATC and (3'-primer) CTCTCGAAAAGTCTGAGGAGG.
S1P1 promoter reporter assay: A -287 S1P1 promoter fragment was generated
by PCR, then ligated into the Nhe I site of the luciferase expression vector
pGL3-basic (Promega), designated as S1P1luc. For S1P1 promoter activity,
transient transfections of Jurkat T cells and HepG2 cells were performed as
previously described (8). pSV β-gal vector was cotransfected as an internal
control. Jurkat T cells were cultured in 10%FBS/RPMI-1640 media, while HepG2
cells were cultured in 10%FBS/DMEM. Cell lysates were prepared 48hrs later,
and luciferase activity was measured using a MonolightTM 3010 Luminometer.
The S1P1 promoter activity was expressed as the ratio of luciferase/β-
galactoside activity. All transfections were performed in triplicate from three
independent experiments.
Statistical Analysis: An unpaired, two-tailed Student’s t test was applied to
determine p-values, using GraphPad Prism software (San Diego, CA). In most
cases, the analysis used untransformed data, however in the case of S1P1 real-
time RT-PCR data (Figs. 3 and S3) and CD62L (Fig. 3), the large difference in
signal values lead to significant differences in variance, and hence those data
were subjected to Log10 transformation before calculation of p-values. In order

to determine p-values in Figure S3, samples showing “infinite” fold-change
between DP and CD4 SP groups were given an arbitrary value of 2000.

References for Methods
1. Wani, M. A., R. T. Means, Jr., and J. B. Lingrel. 1998. Loss of LKLF function results in embryonic lethality in mice. Transgenic Res 7:229-238.
2. Rybak, J. N., A. Ettorre, B. Kaissling, R. Giavazzi, D. Neri, and G. Elia. 2005. In vivo protein biotinylation for identification of organ-specific antigens accessible from the vasculature. Nat Methods 2:291-298.
3. Kimura, S., T. Kawakami, Y. Kawa, Y. Soma, T. Kushimoto, M. Nakamura, H. Watabe, S. Ooka, and M. Mizoguchi. 2005. Bcl-2 reduced and fas activated by the inhibition of stem cell factor/KIT signaling in murine melanocyte precursors. J Invest Dermatol 124:229-234.
4. Mick, V. E., T. K. Starr, T. M. McCaughtry, L. K. McNeil, and K. A. Hogquist. 2004. The regulated expression of a diverse set of genes during thymocyte positive selection in vivo. J Immunol 173:5434-5444.
5. Wells, J., and P. J. Farnham. 2002. Characterizing transcription factor binding sites using formaldehyde crosslinking and immunoprecipitation. Methods 26:48-56.
6. Ahmad, N., and J. B. Lingrel. 2005. Kruppel-like factor 2 transcriptional regulation involves heterogeneous nuclear ribonucleoproteins and acetyltransferases. Biochemistry 44:6276-6285.
7. Wu, J., and J. B. Lingrel. 2005. Kruppel-like factor 2, a novel immediate-early transcriptional factor, regulates IL-2 expression in T lymphocyte activation. J Immunol 175:3060-3066.
8. Wu, J., and J. B. Lingrel. 2004. KLF2 inhibits Jurkat T leukemia cell growth via upregulation of cyclin-dependent kinase inhibitor p21WAF1/CIP1. Oncogene 23:8088-8096.

2) Details of Grant Support
This work was supported by the following grants from the NIH: Immunology training grant T32 AI007313 to B.T.E., R01 AI39560 to K.A.H. and R01 AI38903 to S.C.J. Other support came from the American Cancer Society (RPG-99-264-01 to S.C.J.) and the Cancer Research Institute (postdoctoral fellowship to C.M.C.)

3) Supplementary Figures
Supplementary Figures S1, S2, S3 and S4, with legends, are given on the
following pages.

Fig S1
Fig S1. Normal in vitro survival of KLF2-/- mature thymocytes. KLF2-/- and KLF2+/- FLC were generated as described in the text, except that in this experiment both strains also expressed the OT-I TCR transgene. Bulk thymocytes were cultured in media with or without IL-7 (10ng/ml), and the fraction of live CD8 SP thymocytes was calculated at the indicated time points. Dead cells were excluded by Annexin-V staining. The y-axis shows the fraction of starting CD4-8+ cells recovered. Similar data were obtained in two additional independent experiments.

050416 CD4 D7
Het Spl KO Spl Het LN KO LN Het Tot KO Tot
103
104
105
KO LN
KO Spl
KO Tot
Het LN
Het Spl
Het Tot
Cells and Tissue
CD
4 S
P R
eco
very
Don
or C
D4+
cel
ls
LN Spl. LN+Spl.
050416 CD8 D7
Het Spl KO Spl Het LN KO LN Het Tot KO Tot
103
104
105
KO Spl
KO LN
KO Tot
Het LN
Het Spl
Het Tot
Cells and Tissue
CD
8 S
P R
eco
very
Don
or C
D8+
cel
ls
LN Spl. LN+Spl.
Filled = KLF2-/-Open = KLF2+/-
Fig S2a)
b)
100 101 102 103 104
CD62L
0
20
40
60
80
100
% o
f Max
100 101 102 103 104
CD69
0
20
40
60
80
100
% o
f Max
CD4+ donorcells from:
KLF2+/-
KLF2-/-
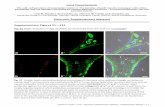
Fig S2. Aberrant trafficking and phenotype of KLF2-/- SP thymocytes after adoptive transfer. KLF2+/- and KLF2-/- FLC were generated as described in Methods and bulk thymocytes were adoptively transferred into B6.SJL recipients as described in Fig 2. (a) At 7 days post transfer, the frequency of donor derived CD4SP (left panel) and CD8SP (right panel) cells were calculated from the indicated tissues. Similar findings were observed with cells harvested at days 1 and 6 post adoptive transfer in two independent experiments. (b) In a different experiment, host animals were sacrificed 6 days after transfer and splenocytes stained for expression of CD69 and CD62L. Data are shown for CD4+ donor T cell in duplicate recipient animals. Similar data were obtained from two independent experiments. Tissue distribution in this experiment was similar to that shown in (a) (data not shown).

-10-100-1000 10 100 10001
Rag1Tdt (L)Tdt (S)PlxnD1CCR9CCR4
Sema4aRgs3Ian1Bcl-2Add3CCR7
β7 IntegrinCD62LS1P1
-1Relative fold change from DP to SP
∞
∞∞
p=0.0087
Fig S3
p=0.045
p=0.0053
Fig S3. Real-time PCR analysis of gene expression in KLF2-/- and KLF2+/- DP and CD4 SP thymocytes. FLC were generated as before and the DP and CD4 SP populations sorted by flow cytometry. Samples were prepared for real-time RT-PCR and assayed for expression of the indicated genes. Expression was monitored relative to Gapdh as a control. The figure shows the average fold change (positive or negative) for the indicated genes between the DP and CD4 SP stages in KLF2+/- (open bars) or KLF2-/- (filled bars) cells, with error bars indicating the SD. The symbol “!” indicates that expression was undetectable in at least one DP
sample but was detectable in the corresponding CD4 SP samples. For the purposes of statistical comparisons, these “infinite fold-change” samples were assigned an arbitrary value of 2000. The p-values for Add-3, CD62L and S1P1 are indicated: all others had a p-value of >0.05. Data derive from at least three independent experiments.
KLF2+/-
KLF2-/-

Mouse: tctgccccagatctttctgggaggctgcttcttagcagcccagaccctgaggggccccct ||||||||||||||||| ||| ||| ||| ||||| |||| || | ||||||||Human: tctgccccagatctttcctggacagtgcgtctcagcagttcagatccgg--gggccccca
Mouse: gccgctggcagagggcggaggagttaaaagcattaacccctcccagtcccttcctagagg || || ||||||||| || | | || ||||||||||||||||| | |||||| ||Human: gc---tgacagagggcgtggggggttaaggcattaacccctcccagcctcttcctgaaga
Mouse: agccacccagcctcggcggggcgctcagagacttcgtcttgcaaa | ||||||||||| |||| |||||| | ||||||| | |||Human: aaccacccagccttggcgcggcgctgggtgacttcgcgtagcagg
Transcription start site
Fig S4
Fig S4. Sequence of human and mouse S1P1 promoters, showing CACCC sequence and location of primers
used for CHiP assay. The figure indicates the proposed transcriptional start site for human S1P1 (arrow) and
the location of the KLF-family consensus binding motif CACCC (underlined). The location of “Primer Set 1”
forward and reverse primers (used for CHiP assay in Fig 3c) is shown by the red bars.