Structure Prediction
-
Upload
victor-moody -
Category
Documents
-
view
50 -
download
5
description
Transcript of Structure Prediction

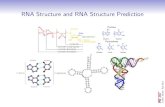
Structure Prediction

Tertiary protein structure: protein folding
Three main approaches:
[1] experimental determination (X-ray crystallography, NMR)
[2] Comparative modeling (based on homology)
[3] Ab initio (de novo) prediction (Dr. Ingo Ruczinski at JHSPH)

Experimental approaches to protein structure
[1] X-ray crystallography-- Used to determine 80% of structures-- Requires high protein concentration-- Requires crystals-- Able to trace amino acid side chains-- Earliest structure solved was myoglobin
[2] NMR-- Magnetic field applied to proteins in solution-- Largest structures: 350 amino acids (40 kD)-- Does not require crystallization

Steps in obtaining a protein structure
Target selection
Obtain, characterize protein
Determine, refine, model the structure
Deposit in database

X-ray crystallography
http://en.wikipedia.org/wiki/X-ray_diffraction
Sperm Whale Myoglobin



PDB
• April 08, 2008 – 50,000 proteins, 25 new experimentally determined structures each day
New folds
Old folds
New
PD
B s
truct
ure
s

Example 1wey

Ab initio protein prediction
• Starts with an attempt to derive secondary structure from the amino acid sequence– Predicting the likelihood that a subsequence will fold into an alpha-
helix, beta-sheet, or coil, using physicochemical parameters or HMMs and ANNs
– Able to accurately predict 3/4 of all local structures

Structure Characteristics

Beta Sheets

Ab Inito Prediction

Secondary structure prediction
Chou and Fasman (1974) developed an algorithmbased on the frequencies of amino acids found in helices, -sheets, and turns.
Proline: occurs at turns, but not in helices.
GOR (Garnier, Osguthorpe, Robson): related algorithm
Modern algorithms: use multiple sequence alignmentsand achieve higher success rate (about 70-75%)
Page 279-280

Table




Frequency Domain

Neural Networks

Training the Network
• Use PDB entries with validated secondary structures
• Measures of accuracy– Q3 Score percentage of protein correctly predicted
(trains to predicting the most abundant structure)– You get 50% if you just predict everything to be a
coil– Most methods get around 60% with this metric

Correlation Coeficient
• How correlated are the predictions for coils, helix and Beta-sheets to the real structures
• This ignores what we really want to get to– If the real structure has 3 coils, do we predict 3
coils?
• Segment overlap score (Sov) gives credit to how protein like the structure is, but it is correlated with Q3

Artificial Neural Network
PredictsStructure at this point

Danger
• You may train the network on your training set, but it may not generalize to other data
• Perhaps we should train several ANNs and then let them vote on the structure

Profile network from HeiDelberg• family (alignment is used as input) instead of just the
new sequence• On the first level, a window of length 13 around the
residue is used • The window slides down the sequence, making a
prediction for each residue• The input includes the frequency of amino acids
occurring in each position in the multiple alignment (In the example, there are 5 sequences in the multiple alignment)
• The second level takes these predictions from neural networks that are centered on neighboring proteins
• The third level does a jury selection

PHD
Predicts 4
Predicts 6Predicts 6
Predicts 5Predicts 5

Fold recognition (structural profiles)
• Attempts to find the best fit of a raw polypeptide sequence onto a library of known protein folds
• A prediction of the secondary structure of the unknown is made and compared with the secondary structure of each member of the library of folds

Threading
• Takes the fold recognition process a step further:– Empirical-energy functions for residue pair
interactions are used to mount the unknown onto the putative backbone in the best possible manner

Fold recognition by threading
Query sequence
Compatibility scores
Fold 1
Fold 2
Fold 3
Fold N

CASP
• http://www.predictioncenter.org/casp8/index.cgi

SCOP
• SCOP: Structural Classification of Proteins.• http://scop.mrc-lmb.cam.ac.uk/scop/

CATH
• CATH: Protein Structure Classification• Class (C), Architecture (A), Topology (T) and
Homologous superfamily (H)