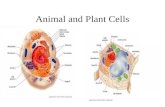
Stable spatial gradients of cytoskeleton assembly regulators
description
Transcript of Stable spatial gradients of cytoskeleton assembly regulators

Stable spatial gradients of cytoskeleton assembly regulators
David Odde
University of Minnesota

Microtubule Structure

Leng
th (
µm
)
Time (minutes)
“Catastrophe”
“Rescue”
Microtubule “Dynamic Instability” (DI)
Vg
Vs
kc
kr
see VanBuren et al., PNAS USA (2002)

Microtubules in Mitosis

Mitotic Spindle
spindle pole body
chromosome
kinetochore
kinetochoremicrotubule
spindle pole body
1.5 µmIn yeast:
~40 MTs10-20 µm
In animal cells:
~1000 MTs
Interpolarmicrotubule

Hypothesis
Dynamic instability alone is sufficient to explain the observed MT length distribution in the yeast mitotic spindle

Results: Cse4p-GFP Distribution
Experimentally Observed
Theoretically Predicted
?
2 µm

Leng
th (
µm
)
Time (minutes)
“Catastrophe”
“Rescue”
Microtubule “Dynamic Instability” (DI)
Vg
Vs
kc
kr

Point Spread Function (PSF)
• A point source of light is spread via diffraction through a circular aperture
• Modeling needs to account for PSF
-0.4-0.20+0.2+0.4 μm

Simulated Image Obtainedby Convolution of PSF and GWN
with Original Distribution
Original FluorophoreDistribution
Model-Convolution

Spindle Geometry

Results: Distribution of Cse4-GFP fluorescence
Experimentally Observed
Theoretically Predicted

Results: Distribution of Cse4-GFP fluorescence
x=0 x=L
QS QSSE

Results: DI Only Model
1000 nm

Results: DI Only Model

Alternative Models

Microtubule Chemotaxis
ImmobileKinase
MobilePhosphatase
Microtubule
A: Phosphorylated Protein Stabilizes MTsB: Unphosphorylated Protein Destabilizes MTs
Concentration
Position
MT Attractant
MT Repellant
X=0 X=L
k*Surface reaction B-->A
kHomogeneous reaction A-->B

Microtubule Chemotaxis:Op18
ImmobilePlx1
MobilePP2A
Microtubule
A: Op18-hi-PB: Op18-low-P Destabilizes MTs
Concentration
Position
Op18-hi-P
Op18-low-P
Chromatin

Microtubule Chemotaxis: RanGTP
ImmobileRCC1
MobileRanGAP
Microtubule
A: RanGTP Stabilizes MTsB: RanGDP
Concentration
Position
RanGTP
RanGDP
Chromatin

Model for Chemotactic Gradients of Phosphoprotein State
cAt
D 2cAx2
kcA Fick’s Second Law with First-Order HomogeneousReaction (A->B)
DcAx x0
k *cB 0 B.C. 1: Surface reaction at x=0 (B->A)
DcAx xL
0 B.C. 2: No net flux at x=L
cA cB cT Conservation of phosphoprotein

Predicted Concentration Profile
where
Y cA cT
X x L
kL2
D
A*e2
e2 1 * 1 e2 B*
e2 1 * 1 e2 * k
*LD
Y Ae X BeX
If k= 1 s-1, D=10-11 m2/s, and L=10 µm, then =3

Model Predictions: Effect of Homogeneous Reaction Rate
1.0
0.8
0.6
0.4
0.2
0.0
Con
cen
trati
on
, Y
1.00.80.60.40.20.0
Position, X

Model Predictions: Effect of Surface Reaction Rate
1.0
0.8
0.6
0.4
0.2
0.0
Con
cen
trati
on
, Y
1.00.80.60.40.20.0
Position, X

Microtubule Chemotaxis: RanGTP
ImmobileRCC1
MobileRanGAP
Microtubule
A: RanGTP Stabilizes MTsB: RanGDP
Concentration
Position
RanGTP
RanGDP
Chromatin



Results: Chemical Gradient and Polar Ejection Force Models
1000 nm

Cse4 Bleach @ end of simulation, mutant “Tension” model
LeftHalfSpindle
RightHalfSpindle
Figure 2

Cse4 Bleach @ End of Simulation, wild-type, “Gradient-Only” Model
RightHalfSpindle
LeftHalfSpindle
Figure 4

Mitotic Spindle
Conclusion: Spatial gradients in MT DI parameter(s)may play a role in mediating budding yeast mitotis
F FF F

X
X
X
Y
Z
Y
Simulated Actin FilamentDendritic Branching
Simulated Image of Actin FilamentDendritic Branching
Model-Convolution: Application to Dendritic Actin Filament Branching

Simulated Image Obtainedby Model-Convolution of
Original Distribution
Original FluorophoreDistribution
Image Obtained by Deconvolution
of Simulated Image
Potential Pitfalls of Deconvolution

Acknowledgements
• Whitaker Foundation
• National Science Foundation

Comparing Models to Microscopy
Molecular Theory Molecular Reality
Microscopic Observations
Model Predictions ???
Fluorescence Microscope
Computer Simulation