RNA Sample preparation and analysis tools from Ambion...
Transcript of RNA Sample preparation and analysis tools from Ambion...
1Life Technologies™ Proprietary |
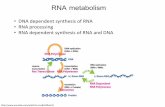
RNA Sample preparation and analysis tools from Ambion and Life Technologies
2Life Technologies™ Proprietary |
Agenda
RNA introductionRNA sample preparation microRNASpecial Samples− Blood− Single Cells− FFPE− ExosomesRNA-Seq
4Life Technologies™ Proprietary |
Working with RNAKeep an “RNAase‐free zone”− set aside an area that is not used for biological samples or enzyme work
− prescriptive steps: clean the surface with RNase ZAP®
− proscriptive remediations: clean all pipet barrels with RNase ZAP®, replace reagents , precious reagents can be checked with RNaseAlert®
5Life Technologies™ Proprietary |
Working with RNAStart with Certified RNase‐free Reagents and Plastics
RNase‐free Water and other common reagents
RNase‐free Tubes and Tips
7Life Technologies™ Proprietary |
RNA Purification Products
Reagents
Columns
Plates
Beads
BrandRNA Type
TRIzol®TRIzol® LS
PureLink™mirVana™
PureLink™mRNA Catcher™
MagMax™RiboMinus™
Total RNA
Total RNAmicroRNA
Total RNAmRNA
mRNATranscriptome
8Life Technologies™ Proprietary |
Trizol®
The most trusted and referenced reagent for preparing high quality and intact RNA− Unparalled lysis capability
− Flexible formulations for difficult samples (ietissues, cells, serum, viruses, and bacteria)
− Scalable
− Single step, monophasic solutions (phenol and guanidine isothiocyanate)
− Low gDNA contamination
9Life Technologies™ Proprietary |
Purelink™ RNA Mini‐ High Yield
Simplified total RNA purification from cells and tissues using spin columns− Incl. animal, plant, yeast,
bacterial, or blood − Purify RNA in less than 20
minutes− No hazardous or toxic
chemical disposal (no phenol)
− Scalable RNA purification up to 1,000 μg of purified RNA from a single prep
10Life Technologies™ Proprietary |
• Combines best extraction with easy to use format; high yield
• Improved purity
• Recommended protocol for Microarray
TRIzol® Plus – Best of both worlds
+
11Life Technologies™ Proprietary | 11
Magnetic bead - MagMax™
Magnetic bead based purification with flexible formats
Manual single tube or 96 well formats
Automated 24 or 96 well formatsIncreased productivity from the significant time savings vs column based purification
process 96 lysates in ~15min
Simple protocol
Specialized protocols for high RNA yield and purity from diverse sample types:
Cell free viral / biological samples, swabs, blood, plants, feces, environmental samples
12Life Technologies™ Proprietary |
TaqMan® Gene Expression Cells-to-CT™ Kit
Complete Optimized Workflow-includes sample prep, DNase, reverse transcription reagents and TaqMan® Gene Expression Master Mix for convenient real-time PCR studies
Compatibility-Components were co-developed to provide the ultimate in sensitivity
Simple and Fast-Formulated master mixes mean only 6 components for entire workflow. Sample preparation can be completed in 2 steps in less than 10 minutes, and is easily
automatable.
13Life Technologies™ Proprietary |
3 Steps
1. Lysis at Room Temp
2. Reverse Transcription
3. Real-time PCR
Sample Prep:
All at room temperature
Genomic DNA removed at same time cells are lysed
Can be done directly in culture plates
No centrifugations
No organics
No filter to clog
Cells-to-CT™ workflow – 10 to 100,000 cells
14Life Technologies™ Proprietary |
Purified RNA
Cells-to-CT™Kit
Cells-to-CT™ Kit
As the sample input is decreased, the percentage loss increases during purification. So at lower cell inputs there is quite a difference compared to purified RNA.
Equal or better performance and sensitivity compared to purified RNA
15Life Technologies™ Proprietary |
•TaqMan ® Fast Cells-to-CT™ Kit lysates and purified RNA from 4000 cells were prepared and evaluated with 163 TaqMan® Gene Expression Assays.
•20% of RT reaction was lysate/RNA.
•20% of the qPCR was RT.
•Similar data seen with miRNA
Equivalent mRNA profiles from Cells-to-CT™ and pure RNA with 163 TaqMan® Assays
16Life Technologies™ Proprietary |
MicroRNAs: Macro Significance
MicroRNAs (miRNAs) are short 18-25 nt RNA molecules, found in abundance in plants and animals
miRNAs are post-transcriptional regulators that bind to complementary sequences on target mRNAs, resulting in translational repression and gene silencing
the human genome encodes about 2000 miRNAs, which may target and regulate about 50% of mammalian genes
each miRNA typically regulates multiple mRNAs, and many mRNAs are regulated by several miRNAs.
17Life Technologies™ Proprietary |
MicroRNA Purification with mirVana™ Kits
mirVana™ Kits enable isolation of total RNA while retaining the small RNA fraction - including microRNAs down to 10-mers.
Ideal for performing RNA-seq, microarrays, TaqMan®/SYBR qPCR analysis
Non-Coding RNA
Easy workflow: Sample to Purified RNA in 30 Minutes:
18Life Technologies™ Proprietary |
TaqMan® MicroRNA Cells-to-CT™ KitFast, simple, and convenient—Prepare samples in ~10 minutes; eliminate tedious RNA isolation
Robust performance—Results equivalent to purified RNA
Broad input dynamic range—Linear detection from 10 to 100,000 cells
• 1000 HeLa cells. RT and subsequent PCR reactions were done for 144 miRNAs. 111 had a CT below 35 and plotted on the graph.
y = 0.9765x + 1.4269R2 = 0.9307
22
24
26
28
30
32
34
36
38
22 24 26 28 30 32 34 36 38Purified RNA CTs
Cel
ls-to
-CT
lysa
te (C
T)
19Life Technologies™ Proprietary |
NEW - TaqMan® miRNA ABC Kit
Targeted microRNA Isolation for Expression Analysis– Achieved by hybridization of miRNAs to anti‐miRNA oligos attached to
Dynabeads®
– Anti-miRNA oligos correspond to the assays in the TaqMan® Array Human MicroRNA A and B Cards
Entire workflow is typically 75 minutesBeads have capacity to capture 128-380 fmol of each miRNA
20Life Technologies™ Proprietary |
TaqMan® miRNA ABC Kit: Data using Human Panel A Kit
Purified MicroRNA from serum, plasma and FFPE samples were analyzed using a set of TaqMan® microRNA (single tube) Assays− Each bar represents duplicate
sample preparations and quadruple PCR replicates.
Excellent Reproducibility
21Life Technologies™ Proprietary |
Pri-miRNA, miRNA & All Non-coding RNA TaqMan® Assays
TaqMan® Assays provide a highly reliable way to measure expression of various RNA transcripts.
Provides high sensitivity, specificity, dynamic range & superior data quality
Delivers faster time-to-results
Requires no design expertise
Works under universal cycling conditions
Detects only Pri-miRNA, miRNA or non-coding RNA transcripts
TaqMan® MicroRNA
Assays
TaqMan® Pri-miRNA
Assays
22Life Technologies™ Proprietary |
MicroRNA Discovery with RNA-Seq
RNA sequencing is the leading hypothesis-free method for microRNA analysis & enables the discovery of new miRNAsThe number of novel miRNAs discovered has increased exponentially since the onset of Next Gen Sequencing
0
5000
10000
15000
1.1
1.3
1.5
2.1 3 4
5.1 7 8
8.2
9.1
10.1 12 14 16 18
Num
ber of Entries
miRBase Version
Sanger miRBase Growth
Distribution Run time Throughput
Ion Torrent(PGM™)
Wide – low barrier to entry, affordable system
2 hours – 35 bases & barcodes
3 barcoded small RNA samples (Ion 318™ Chip)
24Life Technologies™ Proprietary |
Blood-to-Ct™ kit pre-screening and MagMax™ purification
Take 500µL aliquot
Tempus® Stabilized Blood‐to‐CT™ kit*
Perform selected qRT-PCR assays on lysatesto select only tubes of interest:
Do full RNA purification only from these tubes.
Eliminate irrelevant samples
Tempus® MagMAX™ for Stabilized Blood Tubes Kit*
Process 12-96 samples in <1 hour
* Compatible with Tempus® Blood RNA for research use only
25Life Technologies™ Proprietary |
Reverse Transcription
Real-Time PCR
60 minute mark for 12 tubes 2 hr 12 min
Total for 12 tubes
Existing RNA Extraction Protocol
26Life Technologies™ Proprietary |
Collected blood
Aliquot 500 ul
Pellet and Wash 2X
Incubate 8 min at room temp.
Add Stop Solution
Reverse Transcription
Real-Time PCR
2 min. at room temp.
Stabilized Blood-to-CT™ Protocol (tubes and plates)
60 minutes for 12 or 96 samples in comparison to 2 (12) or 4 hrs (96)
Resuspend in Digestion Solution
27Life Technologies™ Proprietary |
5
10
15
20
25
30
35
40
5 10 15 20 25 30 35 40Pure Avg CT
Lysa
te A
vg C
1.5 mL Tube
96-well PlateR2 = 0.97
R2 = 0.97R2 = 0.95
mRNA miRNA
Comparable results to pure RNA
28Life Technologies™ Proprietary |
Conclusions
Stabilized blood− Provides immediate stabilization of host gene transcripts− Allows for delay between collection and analysis− Allows for long term storage of samplesStabilized Blood-to-CT™ Nucleic Acid Preparation Kits for qPCR
Enables expression analysis directly from a 500 μL aliquot of stabilized human whole blood without conventional RNA purification.
MagMAX™ for Stabilized Blood RNA Islolation KitsPurify high quality total RNA, including microRNAs, from blood RNA tubes.
30Life Technologies™ Proprietary |
• 4 functional steps: cell lysis, reverse transcription, cDNA pre-amplification, and Real-Time PCR
• gDNA removal occur at the same time as cell lysis - 5 minutes at room temperature
• All done in same tube (no loss due to tube transfers)
• Enable use of entire lysate sample in subsequent RT/PreAmp rxns
Single Cell-to-Ct™ kit: a complete workflow to obtain statistically relevant single cell data
• Performance benefit with smaller volumes and allows use of 384-well plates
• Increased sensitivity (SuperScript III)• Optimized volumes in workflow
minimize errors in reaction assembly (no water addition in any step)
• Multiple stopping points• microRNA and DNA protocols
31Life Technologies™ Proprietary |
Product Performance
12
14
16
18
20
22
24
26
28C
T
1 Cell Equivalents
Single Cells 100 cells
20.4
21.6
13.6
N=84 single cells
32Life Technologies™ Proprietary |
Conclusions and Impacts
• There are significant differences from cell-to-cell
• Single cells have a large range of gene expression levels.
• Analyzing gene expression en masse gives an average profile and masks differences and variability (gene-to-gene and cell-to-cell)
• Small populations are lost when large populations are averaged
• Average expression patterns of single cell populations are similar to 100 cells.
33Life Technologies™ Proprietary |
The TaqMan® Single Cell-to-Ct™ kit
• Optimized reagents provide a simplified workflow for expression analysis of single cells by qRT-PCR
• Enables transfer of entire cell into each step
• No sample is lost during the reaction which occurs in a single tube
• Sensitivity is maximized by use of reagents such as SuperScript III and PreAmp Master Mix
• Variation introduced by the protocol is very low
• Enables the acquisition of statistically significant data sets
34Life Technologies™ Proprietary |
FFPE Samples
Formaldehyde fixation maintains tissue structure
Most common tissue storage method used today by pathologists
Central to cancer and disease research, biomarker discovery and personalized medicine
Used currently in conjunction with qRT‐PCR or qPCR studies but it is branching out to include:
− Microarrays
− Methylation studies
− miRNA work
− Sequencing
35Life Technologies™ Proprietary |
Challenges with FFPE Samples
Low Yield
‐Nucleic acid is both trapped and modified by extensive protein‐protein and protein‐nucleic acid crosslinks.
Chemical Modification and Fragmentation
‐High temperatures required for paraffin infiltration can accelerate chemical reactions which modify nucleic acid
‐Fragmentation and progressive degradation of nucleic occurs during long‐term storage due to chemical modifications
‐Nucleic acid is often chemically modified to a degree that renders it incompatible with many molecular analysis techniques
35
36Life Technologies™ Proprietary |
Noticeable differences between FFPE and Unfixed
Unfixed FFPE
Agilent Nanochip Results
37Life Technologies™ Proprietary |
MagMAX™ FFPE Isolation Kits ‐ Summary
• A high‐throughput total nucleic acid purification solution from FFPE samples.
• Isolation of both Total RNA and DNA
• Magnetic bead‐based isolation in 96‐well plate format to increase throughput
• Simplified automated workflow completed in 30 minutes.
• Less‐toxic workflow by eliminating xylene
38Life Technologies™ Proprietary |
Between method/tissue concordance ‐ PGM
fresh frozen tumor log2(RP100K)
RP100K = counts per 100k mapped reads (miRBase 16)
fresh frozen
NATlog2(RP1
00K) spearman= 0.843
FFPE tumor log2(RP100K)
FFPE
NATlog2(RP1
00K)
spearman= 0.840
spearman= 0.873
FFPE
tumor
log2(RP1
00K)
fresh frozen tumor log2(RP100K)
spearman= 0.860
fresh frozen NAT log2(RP100K)
FFPE
NAT log2(RP1
00K)
Observation: source of most variation mostly (and subtly) from tissue type and not the tissue preparation/storage method
Present miRNAs only (row sum across all runs >=10 counts)
39Life Technologies™ Proprietary |
• tiny microvesicles with nucleic acid and protein “cargo”
• secreted by all cells
• found in most body fluids• Blood, saliva, urine, breast milk and
cell culture media
• functions are diverse• Cellular communication
• Clean up cell’s trash
What are Exosomes and what do they do?
40Life Technologies™ Proprietary |
Ultracentrifugation is the current “Gold Standard” isolation method for exosomes
Proven methodProduces clean preps
But…Requires a large amount of starting materialYield can be very lowHighly labor intensive and time consuming − 2-3 days depending on sample type,
sample amount and if using gradient or cushion
41Life Technologies™ Proprietary |
Isolation of exosomes
Purification of exosomal RNA &
protein
Analysis of exosomal RNA &
protein
Exosome Immunoprecipitation (Protein A or Protein G)
Optimized Exosome workflow
42Life Technologies™ Proprietary |
Protein Analysis from Exosomes:Further purification using Immuno-magnetic isolation
43Life Technologies™ Proprietary |
Human serum-derived exosomal RNAisolated using Total Exosome Isolation Reagent
• Differential RNA representation of specific miRNAs
• Quantities normalized by the percentage of reads aligned to that transcript out of the total mapped.
• Mapped read count distribution by RNA species
Cell Free Plasma Exosome Isolation Reagent
Donor 1 Donor 2 Donor 1 Donor 2
Cell Free Plasma Exosome Isolation Reagent
44Life Technologies™ Proprietary |
The Ion RNA‐Seq Offering
44
• Preserves strand information
• Small RNA and/or WT libraries
•Optimized workflow
• Template preparation in just 3h with only minutes of hands‐on time
• 200bp read‐lengths
• Intuitive microarray‐like UI and workflows developed for Ion Torrent
RNA purification Library construction Sequencing Data analysis
45Life Technologies™ Proprietary |
RNA-Seq WorkflowSamples to results in a single day
45
2.5HOURS
mRNA
1HOUR
miRNA
46Life Technologies™ Proprietary |
• Library construction in ~ 6 hours• Sequence in as little as 1 hour for small RNAs and 2.5 hours for mRNAs
• 3 Ion Chips to tailor the number of reads by just changing the chip
• 16 barcodes for sample multiplexing• 35 to 200 base read length for sequencing miRNA or mRNA
• Easy library construction with automation friendly magnetic bead clean up steps to replace columns
• No gel size selection needed for small RNA libraries
Simple, Scalable, Fast Ion RNA-Seq v2mRNA and miRNA
Simplicity
Scalability
Speed
46
47Life Technologies™ Proprietary |
Ion Total RNA-Seq Kit v2: Simple & Fast Workflow
Confidential and Proprietary—DO NOT DUPLICATE47
Preparation of small RNAObtain total RNA then determine the quality
Enrich small RNA
Quantitate small RNA sample and determine input amount
Preparation of whole transcriptome RNA25-500 ng rRNA-depleted total RNA or 1-500 ng poly(A)
RNA or 100 ng total RNA
Fragment the RNA
Clean up the RNA
SOLiD™ amplified library constructionHybridize and ligate the RNA adapters
Perform reverse transcription
Purify the cDNA
Assess the yield and size distribution of the amplified DNA
SOLiD™ System templated bead preparation and sequencing
Preparation of small RNAPreparation of whole transcriptome RNA
PGM ™ amplified library construction
Amplify the cDNA (BC addition)
Purify the amplified DNA
IonSphere™ particle preparation and Ion PGM™sequencing
Magnetic beads
Magnetic beads
Magnetic beads
Magnetic beads
Magnetic beads
Magnetic beads
< 6 hours< 5 hours
48Life Technologies™ Proprietary |
Strong Correlation of Ion Proton™ Data with qPCR MAQC Data
TaqM
an™
R=0.958N=657 assays
(UHR/HBR)
49Life Technologies™ Proprietary |
Detection of Novel Exons & Alternate Splicingwith Ion PGM™ Sequencer data
Confidential and Proprietary—DO NOT DUPLICATE49
Ion PGM™ sequencer derived RNA-Seq results from analysis of a Ewings sarcoma cell line (data courtesy T. Triche, Childrens Hospital of Los Angeles).
50Life Technologies™ Proprietary |
Comprehensive quantitation of PGM reads using an iterative mapping strategy
White paper is available on the Ion Community that explains more details on processing Ion RNA-seq data
51Life Technologies™ Proprietary |
Single kit for preparation of either small RNA or whole transcriptome libraries
Maintains strand orientation
Start with as little as 1 ng poly(A) RNA or 25 ng of rRNA-depleted RNA isolated from 100 ng total RNA, or 1 ng enriched miRNA
Easy, fast (6 hr) library construction with automation friendly magnetic bead clean up steps
No gel size selection needed for small RNA libraries
Compatible with Ion Xpress™ RNA-Seq Barcode Kit 1-16
Benefits of Ion Total RNA-Seq Kit v2
Adaptor ligation
Reverse transcription
Amplification
Library Construction
51
52Life Technologies™ Proprietary |
For Research Use Only. Not for use in diagnosticprocedures.
© 2012 Life Technologies Corporation. All rights reserved.
The trademarks mentioned herein are the property of Life Technologies Corporation or their respective owners. TaqMan is a
registered trademark of Roche Molecular Systems, Inc.
CyBi is a registered trademark of CyBio AG. Zephyr is a registered trademark of Caliper Life Sciences, Inc. Biomek is a registered
trademark of Beckman Coulter, Inc. Freedom EVO is a registered trademark of Tecan Group Ltd.
53Life Technologies™ Proprietary |
RiboMinus™ Eukaryote System v2Removing cytoplamic and mitochondrial rRNA
Low Input RiboMinus™ Eukaryote System v2
Includes Mag Bead Cleanup
Or
RiboMinus™ Eukaryote System v2
Includes Mag Bead Cleanup
or
RiboMinus™ Eukaryote Kit v2Cleanup Columns, available separately
54Life Technologies™ Proprietary |
New RiboMinus Eukaryote for RNA-Seq V2
5S
5.8S
18S 28S
HeLa (RIN 9.8) Human Kidney (RIN 6.9)
18S
28S
More efficient removal of ribosomal RNA (rRNA). Effective with compromised RNA. Demonstrated effective prior to RNA‐Seq library generation!