Research Article Identification and Validation of...
-
Upload
nguyenkhue -
Category
Documents
-
view
218 -
download
0
Transcript of Research Article Identification and Validation of...

Research ArticleIdentification and Validation of Expressed Sequence Tags fromPigeonpea (Cajanus cajan L) Root
Ravi Ranjan Kumar12 Shailesh Yadav3 Shourabh Joshi3 Prithviraj P Bhandare2
Vinod Kumar Patil2 Pramod B Kulkarni2 Swati Sonkawade2 and G R Naik2
1 Vidya Pratishthanrsquos School of Biotechnology Vidyanagari Baramati Pune 413133 India2Department of Biotechnology Gulbarga University Gulbarga Karnataka 585106 India3 Institute of Biotechnology Acharya N G Ranga Agricultural University Hyderabad 500030 India
Correspondence should be addressed to G R Naik grnaikbiotechgmailcom
Received 14 November 2013 Accepted 15 April 2014 Published 6 May 2014
Academic Editor Soslashren K Rasmussen
Copyright copy 2014 Ravi Ranjan Kumar et al This is an open access article distributed under the Creative Commons AttributionLicense which permits unrestricted use distribution and reproduction in any medium provided the original work is properlycited
Pigeonpea (Cajanus cajan (L) Millsp) is an important food legume crop of rain fed agriculture in the arid and semiarid tropicsof the world It has deep and extensive root system which serves a number of important physiological and metabolic functions inplant development and growth In order to identify genes associated with pigeonpea root ESTs were generated from the root tissuesof pigeonpea (GRG-295 genotype) by normalized cDNA library A total of 105 high quality ESTs were generated by sequencing of250 random clones which resulted in 72 unigenes comprising 25 contigs and 47 singlets The ESTs were assigned to 9 functionalcategories on the basis of their putative function In order to validate the possible expression of transcripts four genes namelyS-adenosylmethionine synthetase phosphoglycerate kinase serine carboxypeptidase and methionine aminopeptidase were furtheranalyzed by reverse transcriptase PCR The possible role of the identified transcripts and their functions associated with root willalso be a valuable resource for the functional genomics study in legume crop
1 Introduction
Pigeonpea (Cajanus cajan L) Millsp (2119899 = 22) is a majorgrain legume of the arid and semiarid regions of the world [1]Though considered a minor crop pigeonpea is of consider-able importance in areas of South Asia (mainly on the Indiansubcontinent) Africa the Caribbean and Latin Americawhere it is a prominent source of protein in the human dietas well as wood for fuel and light duty structural applicationssuch as thatch for roofing [2] Pigeonpea has nowmoved froman ldquoorphan legume croprdquo to one of the promising pluseswheregenomics-assisted breeding approaches for a sustainable cropimprovement are routine by Pigeonpea Genome Initiativean effort of various researchers [3] The first pigeonpea ESTdataset provides a transcriptomic resource for gene discoveryand development of functionalmarkers associatedwith bioticstress resistance [4] Root is the major part of water andnutrition uptake in pigeonpea which has a deep and extensiveroot system that provides access to water stored deep in
the soil profile when that in the surface layer is depleted thissource of water is particularly important for long durationcrops In order to identify the associated genes in pigeonpearoot tissues a normalized cDNA library was constructedfrom pigeonpea root and expression analysis of the identifiedgenes was carried out by reverse transcriptase PCR (RT-PCR)technique
2 Materials and Methods
The pigeonpea genotype namely GRG-295 was selectedto construct cDNA library and identification of expressedsequence tags (ESTs) The seeds of GRG-295 pigeonpeagenotype were grown in petri dish for 15 days and irrigatedwith water At the end of the 15th day total RNA wasisolated from root tissues by using the Trizol reagent(Invitrogen Carlsbad CA USA) and mRNA was furtherisolated by using the PolyATract mRNA Isolation System
Hindawi Publishing CorporationInternational Journal of Plant GenomicsVolume 2014 Article ID 651912 7 pageshttpdxdoiorg1011552014651912
2 International Journal of Plant Genomics
Table 1 Primer sequences of subset of ESTs for RT-PCR analysis
Accession number Putative function Optimum 119879119898(∘C) Primer sequence (51015840-31015840)
JK973671 S-Adenosylmethionine synthetase 61 F-AGAGGAAAT CGGT GCTGGTGR-GCAGCAATTTGGTCGTTGGT
JK973674 Phosphoglycerate kinase 59 F-TCCCGATCCCGATACCCTACR-CAGCACGCTTTTCAGCAGTT
JK973715 Serine carboxypeptidase 60 F-ACATGAAGCTCAGTGGAGGAGR-AGCCATGGCCTCCAATCTTC
JK973726 Methionine aminopeptidase 60 F-GGCAT TGAAAGTTGGGCAGGR-GATTGCAGCACCGACATCAC
(Promega Madison WI USA) The quantification of RNAwas verified by absorption ratio of OD
260280and by for-
maldehyde gel electrophoresis The first and second strandsof cDNA were synthesized using Clontech SMARTer PCRcDNA Synthesis Kit The cDNAs were purified by the Min-Elute PCR purification kit (Qiagen Valencia CA USA) andligated into a pGEM-T easy vector (Promega Madison WIUSA) Ligated plasmid DNAs were used for transformationinto competent E coli DH5120572 strain Positive clones wereselected on an ampicillinIPTGX-Gal LB plate PlasmidDNA from positive clones were isolated by using REAL96 plasmid isolation kit (Qiagen Netherlands) andpurified DNA was used for single-pass Sanger sequencingby using M13FR universal sequencing primers on ABIsequencing machine 3500XL Genetic analyzer All the ESTswere processed using VecScreen (httpwwwncbinlmnihgovVecScreenVecScreenhtml) to remove vector and clon-ing oligo sequences and various contaminants to trim a highquality region Based on the qualified sequences the pre-dicted amino acid sequences were used to search for similarpeptide sequences to search for similar protein sequences inpublic database NCBI (httpwwwncbinlmnihgov) usingthe BLASTx search algorithm [5] by using default parametersof the program The similarity scores between the cDNAclones and known sequences were represented by BLASTxprobability E values Further the ESTs were classified intodifferent functional categories based on the knowledgeof biochemistry plant physiology and molecular biology(httpwwwMetaCycorg) GO (httpwwwebiacuk)and COG (httpwwwncbinlmnihgovCOG) tools andby searching related abstracts in PubMed
21 RT-PCR Analysis Total root RNAs isolated from pigeon-pea root tissues were used for reverse transcription poly-merase chain reaction (RT-PCR) analysis Genomic DNAcontamination was removed by DNase I First-strand cDNAwas synthesized from each 2120583g of total RNA sample usingClontech SMARTer PCR cDNA Synthesis Kit according tothe manufacturerrsquos protocol The cDNAs were purified usinga commercial column (Qiagen) To determine the expressionof candidate genes PCR was performed with 2120583L of thefirst-strand cDNA template and gene-specific primer pairsGene-specific RT-PCR primers were designed with Primer30 according to the EST sequences and were synthesizedcommercially General PCR was conducted with annealing
as required for the specific primer pairs (Table 1) RT-PCRexperimentswere repeated three times and the PCRproductswere detected on 15 agarose gel
3 Results and Discussion
Plant root systems serve a number of important functionsincluding anchoring the plant absorbingwater and nutrientsproducing amino acids and hormones and secreting organicacids enzymes and alkaloids [6] The physiological signifi-cance of roots is belied by their relative structural simplicityas compared to other plant organs major metabolic path-ways such as photosynthesis lacking in root tissues have astereotypical morphology that is conserved across taxa andthroughout the life cycle of individuals This combination ofphysiological relevance and structural simplicity has maderoots obvious targets for functional genomics analyses [7]As a major grain legume of semiarid tropics and a deep andextensive root system of pigeonpea represents an excellentsource of identification of ESTs associated with their roottissues So the presentworkwas focused on the study of genesassociated with pigeonpea root tissues
In the present investigation the cDNA library was con-structed in order to identify ESTs associated with pigeonpearoots and their functional analysis was carried out Thetotal RNA from the pigeonpea root tissue was isolatedand the first and second cDNA strand was synthesizedThe presence of the gene in plasmid construct of colonieswas confirmed by colony PCR The colony PCR showedthat the size of these inserts ranged from 200 to 800 bpOut of 400 bacterial clones the plasmid construct of 250positive recombinant clones was sequenced in single passedsequencing reaction from 31015840 end using M13 forwardreverseprimer and the sequence data was subjected to BLASTanalysis The leading sequences tailing of the sequence andpoor quality sequences were excluded firstly Finally 105 highquality ESTs were retained which were clustered into 72unigenes comprising 25 contigs and 47 singlets (Table 2) andwere compared with NCBI nonredundant protein databaseusing BLASTx algorithm and default parameters In BLASTxanalysis it was shown that most of the sequences were havinga significant homology with known proteins Sequences thathad no significant homology with protein database werecompared to nucleotide BLAST using default parameters
International Journal of Plant Genomics 3
Table 2 Summary of ESTs library
Total clone sequenced 250ESTs taken for analysis 105Number of unigenes 72Number of contigs 25Number of singlets 47Average length of unigenes 442 bpAverage length of ESTs 437 bp GC content of unigenes 503 GC content of ESTs 512
The ESTs were deposited to NCBI dbEST with the accessionnumber of JK973637 to JK973741 (Table 3)
TheESTswere categorized into 8 diverse functional groupconsisting of 10 transporter genes 6 signal transductiongenes 20 cell growth and transcriptional regulator genesand 26 metabolism genes In other genes 8 genes wereuncharacterized 8 hypothetical genes 10 genes with nosignificant match and 14 genes was involved in otherfunctionsThere were 8 of the ESTs found that did not showany match to known proteins in BLASTx program and thatsuggest novel nature of those genes (Figure 1)
In order to validate the ESTs generated from cDNAlibrary the expression of the four genes which were involvedin different metabolic pathways was analysed by RT-PCRRT-PCR results showed that the expression levels of fourcandidate genes namely S-adenosylmethionine synthetasephosphoglycerate kinase serine carboxypeptidase andmethio-nine aminopeptidase were clearly expressed in pigeonpearoots (Figure 2) It was concluded that overall there was agood agreement between the cDNA library data and the RT-PCR results
S-Adenosylmethionine synthetase (SAMS) comprises oftwo cDNAs in Pinus contorta among which SAMS1 isexpressed in roots and exhibits a specific expression patternin the meristem at the onset of adventitious root devel-opment [8] SAMS also catalyses the nucleophilic substi-tution reaction from between methionine and ATP intoS-adenosylmethionine which have central role in severalbiological process in plants namely methyl group donor intrimethylation of lignin DNA and alkaloids as well as donorof aminopropylmoieties in ethylene and polyamine synthesis[8 9]
Phosphoglycerate kinase superfamily has a diverse func-tion in numerous metabolic processes like generation ofprecursor metabolites and energy carbohydrate metabolismphosphorous metabolism glycolysis kinase activity ATPbinding activity and so forth The presence of phosphoglycer-ate kinase transcripts in the cDNA library supports its diversefunction in pigeonpea root
Serine carboxypeptidase differentially expressed in rootand other tissues is responsible for the synthesis of sinapoyl-choline and sinapoylmalate in Arabidopsis which encodes 51proteins annotated as serine carboxypeptidase-like enzymesand emerged as a new group of acyltransferase enzymesthat are able to modify plant natural products [10ndash12]
Hyp protein8
Transpoter10
Signaltransduction
6
Cell wall synthesis and transcriptional
regulator20
Metabolism26
Uncharacterized protein
8
Other functions14
No hits8
Figure 1 Functional classification of ESTs of pigeonpea generatedby single pass sequencing
M 4321
Figure 2 Validation of cDNA library results by RT-PCR (M-100 bp ladder 1-S-adenosylmethionine synthetase 2-phosphoglyceratekinase 3-serine carboxypeptidase and 4-methionine aminopepti-dase)
Methionine aminopeptidase a ubiquitous enzyme differen-tially expressed in root tissues is one of the central enzymes inprotein synthesis that catalyzes N-terminal methionine fromproteins [13]
A few important ESTs viz type 2 metallothionein TBP-associated factor ABC transporter and phosphatidylcholinetransfer protein were also abundantly found in the cDNAlibrary Reddy et al [14] found abundant metallothioneingenes in normalized library from rice leaves and suggestedthat they might perform essential functions of plant growthbeside metal detoxification TATA-binding protein and TBP-associated factors are transcriptional factors which are pre-dominantly involved in RNA polymerase II mediated tran-scription process [15] ABC transporters play an importantrole in organ growth plant nutrition plant developmentresponse to abiotic stress and the interaction of the plantwith its environment [16] Phosphatidylcholine is usuallythe most abundant phospholipids in animals and plantsoften amounting to almost 50 of the total and as such itis obviously the key building block of membrane bilayersmaking up a very high proportion of the outer leaflet of
4 International Journal of Plant Genomics
Table3Hom
olog
ysearch
oftheE
STsg
enerated
from
pigeon
pear
ootalong
with
their119864
valueu
singBL
AST
xprogramme
Slnum
ber
Gen
bank
accessionnu
mber
Leng
th(in
bp)
Hom
ologou
sprotein
119864-value
1JK
973637
560
Vesic
le-associatedmem
branep
rotein
727-lik
e[Glycinem
ax]
1119890minus111
2JK
973638
231
Mito
chon
drialinn
ermem
branep
rotein
OXA1-like[
Vitis
vinifer
a]5119890minus41
3JK
973639
630
Uncharacterized
protein[Arabidopsisthaliana
]9119890minus159
4JK
973640
621
Hypothetic
alprotein[Sorghum
bicolor]
9119890minus139
5JK
973641
570
GTP
-binding
proteinhfl
x[M
edica
gotru
ncatula]
6119890minus13
6JK
973642
412
Unk
nown[G
lycinem
ax]
11
7JK
973643
264
Secretorycarrier-associated
mem
branep
rotein
3-lik
e[Vitis
vinifer
a]2119890minus55
8JK
973644
438
Histon
e2[Populus
trichocarpa]
5119890minus63
9JK
973645
200
RNArecogn
ition
motif-containing
protein[Arabidopsisthaliana
]4119890minus10
10JK973646
870
Beta-glucosid
ase[
Arabidopsis
thaliana
]00
11JK
973647
555
Epsin
N-te
rminalho
molog
y(ENTH
)dom
ain-containing
protein[M
edica
gotru
ncatula]
4119890minus36
12JK
973648
210
Gam
ma-gliadinprecursor[Ricin
uscommun
is]7119890minus10
13JK
973649
210
AP2
ERF
andB3
domain-containing
transcrip
tionfactor
[Arabidopsisthaliana
]4119890minus40
14JK
973650
474
Glycine
dehydrogenase(decarborcylatingmito
chon
driallike)[G
lycinem
ax]
4119890minus102
15JK
973651
432
Sign
alrecogn
ition
particlesubu
nit[Arabidopsis
thaliana
]9119890minus80
16JK
973652
227
TBP-associated
factor
6B[Arabidopsisthaliana
]0051
17JK
973653
290
TBP-associated
factor
6B[Arabidopsisthaliana
]8119890minus48
18JK
973654
303
Transcrip
tionfactor
[Medica
gotru
ncatula]
6119890minus29
19JK
973655
432
Uncharacterized
protein[G
lycinem
ax]
8119890minus11
20JK
973656
237
Nuclear
cap-bind
ingproteinsubu
nit[Arabidopsis
thaliana
]8119890minus48
21JK
973657
501
Glucanendo
-13-beta-glucosidase-lik
eprotein
3-lik
e[Glycinem
ax]
2119890minus04
22JK
973658
499
DNA-
bind
ingproteinRA
V1[Zeamays]
4119890minus25
23JK
973659
560
Two-compo
nent
respon
seregulator
ARR
9[Vitisv
inifera]
1119890minus45
24JK
973660
210
Ribo
nucle
oside-diph
osph
ater
eductase
smallchain
like[Glycinem
ax]
2119890minus34
25JK
973661
748
Hypothetic
alprotein[O
ryza
sativa]
26JK
973662
742
Cyclin-depend
entp
rotein
kinase
complex
compo
nent
[Aspergillu
skaw
achii]
012
27JK
973663
406
Elon
gatio
nfactor
2[N
icotia
natobacum]
5119890minus39
28JK
973664
841
Expressedprotein[O
ryza
sativa]
73
29JK
973665
385
Hypothetic
alprotein[Vitisv
inifera]
21
30JK
973666
544
ADP-rib
osylationfactor-like
8d[N
icotia
natobacum]
2119890minus105
31JK
973667
510
Jmjcdo
main-containing
protein4-lik
e[Brachypodium
dista
chyon]
6119890minus77
32JK
973668
630
Nosignific
antm
atch
33JK
973669
804
Gag-pro[Pisu
msativ
um]
9119890minus18
34JK
973670
320
HypotheticalproteinMTR
[Medica
gotru
ncatula]
2119890minus13
35JK
973671
509
Putativ
eS-adeno
sylm
ethion
ines
ynthetase[Ca
psicu
mannu
m]
6119890minus114
36JK
973672
371
ABC
transporterF
family
mem
ber1-like
[Brachypodium
dista
chyon]
4119890minus60
37JK
973673
513
Predictedprotein[H
ordeum
vulga
re]
2119890minus89
38JK973674
465
Phosph
oglyceratekinasecytosolic[Triticu
maestivum]
2119890minus97
39JK973675
465
Phosph
oglyceratekinasecytosolic[Triticu
maestivum]
2119890minus97
40JK
973676
457
Unk
nown[Pice
asitchensis]
097
41JK
973677
785
GTP
-binding
signalrecognitio
nparticleSR
P54[M
edica
gotru
ncatula]
7119890minus51
International Journal of Plant Genomics 5
Table3Con
tinued
Slnum
ber
Gen
bank
accessionnu
mber
Leng
th(in
bp)
Hom
ologou
sprotein
119864-value
42JK
973678
241
Sign
alrecogn
ition
particle54
kDAsubu
nitp
recursor
[Pisu
msativ
um]
0044
43JK
973679
475
ProteinSE
T-lik
e[Brachypodium
dista
chyon]
7119890minus33
44JK
973680
350
EIN3-bind
ingF-bo
xprotein[Brachypodium
dista
chyon]
8119890minus65
45JK
973681
389
Predictedprotein[H
ordeum
vulga
re]
4119890minus27
46JK
973682
272
40Srib
osom
alproteinS24-2-lik
e[Brachypodium
dista
chyon]
2119890minus57
47JK973683
210
Putativ
ecalcium
exchanger[Triticum
dicoccoides]
4119890minus27
48JK
973684
478
Thioredo
xin[M
edica
gotru
ncatula]
2119890minus75
49JK
973685
210
Nosignific
antm
atch
50JK
973686
553
Root
nodu
leextension[Pisu
msativa]
5119890minus04
51JK
973687
600
Sign
alrecogn
ition
particleprotein[O
ryza
sativa]
5119890minus08
52JK
973688
544
ADP-rib
osylationfactor-like
8d[N
icotia
natobacum]
2119890minus105
53JK
973689
395
MLO
-like
protein[M
edica
gotru
ncatula]
2119890minus24
54JK973690
250
Phosph
atidylcholinetransferp
rotein
[Ricinu
scom
mun
is]3119890minus09
55JK973691
215
Phosph
atidylcholinetransferp
rotein
[Ricinu
scom
mun
is]3119890minus09
56JK
973692
411
Unk
nown[G
lycinem
ax]
11
57JK973693
246
Phosph
atidylcholinetransferp
rotein
[Ricinu
scom
mun
is]5119890minus09
58JK973694
246
Phosph
atidylcholinetransferp
rotein
[Ricinu
scom
mun
is]5119890minus09
59JK973695
420
Uncharacterized
protein[Zea
mays]
9119890minus83
60JK
973696
350
Uncharacterized
protein[Zea
mays]
5119890minus51
61JK
973697
244
Jmjcdo
main-containing
protein4-lik
e[Brachypodium
dista
chyon]
5119890minus27
62JK
973698
415
Os01g0678900[O
ryza
sativa]
8119890minus11
63JK
973699
282
D-3-Pho
spho
glycerated
ehydrogenase
[Ricinu
scom
mun
is]3119890minus51
64JK
973700
558
60Srib
osom
alprotein[Vitisv
inifera]
8119890minus65
65JK
973701
490
Nosignific
antm
atch
66JK
973702
556
Root
nodu
leextension[Pisu
msativ
um]
3119890minus07
67JK
973703
313
Coiled-coildo
main-containing
protein[Ricinu
scom
mun
is]2119890minus65
68JK
973704
311
Coiled-coildo
main-containing
protein[Ricinu
scom
mun
is]1119890minus64
69JK
973705
320
Hypothetic
alproteinMTR
4g076190
[Medica
gotru
ncatula]
2119890minus13
70JK
973706
509
Putativ
eS-adeno
sylm
ethion
ines
ynthetase[Ca
psicu
mannu
um]
9119890minus119
71JK
973707
460
Transm
embranee
mp24do
main-containing
protein10-like
[Brachypodium
dista
chyon]
2119890minus45
72JK
973708
482
60Srib
osom
alprotein[Zea
mays]
3119890minus52
73JK
973709
395
MLO
5-lik
eprotein
[Medica
gotru
ncatula]
2119890minus24
74JK
973710
480
Type
2metallothionein
[Prosopisjulifl
ora]
4119890minus28
75JK
973711
556
Root
nodu
leextension[Pisu
msativa]
3119890minus07
76JK
973712
174
Hypothetic
alprotein[O
ryza
sativa]
086
77JK
973713
278
Nosignific
antm
atch
78JK
973714
439
Nosignific
antm
atch
79JK
973715
439
Serin
ecarbo
xypeptidase-lik
e19-lik
e[Glycinem
ax]
2119890minus31
80JK
973716
551
Serin
ecarbo
xypeptidase-lik
e19-lik
e[Glycinem
ax]
9119890minus39
81JK
973717
347
Nosignific
antm
atch
82JK
973718
661
Gam
ma-glutam
ylhydrolase[Vitis
vinifer
a]1119890minus54
83JK
973719
499
Type
2metallothionein
[Prosopisjuliflora]
2119890minus30
6 International Journal of Plant Genomics
Table3Con
tinued
Slnum
ber
Gen
bank
accessionnu
mber
Leng
th(in
bp)
Hom
ologou
sprotein
119864-value
84JK
973720
463
Nod
ulin
mtn21eam
a-lik
etranspo
rter
protein[Arabidopsisthaliana
]2119890minus44
85JK
973721
200
RNArecogn
ition
motif-containing
protein[Arabidopsisthaliana
]4119890minus10
86JK
973722
280
Beta-glucosid
ase4
4-lik
e[Glycinem
ax]
3119890minus52
87JK
973723
424
V-type
proton
ATpase
21kD
Aproteolip
idsubu
nit[Medica
gotru
ncatula]
2119890minus29
88JK
973724
210
AP2
ERF
andB3
domain-containing
transcrip
tionfactor
[Arabidopsisthaliana
]4119890minus40
89JK
973725
540
Polyrib
onucleotiden
ucleotidyltransfe
rase
[Vitisv
inifera]
3119890minus07
90JK
973726
649
Methion
inea
minop
eptid
ase2
B-lik
e[Brachypodium
dista
chyon]
3119890minus40
91JK
973727
585
Hypothetic
alprotein[Vitisvinifera]
69
92JK
973728
487
Unk
nown[Lotus
japonica]
1
93JK
973729
558
60Srib
osom
alprotein[Vitisvinifera]
8119890minus65
94JK
973730
314
D-3-Pho
spho
glycerated
ehydrogenase
[Ricinu
scom
mun
is]1119890minus55
95JK
973731
350
EIN3-bind
ingF-bo
xprotein1-like[Brachypodium
dista
chyon]
8119890minus65
96JK
973732
499
DNA-
bind
ingproteinRA
V1[Zeamays]
4119890minus25
97JK
973733
239
NOT2
NOT3
NOT5
family
protein[O
ryza
sativa]
3119890minus62
98JK
973734
272
40Srib
osom
alproteinS24-2-lik
e[Brachypodium
dista
chyon]
2119890minus57
99JK
973735
559
Sign
alrecogn
ition
particlesubu
nit[Arabidopsis
thaliana
]9119890minus80
100
JK973736
793
Vesic
le-associatedmem
branep
rotein
727-lik
e[Glycinem
ax]
1119890minus111
101
JK973737
649
Methion
inea
minop
eptid
ase2
B[Brachypodium
dista
chyon]
1119890minus24
102
JK973738
210
Gam
ma-gliadinprecursor[Ricin
uscommun
is]7119890minus10
103
JK973739
553
Unk
nownproteinprod
uct[Glycinem
ax]
1119890minus04
104
JK973740
600
Glycine
dehydrogenase[Glycinem
ax]
4119890minus102
105
JK973741
572
Glucanendo
-13-beta-glucosidase-lik
eprotein
3-lik
e[Glycinem
ax]
2119890minus04
International Journal of Plant Genomics 7
the plasma membrane [17] Similarly numerous ESTs likeglycine dehydrogenase signal proteins root nodule exten-sions membrane proteins and 120573-glucosidase were observedin the cDNA library constructed from pigeonpea root tissueswhich is thought to play possible roles in plant metabolismgrowth and development
Apart from the knownESTs 6 EST transcripts (JK973668JK973685 JK973701 JK973714 JK973715 and JK973718)wereobserved as not having any significant match in NCBI data-base Along with that uncharacterized and hypothetical pro-teins were also observed in cDNA library which were thoughtto impart in the cardinal role in root tissues of pigeonpea
4 Conclusion
Our present investigation contains a precise repertoire oftranscripts associated with the various metabolic functionsin pigeonpea root These ESTs appear to be involved in mul-tiple metabolism pathways in the plantrsquos physiological andbiochemical processes In addition to known genes someESTs were unknown and uncharacterized whose functionalroles remain unclear and require further investigation infuture The root transcriptome characterized in this studymarkedly provides a unique resource for investigating thefunctional specificities of the root systemThese EST tagsmaybe useful for functional gene annotation analysis of splicesite variants and intronexon determination and evaluationof gene homologies or KEGG pathway confirmation
Abbreviations
ESTs Expressed sequence tagsRT PCR Reverse transcription polymerase chain
reactionSAMS S-Adenosyl methionine synthetase
Conflict of Interests
The authors declare that there is no conflict of interestsregarding the publication of this paper
References
[1] Y L Nene and V K Shiela ldquoPigeon pea geography and impor-tancerdquo inThePigeon Pea Y L Nene S H Hall and V K SheilaEds pp 1ndash14 CAB International Wellingford UK 1990
[2] M T Kassa R V Penmetsa N Carrasquilla-Garcia et alldquoGenetic patterns of domestication in pigeonpea (Cajanus cajan(L) Millsp) and wild Cajanus relativesrdquo PLoS ONE vol 7 no6 Article ID e39563 2012
[3] R K Varshney R V Penmetsa S Dutta et al ldquoPigeonpeagenomics initiative (PGI) an international effort to improvecrop productivity of pigeonpea (Cajanus cajan L)rdquo MolecularBreeding vol 26 no 3 pp 393ndash408 2010
[4] N L Raju B N Gnanesh P Lekha et al ldquoThe first set of ESTresource for gene discovery andmarker development in pigeon-pea (Cajanus cajan L)rdquo BMC Plant Biology vol 10 pp 45ndash512010
[5] S F Altschul T L Madden A A Schaffer et al ldquoGappedBLAST and PSI-BLAST a new generation of protein databasesearch programsrdquo Nucleic Acids Research vol 25 no 17 pp3389ndash3402 1997
[6] R Zhai Y Feng H Wang et al ldquoTranscriptome analysis of riceroot heterosis by RNA-Seqrdquo BMC Genomics vol 14 article 19pp 1ndash14 2013
[7] Y Jiang and M K Deyholos ldquoComprehensive transcriptionalprofiling of NaCl-stressed Arabidopsis roots reveals novelclasses of responsive genesrdquo BMC Plant Biology vol 6 article25 2006
[8] A M Lindroth P Saarikoski G Flygh et al ldquoTwo S-adenosyl-methionine synthetase-encoding genes differentially expressedduring adventitious root development in Pinus contortardquo PlantMolecular Biology vol 46 no 3 pp 335ndash346 2001
[9] P K Chiang R K Gordon J Tal et al ldquoS-adenosylmethionineandmethylationrdquoTheFASEB Journal vol 10 no 4 pp 471ndash4801996
[10] A M Shirley C M McMichael and C Chapple ldquoThe sng2mutant of Arabidopsis is defective in the gene encoding theserine carboxypeptidase-like protein sinapoylglucosecholinesinapoyltransferaserdquo Plant Journal vol 28 no 1 pp 83ndash942001
[11] C M Fraser M G Thompson A M Shirley et al ldquoRelatedArabidopsis serine carboxypeptidase-like sinapoylglucose acyl-transferases display distinct but overlapping substrate specifici-tiesrdquo Plant Physiology vol 144 no 4 pp 1986ndash1999 2007
[12] D Weier J Mittasch D Strack and C Milkowski ldquoThe genesBnSCT1 and BnSCT2 from Brassica napus encoding the finalenzyme of sinapine biosynthesis molecular characterizationand suppressionrdquo Planta vol 227 no 2 pp 375ndash385 2008
[13] W T Lowther and B W Matthews ldquoStructure and functionof the methionine aminopeptidasesrdquo Biochimica et BiophysicaActa vol 1477 no 1-2 pp 157ndash167 2000
[14] V S Reddy G S Ali and A S N Reddy ldquoGenes encodingcalmodulin-binding proteins in the Arabidopsis genomerdquo Jour-nal of Biological Chemistry vol 277 no 12 pp 9840ndash9852 2002
[15] S R Albright and R Tjian ldquoTAFs revisited more data revealnew twists and confirm old ideasrdquo Gene vol 242 no 1-2 pp1ndash13 2000
[16] J Kang J Park H Choi et al ldquoPlant ABC transportersrdquo TheArabidopsis Book vol 9 pp 1ndash25 2010
[17] K A Devor and J B Mudd ldquoStructural analysis of phosphat-idylcholine of plant tissuerdquo Journal of Lipid Research vol 12 no4 pp 396ndash402 1971
Submit your manuscripts athttpwwwhindawicom
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Anatomy Research International
PeptidesInternational Journal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporation httpwwwhindawicom
International Journal of
Volume 2014
Zoology
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Molecular Biology International
GenomicsInternational Journal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
The Scientific World JournalHindawi Publishing Corporation httpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
BioinformaticsAdvances in
Marine BiologyJournal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Signal TransductionJournal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
BioMed Research International
Evolutionary BiologyInternational Journal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Biochemistry Research International
ArchaeaHindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Genetics Research International
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Advances in
Virolog y
Hindawi Publishing Corporationhttpwwwhindawicom
Nucleic AcidsJournal of
Volume 2014
Stem CellsInternational
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Enzyme Research
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
International Journal of
Microbiology

2 International Journal of Plant Genomics
Table 1 Primer sequences of subset of ESTs for RT-PCR analysis
Accession number Putative function Optimum 119879119898(∘C) Primer sequence (51015840-31015840)
JK973671 S-Adenosylmethionine synthetase 61 F-AGAGGAAAT CGGT GCTGGTGR-GCAGCAATTTGGTCGTTGGT
JK973674 Phosphoglycerate kinase 59 F-TCCCGATCCCGATACCCTACR-CAGCACGCTTTTCAGCAGTT
JK973715 Serine carboxypeptidase 60 F-ACATGAAGCTCAGTGGAGGAGR-AGCCATGGCCTCCAATCTTC
JK973726 Methionine aminopeptidase 60 F-GGCAT TGAAAGTTGGGCAGGR-GATTGCAGCACCGACATCAC
(Promega Madison WI USA) The quantification of RNAwas verified by absorption ratio of OD
260280and by for-
maldehyde gel electrophoresis The first and second strandsof cDNA were synthesized using Clontech SMARTer PCRcDNA Synthesis Kit The cDNAs were purified by the Min-Elute PCR purification kit (Qiagen Valencia CA USA) andligated into a pGEM-T easy vector (Promega Madison WIUSA) Ligated plasmid DNAs were used for transformationinto competent E coli DH5120572 strain Positive clones wereselected on an ampicillinIPTGX-Gal LB plate PlasmidDNA from positive clones were isolated by using REAL96 plasmid isolation kit (Qiagen Netherlands) andpurified DNA was used for single-pass Sanger sequencingby using M13FR universal sequencing primers on ABIsequencing machine 3500XL Genetic analyzer All the ESTswere processed using VecScreen (httpwwwncbinlmnihgovVecScreenVecScreenhtml) to remove vector and clon-ing oligo sequences and various contaminants to trim a highquality region Based on the qualified sequences the pre-dicted amino acid sequences were used to search for similarpeptide sequences to search for similar protein sequences inpublic database NCBI (httpwwwncbinlmnihgov) usingthe BLASTx search algorithm [5] by using default parametersof the program The similarity scores between the cDNAclones and known sequences were represented by BLASTxprobability E values Further the ESTs were classified intodifferent functional categories based on the knowledgeof biochemistry plant physiology and molecular biology(httpwwwMetaCycorg) GO (httpwwwebiacuk)and COG (httpwwwncbinlmnihgovCOG) tools andby searching related abstracts in PubMed
21 RT-PCR Analysis Total root RNAs isolated from pigeon-pea root tissues were used for reverse transcription poly-merase chain reaction (RT-PCR) analysis Genomic DNAcontamination was removed by DNase I First-strand cDNAwas synthesized from each 2120583g of total RNA sample usingClontech SMARTer PCR cDNA Synthesis Kit according tothe manufacturerrsquos protocol The cDNAs were purified usinga commercial column (Qiagen) To determine the expressionof candidate genes PCR was performed with 2120583L of thefirst-strand cDNA template and gene-specific primer pairsGene-specific RT-PCR primers were designed with Primer30 according to the EST sequences and were synthesizedcommercially General PCR was conducted with annealing
as required for the specific primer pairs (Table 1) RT-PCRexperimentswere repeated three times and the PCRproductswere detected on 15 agarose gel
3 Results and Discussion
Plant root systems serve a number of important functionsincluding anchoring the plant absorbingwater and nutrientsproducing amino acids and hormones and secreting organicacids enzymes and alkaloids [6] The physiological signifi-cance of roots is belied by their relative structural simplicityas compared to other plant organs major metabolic path-ways such as photosynthesis lacking in root tissues have astereotypical morphology that is conserved across taxa andthroughout the life cycle of individuals This combination ofphysiological relevance and structural simplicity has maderoots obvious targets for functional genomics analyses [7]As a major grain legume of semiarid tropics and a deep andextensive root system of pigeonpea represents an excellentsource of identification of ESTs associated with their roottissues So the presentworkwas focused on the study of genesassociated with pigeonpea root tissues
In the present investigation the cDNA library was con-structed in order to identify ESTs associated with pigeonpearoots and their functional analysis was carried out Thetotal RNA from the pigeonpea root tissue was isolatedand the first and second cDNA strand was synthesizedThe presence of the gene in plasmid construct of colonieswas confirmed by colony PCR The colony PCR showedthat the size of these inserts ranged from 200 to 800 bpOut of 400 bacterial clones the plasmid construct of 250positive recombinant clones was sequenced in single passedsequencing reaction from 31015840 end using M13 forwardreverseprimer and the sequence data was subjected to BLASTanalysis The leading sequences tailing of the sequence andpoor quality sequences were excluded firstly Finally 105 highquality ESTs were retained which were clustered into 72unigenes comprising 25 contigs and 47 singlets (Table 2) andwere compared with NCBI nonredundant protein databaseusing BLASTx algorithm and default parameters In BLASTxanalysis it was shown that most of the sequences were havinga significant homology with known proteins Sequences thathad no significant homology with protein database werecompared to nucleotide BLAST using default parameters
International Journal of Plant Genomics 3
Table 2 Summary of ESTs library
Total clone sequenced 250ESTs taken for analysis 105Number of unigenes 72Number of contigs 25Number of singlets 47Average length of unigenes 442 bpAverage length of ESTs 437 bp GC content of unigenes 503 GC content of ESTs 512
The ESTs were deposited to NCBI dbEST with the accessionnumber of JK973637 to JK973741 (Table 3)
TheESTswere categorized into 8 diverse functional groupconsisting of 10 transporter genes 6 signal transductiongenes 20 cell growth and transcriptional regulator genesand 26 metabolism genes In other genes 8 genes wereuncharacterized 8 hypothetical genes 10 genes with nosignificant match and 14 genes was involved in otherfunctionsThere were 8 of the ESTs found that did not showany match to known proteins in BLASTx program and thatsuggest novel nature of those genes (Figure 1)
In order to validate the ESTs generated from cDNAlibrary the expression of the four genes which were involvedin different metabolic pathways was analysed by RT-PCRRT-PCR results showed that the expression levels of fourcandidate genes namely S-adenosylmethionine synthetasephosphoglycerate kinase serine carboxypeptidase andmethio-nine aminopeptidase were clearly expressed in pigeonpearoots (Figure 2) It was concluded that overall there was agood agreement between the cDNA library data and the RT-PCR results
S-Adenosylmethionine synthetase (SAMS) comprises oftwo cDNAs in Pinus contorta among which SAMS1 isexpressed in roots and exhibits a specific expression patternin the meristem at the onset of adventitious root devel-opment [8] SAMS also catalyses the nucleophilic substi-tution reaction from between methionine and ATP intoS-adenosylmethionine which have central role in severalbiological process in plants namely methyl group donor intrimethylation of lignin DNA and alkaloids as well as donorof aminopropylmoieties in ethylene and polyamine synthesis[8 9]
Phosphoglycerate kinase superfamily has a diverse func-tion in numerous metabolic processes like generation ofprecursor metabolites and energy carbohydrate metabolismphosphorous metabolism glycolysis kinase activity ATPbinding activity and so forth The presence of phosphoglycer-ate kinase transcripts in the cDNA library supports its diversefunction in pigeonpea root
Serine carboxypeptidase differentially expressed in rootand other tissues is responsible for the synthesis of sinapoyl-choline and sinapoylmalate in Arabidopsis which encodes 51proteins annotated as serine carboxypeptidase-like enzymesand emerged as a new group of acyltransferase enzymesthat are able to modify plant natural products [10ndash12]
Hyp protein8
Transpoter10
Signaltransduction
6
Cell wall synthesis and transcriptional
regulator20
Metabolism26
Uncharacterized protein
8
Other functions14
No hits8
Figure 1 Functional classification of ESTs of pigeonpea generatedby single pass sequencing
M 4321
Figure 2 Validation of cDNA library results by RT-PCR (M-100 bp ladder 1-S-adenosylmethionine synthetase 2-phosphoglyceratekinase 3-serine carboxypeptidase and 4-methionine aminopepti-dase)
Methionine aminopeptidase a ubiquitous enzyme differen-tially expressed in root tissues is one of the central enzymes inprotein synthesis that catalyzes N-terminal methionine fromproteins [13]
A few important ESTs viz type 2 metallothionein TBP-associated factor ABC transporter and phosphatidylcholinetransfer protein were also abundantly found in the cDNAlibrary Reddy et al [14] found abundant metallothioneingenes in normalized library from rice leaves and suggestedthat they might perform essential functions of plant growthbeside metal detoxification TATA-binding protein and TBP-associated factors are transcriptional factors which are pre-dominantly involved in RNA polymerase II mediated tran-scription process [15] ABC transporters play an importantrole in organ growth plant nutrition plant developmentresponse to abiotic stress and the interaction of the plantwith its environment [16] Phosphatidylcholine is usuallythe most abundant phospholipids in animals and plantsoften amounting to almost 50 of the total and as such itis obviously the key building block of membrane bilayersmaking up a very high proportion of the outer leaflet of
4 International Journal of Plant Genomics
Table3Hom
olog
ysearch
oftheE
STsg
enerated
from
pigeon
pear
ootalong
with
their119864
valueu
singBL
AST
xprogramme
Slnum
ber
Gen
bank
accessionnu
mber
Leng
th(in
bp)
Hom
ologou
sprotein
119864-value
1JK
973637
560
Vesic
le-associatedmem
branep
rotein
727-lik
e[Glycinem
ax]
1119890minus111
2JK
973638
231
Mito
chon
drialinn
ermem
branep
rotein
OXA1-like[
Vitis
vinifer
a]5119890minus41
3JK
973639
630
Uncharacterized
protein[Arabidopsisthaliana
]9119890minus159
4JK
973640
621
Hypothetic
alprotein[Sorghum
bicolor]
9119890minus139
5JK
973641
570
GTP
-binding
proteinhfl
x[M
edica
gotru
ncatula]
6119890minus13
6JK
973642
412
Unk
nown[G
lycinem
ax]
11
7JK
973643
264
Secretorycarrier-associated
mem
branep
rotein
3-lik
e[Vitis
vinifer
a]2119890minus55
8JK
973644
438
Histon
e2[Populus
trichocarpa]
5119890minus63
9JK
973645
200
RNArecogn
ition
motif-containing
protein[Arabidopsisthaliana
]4119890minus10
10JK973646
870
Beta-glucosid
ase[
Arabidopsis
thaliana
]00
11JK
973647
555
Epsin
N-te
rminalho
molog
y(ENTH
)dom
ain-containing
protein[M
edica
gotru
ncatula]
4119890minus36
12JK
973648
210
Gam
ma-gliadinprecursor[Ricin
uscommun
is]7119890minus10
13JK
973649
210
AP2
ERF
andB3
domain-containing
transcrip
tionfactor
[Arabidopsisthaliana
]4119890minus40
14JK
973650
474
Glycine
dehydrogenase(decarborcylatingmito
chon
driallike)[G
lycinem
ax]
4119890minus102
15JK
973651
432
Sign
alrecogn
ition
particlesubu
nit[Arabidopsis
thaliana
]9119890minus80
16JK
973652
227
TBP-associated
factor
6B[Arabidopsisthaliana
]0051
17JK
973653
290
TBP-associated
factor
6B[Arabidopsisthaliana
]8119890minus48
18JK
973654
303
Transcrip
tionfactor
[Medica
gotru
ncatula]
6119890minus29
19JK
973655
432
Uncharacterized
protein[G
lycinem
ax]
8119890minus11
20JK
973656
237
Nuclear
cap-bind
ingproteinsubu
nit[Arabidopsis
thaliana
]8119890minus48
21JK
973657
501
Glucanendo
-13-beta-glucosidase-lik
eprotein
3-lik
e[Glycinem
ax]
2119890minus04
22JK
973658
499
DNA-
bind
ingproteinRA
V1[Zeamays]
4119890minus25
23JK
973659
560
Two-compo
nent
respon
seregulator
ARR
9[Vitisv
inifera]
1119890minus45
24JK
973660
210
Ribo
nucle
oside-diph
osph
ater
eductase
smallchain
like[Glycinem
ax]
2119890minus34
25JK
973661
748
Hypothetic
alprotein[O
ryza
sativa]
26JK
973662
742
Cyclin-depend
entp
rotein
kinase
complex
compo
nent
[Aspergillu
skaw
achii]
012
27JK
973663
406
Elon
gatio
nfactor
2[N
icotia
natobacum]
5119890minus39
28JK
973664
841
Expressedprotein[O
ryza
sativa]
73
29JK
973665
385
Hypothetic
alprotein[Vitisv
inifera]
21
30JK
973666
544
ADP-rib
osylationfactor-like
8d[N
icotia
natobacum]
2119890minus105
31JK
973667
510
Jmjcdo
main-containing
protein4-lik
e[Brachypodium
dista
chyon]
6119890minus77
32JK
973668
630
Nosignific
antm
atch
33JK
973669
804
Gag-pro[Pisu
msativ
um]
9119890minus18
34JK
973670
320
HypotheticalproteinMTR
[Medica
gotru
ncatula]
2119890minus13
35JK
973671
509
Putativ
eS-adeno
sylm
ethion
ines
ynthetase[Ca
psicu
mannu
m]
6119890minus114
36JK
973672
371
ABC
transporterF
family
mem
ber1-like
[Brachypodium
dista
chyon]
4119890minus60
37JK
973673
513
Predictedprotein[H
ordeum
vulga
re]
2119890minus89
38JK973674
465
Phosph
oglyceratekinasecytosolic[Triticu
maestivum]
2119890minus97
39JK973675
465
Phosph
oglyceratekinasecytosolic[Triticu
maestivum]
2119890minus97
40JK
973676
457
Unk
nown[Pice
asitchensis]
097
41JK
973677
785
GTP
-binding
signalrecognitio
nparticleSR
P54[M
edica
gotru
ncatula]
7119890minus51
International Journal of Plant Genomics 5
Table3Con
tinued
Slnum
ber
Gen
bank
accessionnu
mber
Leng
th(in
bp)
Hom
ologou
sprotein
119864-value
42JK
973678
241
Sign
alrecogn
ition
particle54
kDAsubu
nitp
recursor
[Pisu
msativ
um]
0044
43JK
973679
475
ProteinSE
T-lik
e[Brachypodium
dista
chyon]
7119890minus33
44JK
973680
350
EIN3-bind
ingF-bo
xprotein[Brachypodium
dista
chyon]
8119890minus65
45JK
973681
389
Predictedprotein[H
ordeum
vulga
re]
4119890minus27
46JK
973682
272
40Srib
osom
alproteinS24-2-lik
e[Brachypodium
dista
chyon]
2119890minus57
47JK973683
210
Putativ
ecalcium
exchanger[Triticum
dicoccoides]
4119890minus27
48JK
973684
478
Thioredo
xin[M
edica
gotru
ncatula]
2119890minus75
49JK
973685
210
Nosignific
antm
atch
50JK
973686
553
Root
nodu
leextension[Pisu
msativa]
5119890minus04
51JK
973687
600
Sign
alrecogn
ition
particleprotein[O
ryza
sativa]
5119890minus08
52JK
973688
544
ADP-rib
osylationfactor-like
8d[N
icotia
natobacum]
2119890minus105
53JK
973689
395
MLO
-like
protein[M
edica
gotru
ncatula]
2119890minus24
54JK973690
250
Phosph
atidylcholinetransferp
rotein
[Ricinu
scom
mun
is]3119890minus09
55JK973691
215
Phosph
atidylcholinetransferp
rotein
[Ricinu
scom
mun
is]3119890minus09
56JK
973692
411
Unk
nown[G
lycinem
ax]
11
57JK973693
246
Phosph
atidylcholinetransferp
rotein
[Ricinu
scom
mun
is]5119890minus09
58JK973694
246
Phosph
atidylcholinetransferp
rotein
[Ricinu
scom
mun
is]5119890minus09
59JK973695
420
Uncharacterized
protein[Zea
mays]
9119890minus83
60JK
973696
350
Uncharacterized
protein[Zea
mays]
5119890minus51
61JK
973697
244
Jmjcdo
main-containing
protein4-lik
e[Brachypodium
dista
chyon]
5119890minus27
62JK
973698
415
Os01g0678900[O
ryza
sativa]
8119890minus11
63JK
973699
282
D-3-Pho
spho
glycerated
ehydrogenase
[Ricinu
scom
mun
is]3119890minus51
64JK
973700
558
60Srib
osom
alprotein[Vitisv
inifera]
8119890minus65
65JK
973701
490
Nosignific
antm
atch
66JK
973702
556
Root
nodu
leextension[Pisu
msativ
um]
3119890minus07
67JK
973703
313
Coiled-coildo
main-containing
protein[Ricinu
scom
mun
is]2119890minus65
68JK
973704
311
Coiled-coildo
main-containing
protein[Ricinu
scom
mun
is]1119890minus64
69JK
973705
320
Hypothetic
alproteinMTR
4g076190
[Medica
gotru
ncatula]
2119890minus13
70JK
973706
509
Putativ
eS-adeno
sylm
ethion
ines
ynthetase[Ca
psicu
mannu
um]
9119890minus119
71JK
973707
460
Transm
embranee
mp24do
main-containing
protein10-like
[Brachypodium
dista
chyon]
2119890minus45
72JK
973708
482
60Srib
osom
alprotein[Zea
mays]
3119890minus52
73JK
973709
395
MLO
5-lik
eprotein
[Medica
gotru
ncatula]
2119890minus24
74JK
973710
480
Type
2metallothionein
[Prosopisjulifl
ora]
4119890minus28
75JK
973711
556
Root
nodu
leextension[Pisu
msativa]
3119890minus07
76JK
973712
174
Hypothetic
alprotein[O
ryza
sativa]
086
77JK
973713
278
Nosignific
antm
atch
78JK
973714
439
Nosignific
antm
atch
79JK
973715
439
Serin
ecarbo
xypeptidase-lik
e19-lik
e[Glycinem
ax]
2119890minus31
80JK
973716
551
Serin
ecarbo
xypeptidase-lik
e19-lik
e[Glycinem
ax]
9119890minus39
81JK
973717
347
Nosignific
antm
atch
82JK
973718
661
Gam
ma-glutam
ylhydrolase[Vitis
vinifer
a]1119890minus54
83JK
973719
499
Type
2metallothionein
[Prosopisjuliflora]
2119890minus30
6 International Journal of Plant Genomics
Table3Con
tinued
Slnum
ber
Gen
bank
accessionnu
mber
Leng
th(in
bp)
Hom
ologou
sprotein
119864-value
84JK
973720
463
Nod
ulin
mtn21eam
a-lik
etranspo
rter
protein[Arabidopsisthaliana
]2119890minus44
85JK
973721
200
RNArecogn
ition
motif-containing
protein[Arabidopsisthaliana
]4119890minus10
86JK
973722
280
Beta-glucosid
ase4
4-lik
e[Glycinem
ax]
3119890minus52
87JK
973723
424
V-type
proton
ATpase
21kD
Aproteolip
idsubu
nit[Medica
gotru
ncatula]
2119890minus29
88JK
973724
210
AP2
ERF
andB3
domain-containing
transcrip
tionfactor
[Arabidopsisthaliana
]4119890minus40
89JK
973725
540
Polyrib
onucleotiden
ucleotidyltransfe
rase
[Vitisv
inifera]
3119890minus07
90JK
973726
649
Methion
inea
minop
eptid
ase2
B-lik
e[Brachypodium
dista
chyon]
3119890minus40
91JK
973727
585
Hypothetic
alprotein[Vitisvinifera]
69
92JK
973728
487
Unk
nown[Lotus
japonica]
1
93JK
973729
558
60Srib
osom
alprotein[Vitisvinifera]
8119890minus65
94JK
973730
314
D-3-Pho
spho
glycerated
ehydrogenase
[Ricinu
scom
mun
is]1119890minus55
95JK
973731
350
EIN3-bind
ingF-bo
xprotein1-like[Brachypodium
dista
chyon]
8119890minus65
96JK
973732
499
DNA-
bind
ingproteinRA
V1[Zeamays]
4119890minus25
97JK
973733
239
NOT2
NOT3
NOT5
family
protein[O
ryza
sativa]
3119890minus62
98JK
973734
272
40Srib
osom
alproteinS24-2-lik
e[Brachypodium
dista
chyon]
2119890minus57
99JK
973735
559
Sign
alrecogn
ition
particlesubu
nit[Arabidopsis
thaliana
]9119890minus80
100
JK973736
793
Vesic
le-associatedmem
branep
rotein
727-lik
e[Glycinem
ax]
1119890minus111
101
JK973737
649
Methion
inea
minop
eptid
ase2
B[Brachypodium
dista
chyon]
1119890minus24
102
JK973738
210
Gam
ma-gliadinprecursor[Ricin
uscommun
is]7119890minus10
103
JK973739
553
Unk
nownproteinprod
uct[Glycinem
ax]
1119890minus04
104
JK973740
600
Glycine
dehydrogenase[Glycinem
ax]
4119890minus102
105
JK973741
572
Glucanendo
-13-beta-glucosidase-lik
eprotein
3-lik
e[Glycinem
ax]
2119890minus04
International Journal of Plant Genomics 7
the plasma membrane [17] Similarly numerous ESTs likeglycine dehydrogenase signal proteins root nodule exten-sions membrane proteins and 120573-glucosidase were observedin the cDNA library constructed from pigeonpea root tissueswhich is thought to play possible roles in plant metabolismgrowth and development
Apart from the knownESTs 6 EST transcripts (JK973668JK973685 JK973701 JK973714 JK973715 and JK973718)wereobserved as not having any significant match in NCBI data-base Along with that uncharacterized and hypothetical pro-teins were also observed in cDNA library which were thoughtto impart in the cardinal role in root tissues of pigeonpea
4 Conclusion
Our present investigation contains a precise repertoire oftranscripts associated with the various metabolic functionsin pigeonpea root These ESTs appear to be involved in mul-tiple metabolism pathways in the plantrsquos physiological andbiochemical processes In addition to known genes someESTs were unknown and uncharacterized whose functionalroles remain unclear and require further investigation infuture The root transcriptome characterized in this studymarkedly provides a unique resource for investigating thefunctional specificities of the root systemThese EST tagsmaybe useful for functional gene annotation analysis of splicesite variants and intronexon determination and evaluationof gene homologies or KEGG pathway confirmation
Abbreviations
ESTs Expressed sequence tagsRT PCR Reverse transcription polymerase chain
reactionSAMS S-Adenosyl methionine synthetase
Conflict of Interests
The authors declare that there is no conflict of interestsregarding the publication of this paper
References
[1] Y L Nene and V K Shiela ldquoPigeon pea geography and impor-tancerdquo inThePigeon Pea Y L Nene S H Hall and V K SheilaEds pp 1ndash14 CAB International Wellingford UK 1990
[2] M T Kassa R V Penmetsa N Carrasquilla-Garcia et alldquoGenetic patterns of domestication in pigeonpea (Cajanus cajan(L) Millsp) and wild Cajanus relativesrdquo PLoS ONE vol 7 no6 Article ID e39563 2012
[3] R K Varshney R V Penmetsa S Dutta et al ldquoPigeonpeagenomics initiative (PGI) an international effort to improvecrop productivity of pigeonpea (Cajanus cajan L)rdquo MolecularBreeding vol 26 no 3 pp 393ndash408 2010
[4] N L Raju B N Gnanesh P Lekha et al ldquoThe first set of ESTresource for gene discovery andmarker development in pigeon-pea (Cajanus cajan L)rdquo BMC Plant Biology vol 10 pp 45ndash512010
[5] S F Altschul T L Madden A A Schaffer et al ldquoGappedBLAST and PSI-BLAST a new generation of protein databasesearch programsrdquo Nucleic Acids Research vol 25 no 17 pp3389ndash3402 1997
[6] R Zhai Y Feng H Wang et al ldquoTranscriptome analysis of riceroot heterosis by RNA-Seqrdquo BMC Genomics vol 14 article 19pp 1ndash14 2013
[7] Y Jiang and M K Deyholos ldquoComprehensive transcriptionalprofiling of NaCl-stressed Arabidopsis roots reveals novelclasses of responsive genesrdquo BMC Plant Biology vol 6 article25 2006
[8] A M Lindroth P Saarikoski G Flygh et al ldquoTwo S-adenosyl-methionine synthetase-encoding genes differentially expressedduring adventitious root development in Pinus contortardquo PlantMolecular Biology vol 46 no 3 pp 335ndash346 2001
[9] P K Chiang R K Gordon J Tal et al ldquoS-adenosylmethionineandmethylationrdquoTheFASEB Journal vol 10 no 4 pp 471ndash4801996
[10] A M Shirley C M McMichael and C Chapple ldquoThe sng2mutant of Arabidopsis is defective in the gene encoding theserine carboxypeptidase-like protein sinapoylglucosecholinesinapoyltransferaserdquo Plant Journal vol 28 no 1 pp 83ndash942001
[11] C M Fraser M G Thompson A M Shirley et al ldquoRelatedArabidopsis serine carboxypeptidase-like sinapoylglucose acyl-transferases display distinct but overlapping substrate specifici-tiesrdquo Plant Physiology vol 144 no 4 pp 1986ndash1999 2007
[12] D Weier J Mittasch D Strack and C Milkowski ldquoThe genesBnSCT1 and BnSCT2 from Brassica napus encoding the finalenzyme of sinapine biosynthesis molecular characterizationand suppressionrdquo Planta vol 227 no 2 pp 375ndash385 2008
[13] W T Lowther and B W Matthews ldquoStructure and functionof the methionine aminopeptidasesrdquo Biochimica et BiophysicaActa vol 1477 no 1-2 pp 157ndash167 2000
[14] V S Reddy G S Ali and A S N Reddy ldquoGenes encodingcalmodulin-binding proteins in the Arabidopsis genomerdquo Jour-nal of Biological Chemistry vol 277 no 12 pp 9840ndash9852 2002
[15] S R Albright and R Tjian ldquoTAFs revisited more data revealnew twists and confirm old ideasrdquo Gene vol 242 no 1-2 pp1ndash13 2000
[16] J Kang J Park H Choi et al ldquoPlant ABC transportersrdquo TheArabidopsis Book vol 9 pp 1ndash25 2010
[17] K A Devor and J B Mudd ldquoStructural analysis of phosphat-idylcholine of plant tissuerdquo Journal of Lipid Research vol 12 no4 pp 396ndash402 1971
Submit your manuscripts athttpwwwhindawicom
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Anatomy Research International
PeptidesInternational Journal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporation httpwwwhindawicom
International Journal of
Volume 2014
Zoology
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Molecular Biology International
GenomicsInternational Journal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
The Scientific World JournalHindawi Publishing Corporation httpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
BioinformaticsAdvances in
Marine BiologyJournal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Signal TransductionJournal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
BioMed Research International
Evolutionary BiologyInternational Journal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Biochemistry Research International
ArchaeaHindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Genetics Research International
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Advances in
Virolog y
Hindawi Publishing Corporationhttpwwwhindawicom
Nucleic AcidsJournal of
Volume 2014
Stem CellsInternational
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Enzyme Research
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
International Journal of
Microbiology

International Journal of Plant Genomics 3
Table 2 Summary of ESTs library
Total clone sequenced 250ESTs taken for analysis 105Number of unigenes 72Number of contigs 25Number of singlets 47Average length of unigenes 442 bpAverage length of ESTs 437 bp GC content of unigenes 503 GC content of ESTs 512
The ESTs were deposited to NCBI dbEST with the accessionnumber of JK973637 to JK973741 (Table 3)
TheESTswere categorized into 8 diverse functional groupconsisting of 10 transporter genes 6 signal transductiongenes 20 cell growth and transcriptional regulator genesand 26 metabolism genes In other genes 8 genes wereuncharacterized 8 hypothetical genes 10 genes with nosignificant match and 14 genes was involved in otherfunctionsThere were 8 of the ESTs found that did not showany match to known proteins in BLASTx program and thatsuggest novel nature of those genes (Figure 1)
In order to validate the ESTs generated from cDNAlibrary the expression of the four genes which were involvedin different metabolic pathways was analysed by RT-PCRRT-PCR results showed that the expression levels of fourcandidate genes namely S-adenosylmethionine synthetasephosphoglycerate kinase serine carboxypeptidase andmethio-nine aminopeptidase were clearly expressed in pigeonpearoots (Figure 2) It was concluded that overall there was agood agreement between the cDNA library data and the RT-PCR results
S-Adenosylmethionine synthetase (SAMS) comprises oftwo cDNAs in Pinus contorta among which SAMS1 isexpressed in roots and exhibits a specific expression patternin the meristem at the onset of adventitious root devel-opment [8] SAMS also catalyses the nucleophilic substi-tution reaction from between methionine and ATP intoS-adenosylmethionine which have central role in severalbiological process in plants namely methyl group donor intrimethylation of lignin DNA and alkaloids as well as donorof aminopropylmoieties in ethylene and polyamine synthesis[8 9]
Phosphoglycerate kinase superfamily has a diverse func-tion in numerous metabolic processes like generation ofprecursor metabolites and energy carbohydrate metabolismphosphorous metabolism glycolysis kinase activity ATPbinding activity and so forth The presence of phosphoglycer-ate kinase transcripts in the cDNA library supports its diversefunction in pigeonpea root
Serine carboxypeptidase differentially expressed in rootand other tissues is responsible for the synthesis of sinapoyl-choline and sinapoylmalate in Arabidopsis which encodes 51proteins annotated as serine carboxypeptidase-like enzymesand emerged as a new group of acyltransferase enzymesthat are able to modify plant natural products [10ndash12]
Hyp protein8
Transpoter10
Signaltransduction
6
Cell wall synthesis and transcriptional
regulator20
Metabolism26
Uncharacterized protein
8
Other functions14
No hits8
Figure 1 Functional classification of ESTs of pigeonpea generatedby single pass sequencing
M 4321
Figure 2 Validation of cDNA library results by RT-PCR (M-100 bp ladder 1-S-adenosylmethionine synthetase 2-phosphoglyceratekinase 3-serine carboxypeptidase and 4-methionine aminopepti-dase)
Methionine aminopeptidase a ubiquitous enzyme differen-tially expressed in root tissues is one of the central enzymes inprotein synthesis that catalyzes N-terminal methionine fromproteins [13]
A few important ESTs viz type 2 metallothionein TBP-associated factor ABC transporter and phosphatidylcholinetransfer protein were also abundantly found in the cDNAlibrary Reddy et al [14] found abundant metallothioneingenes in normalized library from rice leaves and suggestedthat they might perform essential functions of plant growthbeside metal detoxification TATA-binding protein and TBP-associated factors are transcriptional factors which are pre-dominantly involved in RNA polymerase II mediated tran-scription process [15] ABC transporters play an importantrole in organ growth plant nutrition plant developmentresponse to abiotic stress and the interaction of the plantwith its environment [16] Phosphatidylcholine is usuallythe most abundant phospholipids in animals and plantsoften amounting to almost 50 of the total and as such itis obviously the key building block of membrane bilayersmaking up a very high proportion of the outer leaflet of
4 International Journal of Plant Genomics
Table3Hom
olog
ysearch
oftheE
STsg
enerated
from
pigeon
pear
ootalong
with
their119864
valueu
singBL
AST
xprogramme
Slnum
ber
Gen
bank
accessionnu
mber
Leng
th(in
bp)
Hom
ologou
sprotein
119864-value
1JK
973637
560
Vesic
le-associatedmem
branep
rotein
727-lik
e[Glycinem
ax]
1119890minus111
2JK
973638
231
Mito
chon
drialinn
ermem
branep
rotein
OXA1-like[
Vitis
vinifer
a]5119890minus41
3JK
973639
630
Uncharacterized
protein[Arabidopsisthaliana
]9119890minus159
4JK
973640
621
Hypothetic
alprotein[Sorghum
bicolor]
9119890minus139
5JK
973641
570
GTP
-binding
proteinhfl
x[M
edica
gotru
ncatula]
6119890minus13
6JK
973642
412
Unk
nown[G
lycinem
ax]
11
7JK
973643
264
Secretorycarrier-associated
mem
branep
rotein
3-lik
e[Vitis
vinifer
a]2119890minus55
8JK
973644
438
Histon
e2[Populus
trichocarpa]
5119890minus63
9JK
973645
200
RNArecogn
ition
motif-containing
protein[Arabidopsisthaliana
]4119890minus10
10JK973646
870
Beta-glucosid
ase[
Arabidopsis
thaliana
]00
11JK
973647
555
Epsin
N-te
rminalho
molog
y(ENTH
)dom
ain-containing
protein[M
edica
gotru
ncatula]
4119890minus36
12JK
973648
210
Gam
ma-gliadinprecursor[Ricin
uscommun
is]7119890minus10
13JK
973649
210
AP2
ERF
andB3
domain-containing
transcrip
tionfactor
[Arabidopsisthaliana
]4119890minus40
14JK
973650
474
Glycine
dehydrogenase(decarborcylatingmito
chon
driallike)[G
lycinem
ax]
4119890minus102
15JK
973651
432
Sign
alrecogn
ition
particlesubu
nit[Arabidopsis
thaliana
]9119890minus80
16JK
973652
227
TBP-associated
factor
6B[Arabidopsisthaliana
]0051
17JK
973653
290
TBP-associated
factor
6B[Arabidopsisthaliana
]8119890minus48
18JK
973654
303
Transcrip
tionfactor
[Medica
gotru
ncatula]
6119890minus29
19JK
973655
432
Uncharacterized
protein[G
lycinem
ax]
8119890minus11
20JK
973656
237
Nuclear
cap-bind
ingproteinsubu
nit[Arabidopsis
thaliana
]8119890minus48
21JK
973657
501
Glucanendo
-13-beta-glucosidase-lik
eprotein
3-lik
e[Glycinem
ax]
2119890minus04
22JK
973658
499
DNA-
bind
ingproteinRA
V1[Zeamays]
4119890minus25
23JK
973659
560
Two-compo
nent
respon
seregulator
ARR
9[Vitisv
inifera]
1119890minus45
24JK
973660
210
Ribo
nucle
oside-diph
osph
ater
eductase
smallchain
like[Glycinem
ax]
2119890minus34
25JK
973661
748
Hypothetic
alprotein[O
ryza
sativa]
26JK
973662
742
Cyclin-depend
entp
rotein
kinase
complex
compo
nent
[Aspergillu
skaw
achii]
012
27JK
973663
406
Elon
gatio
nfactor
2[N
icotia
natobacum]
5119890minus39
28JK
973664
841
Expressedprotein[O
ryza
sativa]
73
29JK
973665
385
Hypothetic
alprotein[Vitisv
inifera]
21
30JK
973666
544
ADP-rib
osylationfactor-like
8d[N
icotia
natobacum]
2119890minus105
31JK
973667
510
Jmjcdo
main-containing
protein4-lik
e[Brachypodium
dista
chyon]
6119890minus77
32JK
973668
630
Nosignific
antm
atch
33JK
973669
804
Gag-pro[Pisu
msativ
um]
9119890minus18
34JK
973670
320
HypotheticalproteinMTR
[Medica
gotru
ncatula]
2119890minus13
35JK
973671
509
Putativ
eS-adeno
sylm
ethion
ines
ynthetase[Ca
psicu
mannu
m]
6119890minus114
36JK
973672
371
ABC
transporterF
family
mem
ber1-like
[Brachypodium
dista
chyon]
4119890minus60
37JK
973673
513
Predictedprotein[H
ordeum
vulga
re]
2119890minus89
38JK973674
465
Phosph
oglyceratekinasecytosolic[Triticu
maestivum]
2119890minus97
39JK973675
465
Phosph
oglyceratekinasecytosolic[Triticu
maestivum]
2119890minus97
40JK
973676
457
Unk
nown[Pice
asitchensis]
097
41JK
973677
785
GTP
-binding
signalrecognitio
nparticleSR
P54[M
edica
gotru
ncatula]
7119890minus51
International Journal of Plant Genomics 5
Table3Con
tinued
Slnum
ber
Gen
bank
accessionnu
mber
Leng
th(in
bp)
Hom
ologou
sprotein
119864-value
42JK
973678
241
Sign
alrecogn
ition
particle54
kDAsubu
nitp
recursor
[Pisu
msativ
um]
0044
43JK
973679
475
ProteinSE
T-lik
e[Brachypodium
dista
chyon]
7119890minus33
44JK
973680
350
EIN3-bind
ingF-bo
xprotein[Brachypodium
dista
chyon]
8119890minus65
45JK
973681
389
Predictedprotein[H
ordeum
vulga
re]
4119890minus27
46JK
973682
272
40Srib
osom
alproteinS24-2-lik
e[Brachypodium
dista
chyon]
2119890minus57
47JK973683
210
Putativ
ecalcium
exchanger[Triticum
dicoccoides]
4119890minus27
48JK
973684
478
Thioredo
xin[M
edica
gotru
ncatula]
2119890minus75
49JK
973685
210
Nosignific
antm
atch
50JK
973686
553
Root
nodu
leextension[Pisu
msativa]
5119890minus04
51JK
973687
600
Sign
alrecogn
ition
particleprotein[O
ryza
sativa]
5119890minus08
52JK
973688
544
ADP-rib
osylationfactor-like
8d[N
icotia
natobacum]
2119890minus105
53JK
973689
395
MLO
-like
protein[M
edica
gotru
ncatula]
2119890minus24
54JK973690
250
Phosph
atidylcholinetransferp
rotein
[Ricinu
scom
mun
is]3119890minus09
55JK973691
215
Phosph
atidylcholinetransferp
rotein
[Ricinu
scom
mun
is]3119890minus09
56JK
973692
411
Unk
nown[G
lycinem
ax]
11
57JK973693
246
Phosph
atidylcholinetransferp
rotein
[Ricinu
scom
mun
is]5119890minus09
58JK973694
246
Phosph
atidylcholinetransferp
rotein
[Ricinu
scom
mun
is]5119890minus09
59JK973695
420
Uncharacterized
protein[Zea
mays]
9119890minus83
60JK
973696
350
Uncharacterized
protein[Zea
mays]
5119890minus51
61JK
973697
244
Jmjcdo
main-containing
protein4-lik
e[Brachypodium
dista
chyon]
5119890minus27
62JK
973698
415
Os01g0678900[O
ryza
sativa]
8119890minus11
63JK
973699
282
D-3-Pho
spho
glycerated
ehydrogenase
[Ricinu
scom
mun
is]3119890minus51
64JK
973700
558
60Srib
osom
alprotein[Vitisv
inifera]
8119890minus65
65JK
973701
490
Nosignific
antm
atch
66JK
973702
556
Root
nodu
leextension[Pisu
msativ
um]
3119890minus07
67JK
973703
313
Coiled-coildo
main-containing
protein[Ricinu
scom
mun
is]2119890minus65
68JK
973704
311
Coiled-coildo
main-containing
protein[Ricinu
scom
mun
is]1119890minus64
69JK
973705
320
Hypothetic
alproteinMTR
4g076190
[Medica
gotru
ncatula]
2119890minus13
70JK
973706
509
Putativ
eS-adeno
sylm
ethion
ines
ynthetase[Ca
psicu
mannu
um]
9119890minus119
71JK
973707
460
Transm
embranee
mp24do
main-containing
protein10-like
[Brachypodium
dista
chyon]
2119890minus45
72JK
973708
482
60Srib
osom
alprotein[Zea
mays]
3119890minus52
73JK
973709
395
MLO
5-lik
eprotein
[Medica
gotru
ncatula]
2119890minus24
74JK
973710
480
Type
2metallothionein
[Prosopisjulifl
ora]
4119890minus28
75JK
973711
556
Root
nodu
leextension[Pisu
msativa]
3119890minus07
76JK
973712
174
Hypothetic
alprotein[O
ryza
sativa]
086
77JK
973713
278
Nosignific
antm
atch
78JK
973714
439
Nosignific
antm
atch
79JK
973715
439
Serin
ecarbo
xypeptidase-lik
e19-lik
e[Glycinem
ax]
2119890minus31
80JK
973716
551
Serin
ecarbo
xypeptidase-lik
e19-lik
e[Glycinem
ax]
9119890minus39
81JK
973717
347
Nosignific
antm
atch
82JK
973718
661
Gam
ma-glutam
ylhydrolase[Vitis
vinifer
a]1119890minus54
83JK
973719
499
Type
2metallothionein
[Prosopisjuliflora]
2119890minus30
6 International Journal of Plant Genomics
Table3Con
tinued
Slnum
ber
Gen
bank
accessionnu
mber
Leng
th(in
bp)
Hom
ologou
sprotein
119864-value
84JK
973720
463
Nod
ulin
mtn21eam
a-lik
etranspo
rter
protein[Arabidopsisthaliana
]2119890minus44
85JK
973721
200
RNArecogn
ition
motif-containing
protein[Arabidopsisthaliana
]4119890minus10
86JK
973722
280
Beta-glucosid
ase4
4-lik
e[Glycinem
ax]
3119890minus52
87JK
973723
424
V-type
proton
ATpase
21kD
Aproteolip
idsubu
nit[Medica
gotru
ncatula]
2119890minus29
88JK
973724
210
AP2
ERF
andB3
domain-containing
transcrip
tionfactor
[Arabidopsisthaliana
]4119890minus40
89JK
973725
540
Polyrib
onucleotiden
ucleotidyltransfe
rase
[Vitisv
inifera]
3119890minus07
90JK
973726
649
Methion
inea
minop
eptid
ase2
B-lik
e[Brachypodium
dista
chyon]
3119890minus40
91JK
973727
585
Hypothetic
alprotein[Vitisvinifera]
69
92JK
973728
487
Unk
nown[Lotus
japonica]
1
93JK
973729
558
60Srib
osom
alprotein[Vitisvinifera]
8119890minus65
94JK
973730
314
D-3-Pho
spho
glycerated
ehydrogenase
[Ricinu
scom
mun
is]1119890minus55
95JK
973731
350
EIN3-bind
ingF-bo
xprotein1-like[Brachypodium
dista
chyon]
8119890minus65
96JK
973732
499
DNA-
bind
ingproteinRA
V1[Zeamays]
4119890minus25
97JK
973733
239
NOT2
NOT3
NOT5
family
protein[O
ryza
sativa]
3119890minus62
98JK
973734
272
40Srib
osom
alproteinS24-2-lik
e[Brachypodium
dista
chyon]
2119890minus57
99JK
973735
559
Sign
alrecogn
ition
particlesubu
nit[Arabidopsis
thaliana
]9119890minus80
100
JK973736
793
Vesic
le-associatedmem
branep
rotein
727-lik
e[Glycinem
ax]
1119890minus111
101
JK973737
649
Methion
inea
minop
eptid
ase2
B[Brachypodium
dista
chyon]
1119890minus24
102
JK973738
210
Gam
ma-gliadinprecursor[Ricin
uscommun
is]7119890minus10
103
JK973739
553
Unk
nownproteinprod
uct[Glycinem
ax]
1119890minus04
104
JK973740
600
Glycine
dehydrogenase[Glycinem
ax]
4119890minus102
105
JK973741
572
Glucanendo
-13-beta-glucosidase-lik
eprotein
3-lik
e[Glycinem
ax]
2119890minus04
International Journal of Plant Genomics 7
the plasma membrane [17] Similarly numerous ESTs likeglycine dehydrogenase signal proteins root nodule exten-sions membrane proteins and 120573-glucosidase were observedin the cDNA library constructed from pigeonpea root tissueswhich is thought to play possible roles in plant metabolismgrowth and development
Apart from the knownESTs 6 EST transcripts (JK973668JK973685 JK973701 JK973714 JK973715 and JK973718)wereobserved as not having any significant match in NCBI data-base Along with that uncharacterized and hypothetical pro-teins were also observed in cDNA library which were thoughtto impart in the cardinal role in root tissues of pigeonpea
4 Conclusion
Our present investigation contains a precise repertoire oftranscripts associated with the various metabolic functionsin pigeonpea root These ESTs appear to be involved in mul-tiple metabolism pathways in the plantrsquos physiological andbiochemical processes In addition to known genes someESTs were unknown and uncharacterized whose functionalroles remain unclear and require further investigation infuture The root transcriptome characterized in this studymarkedly provides a unique resource for investigating thefunctional specificities of the root systemThese EST tagsmaybe useful for functional gene annotation analysis of splicesite variants and intronexon determination and evaluationof gene homologies or KEGG pathway confirmation
Abbreviations
ESTs Expressed sequence tagsRT PCR Reverse transcription polymerase chain
reactionSAMS S-Adenosyl methionine synthetase
Conflict of Interests
The authors declare that there is no conflict of interestsregarding the publication of this paper
References
[1] Y L Nene and V K Shiela ldquoPigeon pea geography and impor-tancerdquo inThePigeon Pea Y L Nene S H Hall and V K SheilaEds pp 1ndash14 CAB International Wellingford UK 1990
[2] M T Kassa R V Penmetsa N Carrasquilla-Garcia et alldquoGenetic patterns of domestication in pigeonpea (Cajanus cajan(L) Millsp) and wild Cajanus relativesrdquo PLoS ONE vol 7 no6 Article ID e39563 2012
[3] R K Varshney R V Penmetsa S Dutta et al ldquoPigeonpeagenomics initiative (PGI) an international effort to improvecrop productivity of pigeonpea (Cajanus cajan L)rdquo MolecularBreeding vol 26 no 3 pp 393ndash408 2010
[4] N L Raju B N Gnanesh P Lekha et al ldquoThe first set of ESTresource for gene discovery andmarker development in pigeon-pea (Cajanus cajan L)rdquo BMC Plant Biology vol 10 pp 45ndash512010
[5] S F Altschul T L Madden A A Schaffer et al ldquoGappedBLAST and PSI-BLAST a new generation of protein databasesearch programsrdquo Nucleic Acids Research vol 25 no 17 pp3389ndash3402 1997
[6] R Zhai Y Feng H Wang et al ldquoTranscriptome analysis of riceroot heterosis by RNA-Seqrdquo BMC Genomics vol 14 article 19pp 1ndash14 2013
[7] Y Jiang and M K Deyholos ldquoComprehensive transcriptionalprofiling of NaCl-stressed Arabidopsis roots reveals novelclasses of responsive genesrdquo BMC Plant Biology vol 6 article25 2006
[8] A M Lindroth P Saarikoski G Flygh et al ldquoTwo S-adenosyl-methionine synthetase-encoding genes differentially expressedduring adventitious root development in Pinus contortardquo PlantMolecular Biology vol 46 no 3 pp 335ndash346 2001
[9] P K Chiang R K Gordon J Tal et al ldquoS-adenosylmethionineandmethylationrdquoTheFASEB Journal vol 10 no 4 pp 471ndash4801996
[10] A M Shirley C M McMichael and C Chapple ldquoThe sng2mutant of Arabidopsis is defective in the gene encoding theserine carboxypeptidase-like protein sinapoylglucosecholinesinapoyltransferaserdquo Plant Journal vol 28 no 1 pp 83ndash942001
[11] C M Fraser M G Thompson A M Shirley et al ldquoRelatedArabidopsis serine carboxypeptidase-like sinapoylglucose acyl-transferases display distinct but overlapping substrate specifici-tiesrdquo Plant Physiology vol 144 no 4 pp 1986ndash1999 2007
[12] D Weier J Mittasch D Strack and C Milkowski ldquoThe genesBnSCT1 and BnSCT2 from Brassica napus encoding the finalenzyme of sinapine biosynthesis molecular characterizationand suppressionrdquo Planta vol 227 no 2 pp 375ndash385 2008
[13] W T Lowther and B W Matthews ldquoStructure and functionof the methionine aminopeptidasesrdquo Biochimica et BiophysicaActa vol 1477 no 1-2 pp 157ndash167 2000
[14] V S Reddy G S Ali and A S N Reddy ldquoGenes encodingcalmodulin-binding proteins in the Arabidopsis genomerdquo Jour-nal of Biological Chemistry vol 277 no 12 pp 9840ndash9852 2002
[15] S R Albright and R Tjian ldquoTAFs revisited more data revealnew twists and confirm old ideasrdquo Gene vol 242 no 1-2 pp1ndash13 2000
[16] J Kang J Park H Choi et al ldquoPlant ABC transportersrdquo TheArabidopsis Book vol 9 pp 1ndash25 2010
[17] K A Devor and J B Mudd ldquoStructural analysis of phosphat-idylcholine of plant tissuerdquo Journal of Lipid Research vol 12 no4 pp 396ndash402 1971
Submit your manuscripts athttpwwwhindawicom
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Anatomy Research International
PeptidesInternational Journal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporation httpwwwhindawicom
International Journal of
Volume 2014
Zoology
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Molecular Biology International
GenomicsInternational Journal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
The Scientific World JournalHindawi Publishing Corporation httpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
BioinformaticsAdvances in
Marine BiologyJournal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Signal TransductionJournal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
BioMed Research International
Evolutionary BiologyInternational Journal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Biochemistry Research International
ArchaeaHindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Genetics Research International
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Advances in
Virolog y
Hindawi Publishing Corporationhttpwwwhindawicom
Nucleic AcidsJournal of
Volume 2014
Stem CellsInternational
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Enzyme Research
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
International Journal of
Microbiology

4 International Journal of Plant Genomics
Table3Hom
olog
ysearch
oftheE
STsg
enerated
from
pigeon
pear
ootalong
with
their119864
valueu
singBL
AST
xprogramme
Slnum
ber
Gen
bank
accessionnu
mber
Leng
th(in
bp)
Hom
ologou
sprotein
119864-value
1JK
973637
560
Vesic
le-associatedmem
branep
rotein
727-lik
e[Glycinem
ax]
1119890minus111
2JK
973638
231
Mito
chon
drialinn
ermem
branep
rotein
OXA1-like[
Vitis
vinifer
a]5119890minus41
3JK
973639
630
Uncharacterized
protein[Arabidopsisthaliana
]9119890minus159
4JK
973640
621
Hypothetic
alprotein[Sorghum
bicolor]
9119890minus139
5JK
973641
570
GTP
-binding
proteinhfl
x[M
edica
gotru
ncatula]
6119890minus13
6JK
973642
412
Unk
nown[G
lycinem
ax]
11
7JK
973643
264
Secretorycarrier-associated
mem
branep
rotein
3-lik
e[Vitis
vinifer
a]2119890minus55
8JK
973644
438
Histon
e2[Populus
trichocarpa]
5119890minus63
9JK
973645
200
RNArecogn
ition
motif-containing
protein[Arabidopsisthaliana
]4119890minus10
10JK973646
870
Beta-glucosid
ase[
Arabidopsis
thaliana
]00
11JK
973647
555
Epsin
N-te
rminalho
molog
y(ENTH
)dom
ain-containing
protein[M
edica
gotru
ncatula]
4119890minus36
12JK
973648
210
Gam
ma-gliadinprecursor[Ricin
uscommun
is]7119890minus10
13JK
973649
210
AP2
ERF
andB3
domain-containing
transcrip
tionfactor
[Arabidopsisthaliana
]4119890minus40
14JK
973650
474
Glycine
dehydrogenase(decarborcylatingmito
chon
driallike)[G
lycinem
ax]
4119890minus102
15JK
973651
432
Sign
alrecogn
ition
particlesubu
nit[Arabidopsis
thaliana
]9119890minus80
16JK
973652
227
TBP-associated
factor
6B[Arabidopsisthaliana
]0051
17JK
973653
290
TBP-associated
factor
6B[Arabidopsisthaliana
]8119890minus48
18JK
973654
303
Transcrip
tionfactor
[Medica
gotru
ncatula]
6119890minus29
19JK
973655
432
Uncharacterized
protein[G
lycinem
ax]
8119890minus11
20JK
973656
237
Nuclear
cap-bind
ingproteinsubu
nit[Arabidopsis
thaliana
]8119890minus48
21JK
973657
501
Glucanendo
-13-beta-glucosidase-lik
eprotein
3-lik
e[Glycinem
ax]
2119890minus04
22JK
973658
499
DNA-
bind
ingproteinRA
V1[Zeamays]
4119890minus25
23JK
973659
560
Two-compo
nent
respon
seregulator
ARR
9[Vitisv
inifera]
1119890minus45
24JK
973660
210
Ribo
nucle
oside-diph
osph
ater
eductase
smallchain
like[Glycinem
ax]
2119890minus34
25JK
973661
748
Hypothetic
alprotein[O
ryza
sativa]
26JK
973662
742
Cyclin-depend
entp
rotein
kinase
complex
compo
nent
[Aspergillu
skaw
achii]
012
27JK
973663
406
Elon
gatio
nfactor
2[N
icotia
natobacum]
5119890minus39
28JK
973664
841
Expressedprotein[O
ryza
sativa]
73
29JK
973665
385
Hypothetic
alprotein[Vitisv
inifera]
21
30JK
973666
544
ADP-rib
osylationfactor-like
8d[N
icotia
natobacum]
2119890minus105
31JK
973667
510
Jmjcdo
main-containing
protein4-lik
e[Brachypodium
dista
chyon]
6119890minus77
32JK
973668
630
Nosignific
antm
atch
33JK
973669
804
Gag-pro[Pisu
msativ
um]
9119890minus18
34JK
973670
320
HypotheticalproteinMTR
[Medica
gotru
ncatula]
2119890minus13
35JK
973671
509
Putativ
eS-adeno
sylm
ethion
ines
ynthetase[Ca
psicu
mannu
m]
6119890minus114
36JK
973672
371
ABC
transporterF
family
mem
ber1-like
[Brachypodium
dista
chyon]
4119890minus60
37JK
973673
513
Predictedprotein[H
ordeum
vulga
re]
2119890minus89
38JK973674
465
Phosph
oglyceratekinasecytosolic[Triticu
maestivum]
2119890minus97
39JK973675
465
Phosph
oglyceratekinasecytosolic[Triticu
maestivum]
2119890minus97
40JK
973676
457
Unk
nown[Pice
asitchensis]
097
41JK
973677
785
GTP
-binding
signalrecognitio
nparticleSR
P54[M
edica
gotru
ncatula]
7119890minus51
International Journal of Plant Genomics 5
Table3Con
tinued
Slnum
ber
Gen
bank
accessionnu
mber
Leng
th(in
bp)
Hom
ologou
sprotein
119864-value
42JK
973678
241
Sign
alrecogn
ition
particle54
kDAsubu
nitp
recursor
[Pisu
msativ
um]
0044
43JK
973679
475
ProteinSE
T-lik
e[Brachypodium
dista
chyon]
7119890minus33
44JK
973680
350
EIN3-bind
ingF-bo
xprotein[Brachypodium
dista
chyon]
8119890minus65
45JK
973681
389
Predictedprotein[H
ordeum
vulga
re]
4119890minus27
46JK
973682
272
40Srib
osom
alproteinS24-2-lik
e[Brachypodium
dista
chyon]
2119890minus57
47JK973683
210
Putativ
ecalcium
exchanger[Triticum
dicoccoides]
4119890minus27
48JK
973684
478
Thioredo
xin[M
edica
gotru
ncatula]
2119890minus75
49JK
973685
210
Nosignific
antm
atch
50JK
973686
553
Root
nodu
leextension[Pisu
msativa]
5119890minus04
51JK
973687
600
Sign
alrecogn
ition
particleprotein[O
ryza
sativa]
5119890minus08
52JK
973688
544
ADP-rib
osylationfactor-like
8d[N
icotia
natobacum]
2119890minus105
53JK
973689
395
MLO
-like
protein[M
edica
gotru
ncatula]
2119890minus24
54JK973690
250
Phosph
atidylcholinetransferp
rotein
[Ricinu
scom
mun
is]3119890minus09
55JK973691
215
Phosph
atidylcholinetransferp
rotein
[Ricinu
scom
mun
is]3119890minus09
56JK
973692
411
Unk
nown[G
lycinem
ax]
11
57JK973693
246
Phosph
atidylcholinetransferp
rotein
[Ricinu
scom
mun
is]5119890minus09
58JK973694
246
Phosph
atidylcholinetransferp
rotein
[Ricinu
scom
mun
is]5119890minus09
59JK973695
420
Uncharacterized
protein[Zea
mays]
9119890minus83
60JK
973696
350
Uncharacterized
protein[Zea
mays]
5119890minus51
61JK
973697
244
Jmjcdo
main-containing
protein4-lik
e[Brachypodium
dista
chyon]
5119890minus27
62JK
973698
415
Os01g0678900[O
ryza
sativa]
8119890minus11
63JK
973699
282
D-3-Pho
spho
glycerated
ehydrogenase
[Ricinu
scom
mun
is]3119890minus51
64JK
973700
558
60Srib
osom
alprotein[Vitisv
inifera]
8119890minus65
65JK
973701
490
Nosignific
antm
atch
66JK
973702
556
Root
nodu
leextension[Pisu
msativ
um]
3119890minus07
67JK
973703
313
Coiled-coildo
main-containing
protein[Ricinu
scom
mun
is]2119890minus65
68JK
973704
311
Coiled-coildo
main-containing
protein[Ricinu
scom
mun
is]1119890minus64
69JK
973705
320
Hypothetic
alproteinMTR
4g076190
[Medica
gotru
ncatula]
2119890minus13
70JK
973706
509
Putativ
eS-adeno
sylm
ethion
ines
ynthetase[Ca
psicu
mannu
um]
9119890minus119
71JK
973707
460
Transm
embranee
mp24do
main-containing
protein10-like
[Brachypodium
dista
chyon]
2119890minus45
72JK
973708
482
60Srib
osom
alprotein[Zea
mays]
3119890minus52
73JK
973709
395
MLO
5-lik
eprotein
[Medica
gotru
ncatula]
2119890minus24
74JK
973710
480
Type
2metallothionein
[Prosopisjulifl
ora]
4119890minus28
75JK
973711
556
Root
nodu
leextension[Pisu
msativa]
3119890minus07
76JK
973712
174
Hypothetic
alprotein[O
ryza
sativa]
086
77JK
973713
278
Nosignific
antm
atch
78JK
973714
439
Nosignific
antm
atch
79JK
973715
439
Serin
ecarbo
xypeptidase-lik
e19-lik
e[Glycinem
ax]
2119890minus31
80JK
973716
551
Serin
ecarbo
xypeptidase-lik
e19-lik
e[Glycinem
ax]
9119890minus39
81JK
973717
347
Nosignific
antm
atch
82JK
973718
661
Gam
ma-glutam
ylhydrolase[Vitis
vinifer
a]1119890minus54
83JK
973719
499
Type
2metallothionein
[Prosopisjuliflora]
2119890minus30
6 International Journal of Plant Genomics
Table3Con
tinued
Slnum
ber
Gen
bank
accessionnu
mber
Leng
th(in
bp)
Hom
ologou
sprotein
119864-value
84JK
973720
463
Nod
ulin
mtn21eam
a-lik
etranspo
rter
protein[Arabidopsisthaliana
]2119890minus44
85JK
973721
200
RNArecogn
ition
motif-containing
protein[Arabidopsisthaliana
]4119890minus10
86JK
973722
280
Beta-glucosid
ase4
4-lik
e[Glycinem
ax]
3119890minus52
87JK
973723
424
V-type
proton
ATpase
21kD
Aproteolip
idsubu
nit[Medica
gotru
ncatula]
2119890minus29
88JK
973724
210
AP2
ERF
andB3
domain-containing
transcrip
tionfactor
[Arabidopsisthaliana
]4119890minus40
89JK
973725
540
Polyrib
onucleotiden
ucleotidyltransfe
rase
[Vitisv
inifera]
3119890minus07
90JK
973726
649
Methion
inea
minop
eptid
ase2
B-lik
e[Brachypodium
dista
chyon]
3119890minus40
91JK
973727
585
Hypothetic
alprotein[Vitisvinifera]
69
92JK
973728
487
Unk
nown[Lotus
japonica]
1
93JK
973729
558
60Srib
osom
alprotein[Vitisvinifera]
8119890minus65
94JK
973730
314
D-3-Pho
spho
glycerated
ehydrogenase
[Ricinu
scom
mun
is]1119890minus55
95JK
973731
350
EIN3-bind
ingF-bo
xprotein1-like[Brachypodium
dista
chyon]
8119890minus65
96JK
973732
499
DNA-
bind
ingproteinRA
V1[Zeamays]
4119890minus25
97JK
973733
239
NOT2
NOT3
NOT5
family
protein[O
ryza
sativa]
3119890minus62
98JK
973734
272
40Srib
osom
alproteinS24-2-lik
e[Brachypodium
dista
chyon]
2119890minus57
99JK
973735
559
Sign
alrecogn
ition
particlesubu
nit[Arabidopsis
thaliana
]9119890minus80
100
JK973736
793
Vesic
le-associatedmem
branep
rotein
727-lik
e[Glycinem
ax]
1119890minus111
101
JK973737
649
Methion
inea
minop
eptid
ase2
B[Brachypodium
dista
chyon]
1119890minus24
102
JK973738
210
Gam
ma-gliadinprecursor[Ricin
uscommun
is]7119890minus10
103
JK973739
553
Unk
nownproteinprod
uct[Glycinem
ax]
1119890minus04
104
JK973740
600
Glycine
dehydrogenase[Glycinem
ax]
4119890minus102
105
JK973741
572
Glucanendo
-13-beta-glucosidase-lik
eprotein
3-lik
e[Glycinem
ax]
2119890minus04
International Journal of Plant Genomics 7
the plasma membrane [17] Similarly numerous ESTs likeglycine dehydrogenase signal proteins root nodule exten-sions membrane proteins and 120573-glucosidase were observedin the cDNA library constructed from pigeonpea root tissueswhich is thought to play possible roles in plant metabolismgrowth and development
Apart from the knownESTs 6 EST transcripts (JK973668JK973685 JK973701 JK973714 JK973715 and JK973718)wereobserved as not having any significant match in NCBI data-base Along with that uncharacterized and hypothetical pro-teins were also observed in cDNA library which were thoughtto impart in the cardinal role in root tissues of pigeonpea
4 Conclusion
Our present investigation contains a precise repertoire oftranscripts associated with the various metabolic functionsin pigeonpea root These ESTs appear to be involved in mul-tiple metabolism pathways in the plantrsquos physiological andbiochemical processes In addition to known genes someESTs were unknown and uncharacterized whose functionalroles remain unclear and require further investigation infuture The root transcriptome characterized in this studymarkedly provides a unique resource for investigating thefunctional specificities of the root systemThese EST tagsmaybe useful for functional gene annotation analysis of splicesite variants and intronexon determination and evaluationof gene homologies or KEGG pathway confirmation
Abbreviations
ESTs Expressed sequence tagsRT PCR Reverse transcription polymerase chain
reactionSAMS S-Adenosyl methionine synthetase
Conflict of Interests
The authors declare that there is no conflict of interestsregarding the publication of this paper
References
[1] Y L Nene and V K Shiela ldquoPigeon pea geography and impor-tancerdquo inThePigeon Pea Y L Nene S H Hall and V K SheilaEds pp 1ndash14 CAB International Wellingford UK 1990
[2] M T Kassa R V Penmetsa N Carrasquilla-Garcia et alldquoGenetic patterns of domestication in pigeonpea (Cajanus cajan(L) Millsp) and wild Cajanus relativesrdquo PLoS ONE vol 7 no6 Article ID e39563 2012
[3] R K Varshney R V Penmetsa S Dutta et al ldquoPigeonpeagenomics initiative (PGI) an international effort to improvecrop productivity of pigeonpea (Cajanus cajan L)rdquo MolecularBreeding vol 26 no 3 pp 393ndash408 2010
[4] N L Raju B N Gnanesh P Lekha et al ldquoThe first set of ESTresource for gene discovery andmarker development in pigeon-pea (Cajanus cajan L)rdquo BMC Plant Biology vol 10 pp 45ndash512010
[5] S F Altschul T L Madden A A Schaffer et al ldquoGappedBLAST and PSI-BLAST a new generation of protein databasesearch programsrdquo Nucleic Acids Research vol 25 no 17 pp3389ndash3402 1997
[6] R Zhai Y Feng H Wang et al ldquoTranscriptome analysis of riceroot heterosis by RNA-Seqrdquo BMC Genomics vol 14 article 19pp 1ndash14 2013
[7] Y Jiang and M K Deyholos ldquoComprehensive transcriptionalprofiling of NaCl-stressed Arabidopsis roots reveals novelclasses of responsive genesrdquo BMC Plant Biology vol 6 article25 2006
[8] A M Lindroth P Saarikoski G Flygh et al ldquoTwo S-adenosyl-methionine synthetase-encoding genes differentially expressedduring adventitious root development in Pinus contortardquo PlantMolecular Biology vol 46 no 3 pp 335ndash346 2001
[9] P K Chiang R K Gordon J Tal et al ldquoS-adenosylmethionineandmethylationrdquoTheFASEB Journal vol 10 no 4 pp 471ndash4801996
[10] A M Shirley C M McMichael and C Chapple ldquoThe sng2mutant of Arabidopsis is defective in the gene encoding theserine carboxypeptidase-like protein sinapoylglucosecholinesinapoyltransferaserdquo Plant Journal vol 28 no 1 pp 83ndash942001
[11] C M Fraser M G Thompson A M Shirley et al ldquoRelatedArabidopsis serine carboxypeptidase-like sinapoylglucose acyl-transferases display distinct but overlapping substrate specifici-tiesrdquo Plant Physiology vol 144 no 4 pp 1986ndash1999 2007
[12] D Weier J Mittasch D Strack and C Milkowski ldquoThe genesBnSCT1 and BnSCT2 from Brassica napus encoding the finalenzyme of sinapine biosynthesis molecular characterizationand suppressionrdquo Planta vol 227 no 2 pp 375ndash385 2008
[13] W T Lowther and B W Matthews ldquoStructure and functionof the methionine aminopeptidasesrdquo Biochimica et BiophysicaActa vol 1477 no 1-2 pp 157ndash167 2000
[14] V S Reddy G S Ali and A S N Reddy ldquoGenes encodingcalmodulin-binding proteins in the Arabidopsis genomerdquo Jour-nal of Biological Chemistry vol 277 no 12 pp 9840ndash9852 2002
[15] S R Albright and R Tjian ldquoTAFs revisited more data revealnew twists and confirm old ideasrdquo Gene vol 242 no 1-2 pp1ndash13 2000
[16] J Kang J Park H Choi et al ldquoPlant ABC transportersrdquo TheArabidopsis Book vol 9 pp 1ndash25 2010
[17] K A Devor and J B Mudd ldquoStructural analysis of phosphat-idylcholine of plant tissuerdquo Journal of Lipid Research vol 12 no4 pp 396ndash402 1971
Submit your manuscripts athttpwwwhindawicom
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Anatomy Research International
PeptidesInternational Journal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporation httpwwwhindawicom
International Journal of
Volume 2014
Zoology
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Molecular Biology International
GenomicsInternational Journal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
The Scientific World JournalHindawi Publishing Corporation httpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
BioinformaticsAdvances in
Marine BiologyJournal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Signal TransductionJournal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
BioMed Research International
Evolutionary BiologyInternational Journal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Biochemistry Research International
ArchaeaHindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Genetics Research International
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Advances in
Virolog y
Hindawi Publishing Corporationhttpwwwhindawicom
Nucleic AcidsJournal of
Volume 2014
Stem CellsInternational
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Enzyme Research
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
International Journal of
Microbiology

International Journal of Plant Genomics 5
Table3Con
tinued
Slnum
ber
Gen
bank
accessionnu
mber
Leng
th(in
bp)
Hom
ologou
sprotein
119864-value
42JK
973678
241
Sign
alrecogn
ition
particle54
kDAsubu
nitp
recursor
[Pisu
msativ
um]
0044
43JK
973679
475
ProteinSE
T-lik
e[Brachypodium
dista
chyon]
7119890minus33
44JK
973680
350
EIN3-bind
ingF-bo
xprotein[Brachypodium
dista
chyon]
8119890minus65
45JK
973681
389
Predictedprotein[H
ordeum
vulga
re]
4119890minus27
46JK
973682
272
40Srib
osom
alproteinS24-2-lik
e[Brachypodium
dista
chyon]
2119890minus57
47JK973683
210
Putativ
ecalcium
exchanger[Triticum
dicoccoides]
4119890minus27
48JK
973684
478
Thioredo
xin[M
edica
gotru
ncatula]
2119890minus75
49JK
973685
210
Nosignific
antm
atch
50JK
973686
553
Root
nodu
leextension[Pisu
msativa]
5119890minus04
51JK
973687
600
Sign
alrecogn
ition
particleprotein[O
ryza
sativa]
5119890minus08
52JK
973688
544
ADP-rib
osylationfactor-like
8d[N
icotia
natobacum]
2119890minus105
53JK
973689
395
MLO
-like
protein[M
edica
gotru
ncatula]
2119890minus24
54JK973690
250
Phosph
atidylcholinetransferp
rotein
[Ricinu
scom
mun
is]3119890minus09
55JK973691
215
Phosph
atidylcholinetransferp
rotein
[Ricinu
scom
mun
is]3119890minus09
56JK
973692
411
Unk
nown[G
lycinem
ax]
11
57JK973693
246
Phosph
atidylcholinetransferp
rotein
[Ricinu
scom
mun
is]5119890minus09
58JK973694
246
Phosph
atidylcholinetransferp
rotein
[Ricinu
scom
mun
is]5119890minus09
59JK973695
420
Uncharacterized
protein[Zea
mays]
9119890minus83
60JK
973696
350
Uncharacterized
protein[Zea
mays]
5119890minus51
61JK
973697
244
Jmjcdo
main-containing
protein4-lik
e[Brachypodium
dista
chyon]
5119890minus27
62JK
973698
415
Os01g0678900[O
ryza
sativa]
8119890minus11
63JK
973699
282
D-3-Pho
spho
glycerated
ehydrogenase
[Ricinu
scom
mun
is]3119890minus51
64JK
973700
558
60Srib
osom
alprotein[Vitisv
inifera]
8119890minus65
65JK
973701
490
Nosignific
antm
atch
66JK
973702
556
Root
nodu
leextension[Pisu
msativ
um]
3119890minus07
67JK
973703
313
Coiled-coildo
main-containing
protein[Ricinu
scom
mun
is]2119890minus65
68JK
973704
311
Coiled-coildo
main-containing
protein[Ricinu
scom
mun
is]1119890minus64
69JK
973705
320
Hypothetic
alproteinMTR
4g076190
[Medica
gotru
ncatula]
2119890minus13
70JK
973706
509
Putativ
eS-adeno
sylm
ethion
ines
ynthetase[Ca
psicu
mannu
um]
9119890minus119
71JK
973707
460
Transm
embranee
mp24do
main-containing
protein10-like
[Brachypodium
dista
chyon]
2119890minus45
72JK
973708
482
60Srib
osom
alprotein[Zea
mays]
3119890minus52
73JK
973709
395
MLO
5-lik
eprotein
[Medica
gotru
ncatula]
2119890minus24
74JK
973710
480
Type
2metallothionein
[Prosopisjulifl
ora]
4119890minus28
75JK
973711
556
Root
nodu
leextension[Pisu
msativa]
3119890minus07
76JK
973712
174
Hypothetic
alprotein[O
ryza
sativa]
086
77JK
973713
278
Nosignific
antm
atch
78JK
973714
439
Nosignific
antm
atch
79JK
973715
439
Serin
ecarbo
xypeptidase-lik
e19-lik
e[Glycinem
ax]
2119890minus31
80JK
973716
551
Serin
ecarbo
xypeptidase-lik
e19-lik
e[Glycinem
ax]
9119890minus39
81JK
973717
347
Nosignific
antm
atch
82JK
973718
661
Gam
ma-glutam
ylhydrolase[Vitis
vinifer
a]1119890minus54
83JK
973719
499
Type
2metallothionein
[Prosopisjuliflora]
2119890minus30
6 International Journal of Plant Genomics
Table3Con
tinued
Slnum
ber
Gen
bank
accessionnu
mber
Leng
th(in
bp)
Hom
ologou
sprotein
119864-value
84JK
973720
463
Nod
ulin
mtn21eam
a-lik
etranspo
rter
protein[Arabidopsisthaliana
]2119890minus44
85JK
973721
200
RNArecogn
ition
motif-containing
protein[Arabidopsisthaliana
]4119890minus10
86JK
973722
280
Beta-glucosid
ase4
4-lik
e[Glycinem
ax]
3119890minus52
87JK
973723
424
V-type
proton
ATpase
21kD
Aproteolip
idsubu
nit[Medica
gotru
ncatula]
2119890minus29
88JK
973724
210
AP2
ERF
andB3
domain-containing
transcrip
tionfactor
[Arabidopsisthaliana
]4119890minus40
89JK
973725
540
Polyrib
onucleotiden
ucleotidyltransfe
rase
[Vitisv
inifera]
3119890minus07
90JK
973726
649
Methion
inea
minop
eptid
ase2
B-lik
e[Brachypodium
dista
chyon]
3119890minus40
91JK
973727
585
Hypothetic
alprotein[Vitisvinifera]
69
92JK
973728
487
Unk
nown[Lotus
japonica]
1
93JK
973729
558
60Srib
osom
alprotein[Vitisvinifera]
8119890minus65
94JK
973730
314
D-3-Pho
spho
glycerated
ehydrogenase
[Ricinu
scom
mun
is]1119890minus55
95JK
973731
350
EIN3-bind
ingF-bo
xprotein1-like[Brachypodium
dista
chyon]
8119890minus65
96JK
973732
499
DNA-
bind
ingproteinRA
V1[Zeamays]
4119890minus25
97JK
973733
239
NOT2
NOT3
NOT5
family
protein[O
ryza
sativa]
3119890minus62
98JK
973734
272
40Srib
osom
alproteinS24-2-lik
e[Brachypodium
dista
chyon]
2119890minus57
99JK
973735
559
Sign
alrecogn
ition
particlesubu
nit[Arabidopsis
thaliana
]9119890minus80
100
JK973736
793
Vesic
le-associatedmem
branep
rotein
727-lik
e[Glycinem
ax]
1119890minus111
101
JK973737
649
Methion
inea
minop
eptid
ase2
B[Brachypodium
dista
chyon]
1119890minus24
102
JK973738
210
Gam
ma-gliadinprecursor[Ricin
uscommun
is]7119890minus10
103
JK973739
553
Unk
nownproteinprod
uct[Glycinem
ax]
1119890minus04
104
JK973740
600
Glycine
dehydrogenase[Glycinem
ax]
4119890minus102
105
JK973741
572
Glucanendo
-13-beta-glucosidase-lik
eprotein
3-lik
e[Glycinem
ax]
2119890minus04
International Journal of Plant Genomics 7
the plasma membrane [17] Similarly numerous ESTs likeglycine dehydrogenase signal proteins root nodule exten-sions membrane proteins and 120573-glucosidase were observedin the cDNA library constructed from pigeonpea root tissueswhich is thought to play possible roles in plant metabolismgrowth and development
Apart from the knownESTs 6 EST transcripts (JK973668JK973685 JK973701 JK973714 JK973715 and JK973718)wereobserved as not having any significant match in NCBI data-base Along with that uncharacterized and hypothetical pro-teins were also observed in cDNA library which were thoughtto impart in the cardinal role in root tissues of pigeonpea
4 Conclusion
Our present investigation contains a precise repertoire oftranscripts associated with the various metabolic functionsin pigeonpea root These ESTs appear to be involved in mul-tiple metabolism pathways in the plantrsquos physiological andbiochemical processes In addition to known genes someESTs were unknown and uncharacterized whose functionalroles remain unclear and require further investigation infuture The root transcriptome characterized in this studymarkedly provides a unique resource for investigating thefunctional specificities of the root systemThese EST tagsmaybe useful for functional gene annotation analysis of splicesite variants and intronexon determination and evaluationof gene homologies or KEGG pathway confirmation
Abbreviations
ESTs Expressed sequence tagsRT PCR Reverse transcription polymerase chain
reactionSAMS S-Adenosyl methionine synthetase
Conflict of Interests
The authors declare that there is no conflict of interestsregarding the publication of this paper
References
[1] Y L Nene and V K Shiela ldquoPigeon pea geography and impor-tancerdquo inThePigeon Pea Y L Nene S H Hall and V K SheilaEds pp 1ndash14 CAB International Wellingford UK 1990
[2] M T Kassa R V Penmetsa N Carrasquilla-Garcia et alldquoGenetic patterns of domestication in pigeonpea (Cajanus cajan(L) Millsp) and wild Cajanus relativesrdquo PLoS ONE vol 7 no6 Article ID e39563 2012
[3] R K Varshney R V Penmetsa S Dutta et al ldquoPigeonpeagenomics initiative (PGI) an international effort to improvecrop productivity of pigeonpea (Cajanus cajan L)rdquo MolecularBreeding vol 26 no 3 pp 393ndash408 2010
[4] N L Raju B N Gnanesh P Lekha et al ldquoThe first set of ESTresource for gene discovery andmarker development in pigeon-pea (Cajanus cajan L)rdquo BMC Plant Biology vol 10 pp 45ndash512010
[5] S F Altschul T L Madden A A Schaffer et al ldquoGappedBLAST and PSI-BLAST a new generation of protein databasesearch programsrdquo Nucleic Acids Research vol 25 no 17 pp3389ndash3402 1997
[6] R Zhai Y Feng H Wang et al ldquoTranscriptome analysis of riceroot heterosis by RNA-Seqrdquo BMC Genomics vol 14 article 19pp 1ndash14 2013
[7] Y Jiang and M K Deyholos ldquoComprehensive transcriptionalprofiling of NaCl-stressed Arabidopsis roots reveals novelclasses of responsive genesrdquo BMC Plant Biology vol 6 article25 2006
[8] A M Lindroth P Saarikoski G Flygh et al ldquoTwo S-adenosyl-methionine synthetase-encoding genes differentially expressedduring adventitious root development in Pinus contortardquo PlantMolecular Biology vol 46 no 3 pp 335ndash346 2001
[9] P K Chiang R K Gordon J Tal et al ldquoS-adenosylmethionineandmethylationrdquoTheFASEB Journal vol 10 no 4 pp 471ndash4801996
[10] A M Shirley C M McMichael and C Chapple ldquoThe sng2mutant of Arabidopsis is defective in the gene encoding theserine carboxypeptidase-like protein sinapoylglucosecholinesinapoyltransferaserdquo Plant Journal vol 28 no 1 pp 83ndash942001
[11] C M Fraser M G Thompson A M Shirley et al ldquoRelatedArabidopsis serine carboxypeptidase-like sinapoylglucose acyl-transferases display distinct but overlapping substrate specifici-tiesrdquo Plant Physiology vol 144 no 4 pp 1986ndash1999 2007
[12] D Weier J Mittasch D Strack and C Milkowski ldquoThe genesBnSCT1 and BnSCT2 from Brassica napus encoding the finalenzyme of sinapine biosynthesis molecular characterizationand suppressionrdquo Planta vol 227 no 2 pp 375ndash385 2008
[13] W T Lowther and B W Matthews ldquoStructure and functionof the methionine aminopeptidasesrdquo Biochimica et BiophysicaActa vol 1477 no 1-2 pp 157ndash167 2000
[14] V S Reddy G S Ali and A S N Reddy ldquoGenes encodingcalmodulin-binding proteins in the Arabidopsis genomerdquo Jour-nal of Biological Chemistry vol 277 no 12 pp 9840ndash9852 2002
[15] S R Albright and R Tjian ldquoTAFs revisited more data revealnew twists and confirm old ideasrdquo Gene vol 242 no 1-2 pp1ndash13 2000
[16] J Kang J Park H Choi et al ldquoPlant ABC transportersrdquo TheArabidopsis Book vol 9 pp 1ndash25 2010
[17] K A Devor and J B Mudd ldquoStructural analysis of phosphat-idylcholine of plant tissuerdquo Journal of Lipid Research vol 12 no4 pp 396ndash402 1971
Submit your manuscripts athttpwwwhindawicom
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Anatomy Research International
PeptidesInternational Journal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporation httpwwwhindawicom
International Journal of
Volume 2014
Zoology
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Molecular Biology International
GenomicsInternational Journal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
The Scientific World JournalHindawi Publishing Corporation httpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
BioinformaticsAdvances in
Marine BiologyJournal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Signal TransductionJournal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
BioMed Research International
Evolutionary BiologyInternational Journal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Biochemistry Research International
ArchaeaHindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Genetics Research International
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Advances in
Virolog y
Hindawi Publishing Corporationhttpwwwhindawicom
Nucleic AcidsJournal of
Volume 2014
Stem CellsInternational
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Enzyme Research
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
International Journal of
Microbiology

6 International Journal of Plant Genomics
Table3Con
tinued
Slnum
ber
Gen
bank
accessionnu
mber
Leng
th(in
bp)
Hom
ologou
sprotein
119864-value
84JK
973720
463
Nod
ulin
mtn21eam
a-lik
etranspo
rter
protein[Arabidopsisthaliana
]2119890minus44
85JK
973721
200
RNArecogn
ition
motif-containing
protein[Arabidopsisthaliana
]4119890minus10
86JK
973722
280
Beta-glucosid
ase4
4-lik
e[Glycinem
ax]
3119890minus52
87JK
973723
424
V-type
proton
ATpase
21kD
Aproteolip
idsubu
nit[Medica
gotru
ncatula]
2119890minus29
88JK
973724
210
AP2
ERF
andB3
domain-containing
transcrip
tionfactor
[Arabidopsisthaliana
]4119890minus40
89JK
973725
540
Polyrib
onucleotiden
ucleotidyltransfe
rase
[Vitisv
inifera]
3119890minus07
90JK
973726
649
Methion
inea
minop
eptid
ase2
B-lik
e[Brachypodium
dista
chyon]
3119890minus40
91JK
973727
585
Hypothetic
alprotein[Vitisvinifera]
69
92JK
973728
487
Unk
nown[Lotus
japonica]
1
93JK
973729
558
60Srib
osom
alprotein[Vitisvinifera]
8119890minus65
94JK
973730
314
D-3-Pho
spho
glycerated
ehydrogenase
[Ricinu
scom
mun
is]1119890minus55
95JK
973731
350
EIN3-bind
ingF-bo
xprotein1-like[Brachypodium
dista
chyon]
8119890minus65
96JK
973732
499
DNA-
bind
ingproteinRA
V1[Zeamays]
4119890minus25
97JK
973733
239
NOT2
NOT3
NOT5
family
protein[O
ryza
sativa]
3119890minus62
98JK
973734
272
40Srib
osom
alproteinS24-2-lik
e[Brachypodium
dista
chyon]
2119890minus57
99JK
973735
559
Sign
alrecogn
ition
particlesubu
nit[Arabidopsis
thaliana
]9119890minus80
100
JK973736
793
Vesic
le-associatedmem
branep
rotein
727-lik
e[Glycinem
ax]
1119890minus111
101
JK973737
649
Methion
inea
minop
eptid
ase2
B[Brachypodium
dista
chyon]
1119890minus24
102
JK973738
210
Gam
ma-gliadinprecursor[Ricin
uscommun
is]7119890minus10
103
JK973739
553
Unk
nownproteinprod
uct[Glycinem
ax]
1119890minus04
104
JK973740
600
Glycine
dehydrogenase[Glycinem
ax]
4119890minus102
105
JK973741
572
Glucanendo
-13-beta-glucosidase-lik
eprotein
3-lik
e[Glycinem
ax]
2119890minus04
International Journal of Plant Genomics 7
the plasma membrane [17] Similarly numerous ESTs likeglycine dehydrogenase signal proteins root nodule exten-sions membrane proteins and 120573-glucosidase were observedin the cDNA library constructed from pigeonpea root tissueswhich is thought to play possible roles in plant metabolismgrowth and development
Apart from the knownESTs 6 EST transcripts (JK973668JK973685 JK973701 JK973714 JK973715 and JK973718)wereobserved as not having any significant match in NCBI data-base Along with that uncharacterized and hypothetical pro-teins were also observed in cDNA library which were thoughtto impart in the cardinal role in root tissues of pigeonpea
4 Conclusion
Our present investigation contains a precise repertoire oftranscripts associated with the various metabolic functionsin pigeonpea root These ESTs appear to be involved in mul-tiple metabolism pathways in the plantrsquos physiological andbiochemical processes In addition to known genes someESTs were unknown and uncharacterized whose functionalroles remain unclear and require further investigation infuture The root transcriptome characterized in this studymarkedly provides a unique resource for investigating thefunctional specificities of the root systemThese EST tagsmaybe useful for functional gene annotation analysis of splicesite variants and intronexon determination and evaluationof gene homologies or KEGG pathway confirmation
Abbreviations
ESTs Expressed sequence tagsRT PCR Reverse transcription polymerase chain
reactionSAMS S-Adenosyl methionine synthetase
Conflict of Interests
The authors declare that there is no conflict of interestsregarding the publication of this paper
References
[1] Y L Nene and V K Shiela ldquoPigeon pea geography and impor-tancerdquo inThePigeon Pea Y L Nene S H Hall and V K SheilaEds pp 1ndash14 CAB International Wellingford UK 1990
[2] M T Kassa R V Penmetsa N Carrasquilla-Garcia et alldquoGenetic patterns of domestication in pigeonpea (Cajanus cajan(L) Millsp) and wild Cajanus relativesrdquo PLoS ONE vol 7 no6 Article ID e39563 2012
[3] R K Varshney R V Penmetsa S Dutta et al ldquoPigeonpeagenomics initiative (PGI) an international effort to improvecrop productivity of pigeonpea (Cajanus cajan L)rdquo MolecularBreeding vol 26 no 3 pp 393ndash408 2010
[4] N L Raju B N Gnanesh P Lekha et al ldquoThe first set of ESTresource for gene discovery andmarker development in pigeon-pea (Cajanus cajan L)rdquo BMC Plant Biology vol 10 pp 45ndash512010
[5] S F Altschul T L Madden A A Schaffer et al ldquoGappedBLAST and PSI-BLAST a new generation of protein databasesearch programsrdquo Nucleic Acids Research vol 25 no 17 pp3389ndash3402 1997
[6] R Zhai Y Feng H Wang et al ldquoTranscriptome analysis of riceroot heterosis by RNA-Seqrdquo BMC Genomics vol 14 article 19pp 1ndash14 2013
[7] Y Jiang and M K Deyholos ldquoComprehensive transcriptionalprofiling of NaCl-stressed Arabidopsis roots reveals novelclasses of responsive genesrdquo BMC Plant Biology vol 6 article25 2006
[8] A M Lindroth P Saarikoski G Flygh et al ldquoTwo S-adenosyl-methionine synthetase-encoding genes differentially expressedduring adventitious root development in Pinus contortardquo PlantMolecular Biology vol 46 no 3 pp 335ndash346 2001
[9] P K Chiang R K Gordon J Tal et al ldquoS-adenosylmethionineandmethylationrdquoTheFASEB Journal vol 10 no 4 pp 471ndash4801996
[10] A M Shirley C M McMichael and C Chapple ldquoThe sng2mutant of Arabidopsis is defective in the gene encoding theserine carboxypeptidase-like protein sinapoylglucosecholinesinapoyltransferaserdquo Plant Journal vol 28 no 1 pp 83ndash942001
[11] C M Fraser M G Thompson A M Shirley et al ldquoRelatedArabidopsis serine carboxypeptidase-like sinapoylglucose acyl-transferases display distinct but overlapping substrate specifici-tiesrdquo Plant Physiology vol 144 no 4 pp 1986ndash1999 2007
[12] D Weier J Mittasch D Strack and C Milkowski ldquoThe genesBnSCT1 and BnSCT2 from Brassica napus encoding the finalenzyme of sinapine biosynthesis molecular characterizationand suppressionrdquo Planta vol 227 no 2 pp 375ndash385 2008
[13] W T Lowther and B W Matthews ldquoStructure and functionof the methionine aminopeptidasesrdquo Biochimica et BiophysicaActa vol 1477 no 1-2 pp 157ndash167 2000
[14] V S Reddy G S Ali and A S N Reddy ldquoGenes encodingcalmodulin-binding proteins in the Arabidopsis genomerdquo Jour-nal of Biological Chemistry vol 277 no 12 pp 9840ndash9852 2002
[15] S R Albright and R Tjian ldquoTAFs revisited more data revealnew twists and confirm old ideasrdquo Gene vol 242 no 1-2 pp1ndash13 2000
[16] J Kang J Park H Choi et al ldquoPlant ABC transportersrdquo TheArabidopsis Book vol 9 pp 1ndash25 2010
[17] K A Devor and J B Mudd ldquoStructural analysis of phosphat-idylcholine of plant tissuerdquo Journal of Lipid Research vol 12 no4 pp 396ndash402 1971
Submit your manuscripts athttpwwwhindawicom
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Anatomy Research International
PeptidesInternational Journal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporation httpwwwhindawicom
International Journal of
Volume 2014
Zoology
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Molecular Biology International
GenomicsInternational Journal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
The Scientific World JournalHindawi Publishing Corporation httpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
BioinformaticsAdvances in
Marine BiologyJournal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Signal TransductionJournal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
BioMed Research International
Evolutionary BiologyInternational Journal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Biochemistry Research International
ArchaeaHindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Genetics Research International
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Advances in
Virolog y
Hindawi Publishing Corporationhttpwwwhindawicom
Nucleic AcidsJournal of
Volume 2014
Stem CellsInternational
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Enzyme Research
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
International Journal of
Microbiology

International Journal of Plant Genomics 7
the plasma membrane [17] Similarly numerous ESTs likeglycine dehydrogenase signal proteins root nodule exten-sions membrane proteins and 120573-glucosidase were observedin the cDNA library constructed from pigeonpea root tissueswhich is thought to play possible roles in plant metabolismgrowth and development
Apart from the knownESTs 6 EST transcripts (JK973668JK973685 JK973701 JK973714 JK973715 and JK973718)wereobserved as not having any significant match in NCBI data-base Along with that uncharacterized and hypothetical pro-teins were also observed in cDNA library which were thoughtto impart in the cardinal role in root tissues of pigeonpea
4 Conclusion
Our present investigation contains a precise repertoire oftranscripts associated with the various metabolic functionsin pigeonpea root These ESTs appear to be involved in mul-tiple metabolism pathways in the plantrsquos physiological andbiochemical processes In addition to known genes someESTs were unknown and uncharacterized whose functionalroles remain unclear and require further investigation infuture The root transcriptome characterized in this studymarkedly provides a unique resource for investigating thefunctional specificities of the root systemThese EST tagsmaybe useful for functional gene annotation analysis of splicesite variants and intronexon determination and evaluationof gene homologies or KEGG pathway confirmation
Abbreviations
ESTs Expressed sequence tagsRT PCR Reverse transcription polymerase chain
reactionSAMS S-Adenosyl methionine synthetase
Conflict of Interests
The authors declare that there is no conflict of interestsregarding the publication of this paper
References
[1] Y L Nene and V K Shiela ldquoPigeon pea geography and impor-tancerdquo inThePigeon Pea Y L Nene S H Hall and V K SheilaEds pp 1ndash14 CAB International Wellingford UK 1990
[2] M T Kassa R V Penmetsa N Carrasquilla-Garcia et alldquoGenetic patterns of domestication in pigeonpea (Cajanus cajan(L) Millsp) and wild Cajanus relativesrdquo PLoS ONE vol 7 no6 Article ID e39563 2012
[3] R K Varshney R V Penmetsa S Dutta et al ldquoPigeonpeagenomics initiative (PGI) an international effort to improvecrop productivity of pigeonpea (Cajanus cajan L)rdquo MolecularBreeding vol 26 no 3 pp 393ndash408 2010
[4] N L Raju B N Gnanesh P Lekha et al ldquoThe first set of ESTresource for gene discovery andmarker development in pigeon-pea (Cajanus cajan L)rdquo BMC Plant Biology vol 10 pp 45ndash512010
[5] S F Altschul T L Madden A A Schaffer et al ldquoGappedBLAST and PSI-BLAST a new generation of protein databasesearch programsrdquo Nucleic Acids Research vol 25 no 17 pp3389ndash3402 1997
[6] R Zhai Y Feng H Wang et al ldquoTranscriptome analysis of riceroot heterosis by RNA-Seqrdquo BMC Genomics vol 14 article 19pp 1ndash14 2013
[7] Y Jiang and M K Deyholos ldquoComprehensive transcriptionalprofiling of NaCl-stressed Arabidopsis roots reveals novelclasses of responsive genesrdquo BMC Plant Biology vol 6 article25 2006
[8] A M Lindroth P Saarikoski G Flygh et al ldquoTwo S-adenosyl-methionine synthetase-encoding genes differentially expressedduring adventitious root development in Pinus contortardquo PlantMolecular Biology vol 46 no 3 pp 335ndash346 2001
[9] P K Chiang R K Gordon J Tal et al ldquoS-adenosylmethionineandmethylationrdquoTheFASEB Journal vol 10 no 4 pp 471ndash4801996
[10] A M Shirley C M McMichael and C Chapple ldquoThe sng2mutant of Arabidopsis is defective in the gene encoding theserine carboxypeptidase-like protein sinapoylglucosecholinesinapoyltransferaserdquo Plant Journal vol 28 no 1 pp 83ndash942001
[11] C M Fraser M G Thompson A M Shirley et al ldquoRelatedArabidopsis serine carboxypeptidase-like sinapoylglucose acyl-transferases display distinct but overlapping substrate specifici-tiesrdquo Plant Physiology vol 144 no 4 pp 1986ndash1999 2007
[12] D Weier J Mittasch D Strack and C Milkowski ldquoThe genesBnSCT1 and BnSCT2 from Brassica napus encoding the finalenzyme of sinapine biosynthesis molecular characterizationand suppressionrdquo Planta vol 227 no 2 pp 375ndash385 2008
[13] W T Lowther and B W Matthews ldquoStructure and functionof the methionine aminopeptidasesrdquo Biochimica et BiophysicaActa vol 1477 no 1-2 pp 157ndash167 2000
[14] V S Reddy G S Ali and A S N Reddy ldquoGenes encodingcalmodulin-binding proteins in the Arabidopsis genomerdquo Jour-nal of Biological Chemistry vol 277 no 12 pp 9840ndash9852 2002
[15] S R Albright and R Tjian ldquoTAFs revisited more data revealnew twists and confirm old ideasrdquo Gene vol 242 no 1-2 pp1ndash13 2000
[16] J Kang J Park H Choi et al ldquoPlant ABC transportersrdquo TheArabidopsis Book vol 9 pp 1ndash25 2010
[17] K A Devor and J B Mudd ldquoStructural analysis of phosphat-idylcholine of plant tissuerdquo Journal of Lipid Research vol 12 no4 pp 396ndash402 1971
Submit your manuscripts athttpwwwhindawicom
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Anatomy Research International
PeptidesInternational Journal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporation httpwwwhindawicom
International Journal of
Volume 2014
Zoology
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Molecular Biology International
GenomicsInternational Journal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
The Scientific World JournalHindawi Publishing Corporation httpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
BioinformaticsAdvances in
Marine BiologyJournal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Signal TransductionJournal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
BioMed Research International
Evolutionary BiologyInternational Journal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Biochemistry Research International
ArchaeaHindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Genetics Research International
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Advances in
Virolog y
Hindawi Publishing Corporationhttpwwwhindawicom
Nucleic AcidsJournal of
Volume 2014
Stem CellsInternational
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Enzyme Research
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
International Journal of
Microbiology

Submit your manuscripts athttpwwwhindawicom
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Anatomy Research International
PeptidesInternational Journal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporation httpwwwhindawicom
International Journal of
Volume 2014
Zoology
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Molecular Biology International
GenomicsInternational Journal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
The Scientific World JournalHindawi Publishing Corporation httpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
BioinformaticsAdvances in
Marine BiologyJournal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Signal TransductionJournal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
BioMed Research International
Evolutionary BiologyInternational Journal of
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Biochemistry Research International
ArchaeaHindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Genetics Research International
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Advances in
Virolog y
Hindawi Publishing Corporationhttpwwwhindawicom
Nucleic AcidsJournal of
Volume 2014
Stem CellsInternational
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
Enzyme Research
Hindawi Publishing Corporationhttpwwwhindawicom Volume 2014
International Journal of
Microbiology



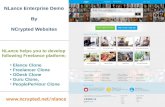




![Comparison of 61 Sequenced Escherichia coli Genomes · Comparison of 61 Sequenced Escherichia coli Genomes ... O103:H2 [37] 15578 E. coli E110019 ... Comparison of 61 Sequenced Escherichia](https://static.fdocuments.in/doc/165x107/5af461b97f8b9a92718d78d2/comparison-of-61-sequenced-escherichia-coli-of-61-sequenced-escherichia-coli-genomes.jpg)










