Lecture Transcriptional Control in Prokaryotes and Eukaryotes
Regulating Gene Expression Proteins are not required by all cells at all times Eukaryotes – 4 ways...
-
Upload
griffin-heath -
Category
Documents
-
view
215 -
download
0
Transcript of Regulating Gene Expression Proteins are not required by all cells at all times Eukaryotes – 4 ways...

Regulating Gene Expression• Proteins are not
required by all cells at all times
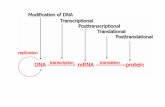
• Eukaryotes – 4 ways– Transcriptional (as
mRNA is being synthesized)
– Post-transcriptional (as mRNA is being processed)
– Translational (as proteins are made)
– Post-translational (after protein has been made)

Transcriptional Regulation• Activating gene transcription • DNA Acetylation • DNA wrapped around histones keep gene promoters inactive• Activator molecule is used (2 ways)
1. Signals a protein remodelling complex which loosen the histones exposing promoter
2. Signals an enzyme that adds an acetyl group to histones exposing promoter region (acetylation)

Transcriptional Regulation• Inhibiting gene transcription• DNA methylation (Silencing)– Methyl groups are added to the cytosine bases in the
promoter of a gene (transcription initiation complex) – Inhibits transcription – silencing– Genes are placed “on hold” until they are needed

Post Transcriptional Regulation• RBP (RNA binding proteins) • Used in:
– Pre-mRNA processing– Alternative splicing– Polyadenylation (to 3’ end)
• Rate of mRNA degradation– Masking proteins used to degrade mRNA – Translation does not occur
• Hormones – Casein – milk protein in mammary gland– When casein is needed, prolactin is produced
extending lifespan of casein mRNA– Translation continues to occur
• MicroRNA– Produced by DICER protein – Block protein production – Studied for being early cancer detection

Translational Regulation• Occurs during protein
synthesis by a ribosome• Polyadenylation
– Changes in length of poly(A) tail
– Enzymes add or delete adenines
– Increases or decreases time required to translate mRNA into protein
• Deadenylase– Removal of poly (A) tail
(polyadenylation)• Exonuclease
– Degrade mRNA after removal of poly (A) tail

Post-Translational Regulation• Proteolysis
– Removes sections of protein to make it active or inactive
• Inactivating– Removal of N-methionine
• Chemical modification– Chemical groups are added
or deleted – Puts the protein “on hold”– Phosphorylation
• Ubiquitination– Proteins tagged with
ubiquitin are degraded via proteasome

Cancer• Lack regulatory mechanisms • Mutations in genetic code
(mutagens)– Probability increases over
lifetime– Radiation, smoking, chemicals
• Mutations are passed on to daughter cells – Can lead to a mass of
undifferentiated cells (tumor)– Benign and malignant
• Oncogenes– Mutated genes that once served
to stimulate cell growth– Cause undifferentiated cell
division

Genetic Mutations• Positive and negative – Natural selection/
evolution – Cancer –death
• Small-Scale – Single base pair
• Large-Scale – Multiple base
pairs/whole genes

Small-Scale Mutations• Four groups – Missense, nonsense, silent, frameshift• Lactose, sickle cell anemia
– SNPs – single nucleotide polymorphisms• Caused by point mutations

Missense mutation• Change of a single base pair or group of base pairs• Results in the code for a different amino acid • Protein will have different sequence and structure
and may be non-functional or function differently

Nonsense mutation• Change in single base pair or group of base pairs • Results in premature stop codon • Protein will not be able to function

Silent Mutation• Change in one or more base pairs• Does not affect functioning of a gene• Mutated DNA sequence codes for same amino acid • Protein is not altered

Frameshift mutation• One or more nucleotides are inserted/deleted from a DNA
sequence• Reading frame of codons shifts resulting in multiple missense
and/or nonsense effects• Any deletion or insertion of base pairs in multiples of 3 does
not cause frameshift

Large-scale mutations • Multiple nucleotides,
entire genes, whole regions of chromosomes

Large-scale mutations • Amplification – gene
duplication– Entire genes are copied to
multiple regions of chromosomes

Large-scale mutations • Large-scale deletions – Entire coding regions of DNA are removed • Muscular Dystrophy

Large-scale mutations • Chromosomal translocation– Entire genes or groups of genes are moved from one
chromosome to another – Enhance, disrupt expression of gene

Large-scale mutations • Inversion– Portion of a DNA molecule reverses its direction in the
genome– No direct result but reversal could occur in the middle of
a coding sequence compromising the gene

Causes of genetic mutations• Spontaneous mutations– Inaccurate DNA replication
• Induced mutations – Caused by environmental agent – mutagen – Directly alter DNA – entering cell nucleus – Chemicals, radiation

Chemical Mutagens • Nitrous Acid
– Modify individual nucleotides– Nucleotides resemble other
base pairs– Confuses replication machinery
– inaccurate copying • Ethidium bromide
– Used to dye DNA/RNA– insert itself into DNA
• Aflatoxin produced by aspergillus (fungi) – found in peanut butter, corn– Low levels approved by FDA – Causes mutation of p53 gene
(acts as tumor suppressor)– Cancer causing?

Radiation - Low energy • UV B rays • Non-homologous end joining– Bonds form between adjacent nucleotides along DNA
strand – Form kinks in backbone – Skin cancer

Radiation – high energy • Ionizing radiation – x-ray, gamma rays • Strip molecules of electrons • Break bonds within DNA– Delete portions of chromosomes
• Development of tumors

Large-scale mutations • Trinucleotide repeat
expansion• Increases number of
repeats in genetic code • CAG CAG CAG CAG CAG
CAG CAG CAG • Occurs during DNA
repair/replication• “loop out” structures may
form due to repetitive nature of DNA
• Increase in expansion could cause disease or increase severity of disease– Neuromuscular/
neurodegenerative disorders

Transposable Elements (TE)• Roughly half the Human
Genome is made up TE’s• Result in mutation • DNA Transposons– Jumping genes “cut and
paste” mechanism – Move from one location of
the genome to another– Encode protein transposase
which is required for insertion and excision
– Terminal inverted repeats (9-40) base pairs long
– Less than 2% of human genome

Transposable Elements (TE)• Retrotransposons (RNA transposons)
• move through action of RNA intermediates • Produce RNA transcripts • Reverse transcriptase enzymes reverse RNA back to DNA and inserted • Give rise to variation in organism• Evolution of species 1. LINE – long interspersed transposable elements
• 6 kilobases long 2. SINE – short interspersed transposable elements (not in humans)
• Few hundred bases long

Genomes and Gene organization• Human Body– 22 autosomal chromosomes– 1 pair of each sex chromosome (XX, YY)


















![[VI]. Post-Transcriptional Processing and Post- Transcriptional Control of Gene Expression Processing of eukaryotic pre-mRNA About 60% of human genes.](https://static.fdocuments.in/doc/165x107/56649ddb5503460f94ad2a9e/vi-post-transcriptional-processing-and-post-transcriptional-control-of.jpg)