M.sc.Micro. R DNA Tech.unit 5
-
Upload
hanumant-suryawanshi -
Category
Documents
-
view
214 -
download
0
Transcript of M.sc.Micro. R DNA Tech.unit 5
-
7/29/2019 M.sc.Micro. R DNA Tech.unit 5
1/41
M.Sc.Microbiology R- DNA Technology Unit 5
1
Applications of PCRMedical applications
PCR has been applied to a large number of medical procedures:
The first application of PCR was forgenetic testing, where a sample of DNA is
analyzed for the presence ofgenetic diseasemutations. Prospective parents can be
tested for beinggenetic carriers, or their children might be tested for actually being
affected by adisease. DNA samples forPrenatal testingcan be obtained
byamniocentesis,chorionic villus sampling, or even by the analysis of rare fetal cells
circulating in the mother's bloodstream. PCR analysis is also essential
toPreimplantation genetic diagnosis, where individual cells of a developing embryo are
tested for mutations.
PCR can also be used as part of a sensitive test fortissue typing,vital toorgan
transplantation. As of 2008, there is even a proposal to replace the traditional antibody-
based tests forblood typewith PCR-based tests.
Many forms of cancer involve alterations tooncogenes. By using PCR-based tests to
study these mutations, therapy regimens can sometimes be individually customized to a
patient.
Infectious disease applications
Characterization and detection of infectious disease organisms have been revolutionized by
PCR:
The Human Immunodeficiency Virus (orHIV), of antibodies to the virus circulating in
the bloodstream. However, antibodies don't appear until many weeks after infection,
maternal antibodies mask the infection of a newborn, and therapeutic agents to fight the
infection don't affect the antibodies. PCRtestshave been developed that can detect as
little as one viral genome among the DNA of over 50,000 host cells. Infections can be
detected earlier, donated blood can be screened directly for the virus, newborns can be
immediately tested for infection, and the effects of antiviral treatments can bequantified.
http://en.wikipedia.org/wiki/Genetic_testinghttp://en.wikipedia.org/wiki/Genetic_testinghttp://en.wikipedia.org/wiki/Genetic_testinghttp://en.wikipedia.org/wiki/Genetic_diseasehttp://en.wikipedia.org/wiki/Genetic_diseasehttp://en.wikipedia.org/wiki/Genetic_mutationhttp://en.wikipedia.org/wiki/Genetic_mutationhttp://en.wikipedia.org/wiki/Genetic_mutationhttp://en.wikipedia.org/wiki/Genetic_carrierhttp://en.wikipedia.org/wiki/Genetic_carrierhttp://en.wikipedia.org/wiki/Genetic_carrierhttp://en.wikipedia.org/wiki/Cystic_Fibrosishttp://en.wikipedia.org/wiki/Cystic_Fibrosishttp://en.wikipedia.org/wiki/Cystic_Fibrosishttp://en.wikipedia.org/wiki/Prenatal_diagnosishttp://en.wikipedia.org/wiki/Prenatal_diagnosishttp://en.wikipedia.org/wiki/Prenatal_diagnosishttp://en.wikipedia.org/wiki/Amniocentesishttp://en.wikipedia.org/wiki/Amniocentesishttp://en.wikipedia.org/wiki/Amniocentesishttp://en.wikipedia.org/wiki/Chorionic_villus_samplinghttp://en.wikipedia.org/wiki/Chorionic_villus_samplinghttp://en.wikipedia.org/wiki/Chorionic_villus_samplinghttp://en.wikipedia.org/wiki/Preimplantation_genetic_diagnosishttp://en.wikipedia.org/wiki/Preimplantation_genetic_diagnosishttp://en.wikipedia.org/wiki/Preimplantation_genetic_diagnosishttp://en.wikipedia.org/wiki/Tissue_typinghttp://en.wikipedia.org/wiki/Tissue_typinghttp://en.wikipedia.org/wiki/Tissue_typinghttp://en.wikipedia.org/wiki/Organ_transplantationhttp://en.wikipedia.org/wiki/Organ_transplantationhttp://en.wikipedia.org/wiki/Organ_transplantationhttp://en.wikipedia.org/wiki/Organ_transplantationhttp://en.wikipedia.org/wiki/Blood_typehttp://en.wikipedia.org/wiki/Blood_typehttp://en.wikipedia.org/wiki/Blood_typehttp://en.wikipedia.org/wiki/Oncogenehttp://en.wikipedia.org/wiki/Oncogenehttp://en.wikipedia.org/wiki/Oncogenehttp://en.wikipedia.org/wiki/HIVhttp://en.wikipedia.org/wiki/HIVhttp://en.wikipedia.org/wiki/HIVhttp://en.wikipedia.org/wiki/HIV_testhttp://en.wikipedia.org/wiki/HIV_testhttp://en.wikipedia.org/wiki/HIV_testhttp://en.wikipedia.org/wiki/Viral_loadhttp://en.wikipedia.org/wiki/Viral_loadhttp://en.wikipedia.org/wiki/Viral_loadhttp://en.wikipedia.org/wiki/Viral_loadhttp://en.wikipedia.org/wiki/HIV_testhttp://en.wikipedia.org/wiki/HIVhttp://en.wikipedia.org/wiki/Oncogenehttp://en.wikipedia.org/wiki/Blood_typehttp://en.wikipedia.org/wiki/Organ_transplantationhttp://en.wikipedia.org/wiki/Organ_transplantationhttp://en.wikipedia.org/wiki/Tissue_typinghttp://en.wikipedia.org/wiki/Preimplantation_genetic_diagnosishttp://en.wikipedia.org/wiki/Chorionic_villus_samplinghttp://en.wikipedia.org/wiki/Amniocentesishttp://en.wikipedia.org/wiki/Prenatal_diagnosishttp://en.wikipedia.org/wiki/Cystic_Fibrosishttp://en.wikipedia.org/wiki/Genetic_carrierhttp://en.wikipedia.org/wiki/Genetic_mutationhttp://en.wikipedia.org/wiki/Genetic_diseasehttp://en.wikipedia.org/wiki/Genetic_testing -
7/29/2019 M.sc.Micro. R DNA Tech.unit 5
2/41
M.Sc.Microbiology R- DNA Technology Unit 5
2
Some disease organisms, such as that forTuberculosis, are difficult to sample from
patients and slow to begrownin the laboratory. PCR-based tests have allowed
detection of small numbers of disease organisms (both live or dead), in
convenientsamples. Detailed genetic analysis can also be used to detect antibiotic
resistance, allowing immediate and effective therapy. The effects of therapy can also be
immediately evaluated.
The spread of adiseaseorganism through populations ofdomesticorwildanimals can
be monitored by PCR testing. In many cases, the appearance of new virulent sub-
typescan be detected and monitored. The sub-types of an organism that were
responsible forearlier epidemicscan also be determined by PCR analysis.
Forensic applications]
The development of PCR-basedgenetic(orDNA) fingerprinting protocols has seen widespread
application inforensics:
In its most discriminating form,Genetic fingerprintingcan uniquely discriminate any
one person from the entire population of theworld. Minute samples of DNA can be
isolated from acrime scene, andcomparedto that from suspects, or from aDNA
databaseof earlier evidence or convicts. Simpler versions of these tests are often used
to rapidly rule out suspects during a criminal investigation. Evidence from decades-old
crimes can be tested, confirming orexoneratingthe people originally convicted.
Less discriminating forms ofDNA fingerprintingcan help inParental testing, where an
individual is matched with their close relatives. DNA from unidentified human remains
can be tested, and compared with that from possible parents, siblings, or children.
Similar testing can be used to confirm the biological parents of an adopted (or
kidnapped) child. The actual biological father of anewborncan also beconfirmed(or
ruled out).
Research applications
PCR has been applied to many areas of research in molecular genetics:
PCR allows rapid production of short pieces of DNA, even when nothing more than the
sequence of the two primers is known. This ability of PCR augments many methods,
http://en.wikipedia.org/wiki/Tuberculosishttp://en.wikipedia.org/wiki/Tuberculosishttp://en.wikipedia.org/wiki/Tuberculosishttp://en.wikipedia.org/wiki/Tuberculosis_diagnosishttp://en.wikipedia.org/wiki/Tuberculosis_diagnosishttp://en.wikipedia.org/wiki/Tuberculosis_diagnosishttp://en.wikipedia.org/wiki/Sputumhttp://en.wikipedia.org/wiki/Sputumhttp://en.wikipedia.org/wiki/Sputumhttp://en.wikipedia.org/wiki/Influenzahttp://en.wikipedia.org/wiki/Influenzahttp://en.wikipedia.org/wiki/Influenzahttp://en.wikipedia.org/wiki/Chickenhttp://en.wikipedia.org/wiki/Chickenhttp://en.wikipedia.org/wiki/Chickenhttp://en.wikipedia.org/wiki/Goosehttp://en.wikipedia.org/wiki/Goosehttp://en.wikipedia.org/wiki/Goosehttp://en.wikipedia.org/wiki/H5N1http://en.wikipedia.org/wiki/H5N1http://en.wikipedia.org/wiki/H5N1http://en.wikipedia.org/wiki/Influenza_pandemichttp://en.wikipedia.org/wiki/Influenza_pandemichttp://en.wikipedia.org/wiki/Influenza_pandemichttp://en.wikipedia.org/wiki/Genetic_fingerprintinghttp://en.wikipedia.org/wiki/Genetic_fingerprintinghttp://en.wikipedia.org/wiki/Genetic_fingerprintinghttp://en.wikipedia.org/wiki/DNA_fingerprintinghttp://en.wikipedia.org/wiki/DNA_fingerprintinghttp://en.wikipedia.org/wiki/DNA_fingerprintinghttp://en.wikipedia.org/wiki/Forensicshttp://en.wikipedia.org/wiki/Forensicshttp://en.wikipedia.org/wiki/Forensicshttp://en.wikipedia.org/wiki/Genetic_fingerprintinghttp://en.wikipedia.org/wiki/Genetic_fingerprintinghttp://en.wikipedia.org/wiki/Genetic_fingerprintinghttp://en.wikipedia.org/wiki/Earthhttp://en.wikipedia.org/wiki/Earthhttp://en.wikipedia.org/wiki/Earthhttp://en.wikipedia.org/wiki/O._J._Simpson_murder_casehttp://en.wikipedia.org/wiki/O._J._Simpson_murder_casehttp://en.wikipedia.org/wiki/O._J._Simpson_murder_casehttp://en.wikipedia.org/wiki/Combined_DNA_Index_Systemhttp://en.wikipedia.org/wiki/Combined_DNA_Index_Systemhttp://en.wikipedia.org/wiki/Combined_DNA_Index_Systemhttp://en.wikipedia.org/wiki/National_DNA_databasehttp://en.wikipedia.org/wiki/National_DNA_databasehttp://en.wikipedia.org/wiki/National_DNA_databasehttp://en.wikipedia.org/wiki/National_DNA_databasehttp://en.wikipedia.org/wiki/Innocence_Projecthttp://en.wikipedia.org/wiki/Innocence_Projecthttp://en.wikipedia.org/wiki/Innocence_Projecthttp://en.wikipedia.org/wiki/DNA_fingerprintinghttp://en.wikipedia.org/wiki/DNA_fingerprintinghttp://en.wikipedia.org/wiki/DNA_fingerprintinghttp://en.wikipedia.org/wiki/Parental_testinghttp://en.wikipedia.org/wiki/Parental_testinghttp://en.wikipedia.org/wiki/Parental_testinghttp://en.wikipedia.org/wiki/Anna_Nicole_Smithhttp://en.wikipedia.org/wiki/Anna_Nicole_Smithhttp://en.wikipedia.org/wiki/Anna_Nicole_Smithhttp://en.wikipedia.org/wiki/Paternity_testinghttp://en.wikipedia.org/wiki/Paternity_testinghttp://en.wikipedia.org/wiki/Paternity_testinghttp://en.wikipedia.org/wiki/Paternity_testinghttp://en.wikipedia.org/wiki/Anna_Nicole_Smithhttp://en.wikipedia.org/wiki/Parental_testinghttp://en.wikipedia.org/wiki/DNA_fingerprintinghttp://en.wikipedia.org/wiki/Innocence_Projecthttp://en.wikipedia.org/wiki/National_DNA_databasehttp://en.wikipedia.org/wiki/National_DNA_databasehttp://en.wikipedia.org/wiki/Combined_DNA_Index_Systemhttp://en.wikipedia.org/wiki/O._J._Simpson_murder_casehttp://en.wikipedia.org/wiki/Earthhttp://en.wikipedia.org/wiki/Genetic_fingerprintinghttp://en.wikipedia.org/wiki/Forensicshttp://en.wikipedia.org/wiki/DNA_fingerprintinghttp://en.wikipedia.org/wiki/Genetic_fingerprintinghttp://en.wikipedia.org/wiki/Influenza_pandemichttp://en.wikipedia.org/wiki/H5N1http://en.wikipedia.org/wiki/H5N1http://en.wikipedia.org/wiki/Goosehttp://en.wikipedia.org/wiki/Chickenhttp://en.wikipedia.org/wiki/Influenzahttp://en.wikipedia.org/wiki/Sputumhttp://en.wikipedia.org/wiki/Tuberculosis_diagnosishttp://en.wikipedia.org/wiki/Tuberculosis -
7/29/2019 M.sc.Micro. R DNA Tech.unit 5
3/41
M.Sc.Microbiology R- DNA Technology Unit 5
3
such as generatinghybridizationprobesforSouthernornorthern blothybridization.
PCR supplies these techniques with large amounts of pure DNA, sometimes as a single
strand, enabling analysis even from very small amounts of starting material.
The task ofDNA sequencingcan also be assisted by PCR. Known segments of DNAcan easily be produced from a patient with a genetic disease mutation. Modifications to
the amplification technique can extract segments from a completely unknown genome,
or can generate just a single strand of an area of interest.
PCR has numerous applications to the more traditional process ofDNA cloning. It can
extract segments for insertion into a vector from a larger genome, which may be only
available in small quantities. Using a single set of 'vector primers', it can also analyze or
extract fragments that have already been inserted into vectors. Some alterations to the
PCR protocol can generate mutations (general or site-directed) of an inserted
fragment.
Sequence-tagged sitesis a process where PCR is used as an indicator that a
particular segment of a genome is present in a particular clone. TheHuman Genome
Projectfound this application vital to mapping the cosmid clones they were sequencing,
and to coordinating the results from different laboratories.
An exciting application of PCR is thephylogenicanalysis of DNA fromancient sources,
such as that found in the recovered bones ofNeanderthals, or from frozen tissues
ofMammoths. In some cases the highly degraded DNA from these sources might be
reassembled during the early stages of amplification.
A common application of PCR is the study of patterns ofgene expression. Tissues (or
even individual cells) can be analyzed at different stages to see which genes have
become active, or which have been switched off. This application can also useQ-
PCRto quantitate the actual levels of expression
The ability of PCR to simultaneously amplify several loci from individual sperm[4]has
greatly enhanced the more traditional task ofgenetic mappingby
studyingchromosomal crossoversaftermeiosis. Rare crossover events between very
close loci have been directly observed by analyzing thousands of individual sperms.
Similarly, unusual deletions, insertions, translocations, or inversions can be analyzed, all
http://en.wikipedia.org/wiki/Nucleic_acid_hybridizationhttp://en.wikipedia.org/wiki/Nucleic_acid_hybridizationhttp://en.wikipedia.org/wiki/Hybridization_probehttp://en.wikipedia.org/wiki/Hybridization_probehttp://en.wikipedia.org/wiki/Hybridization_probehttp://en.wikipedia.org/wiki/Southern_blothttp://en.wikipedia.org/wiki/Southern_blothttp://en.wikipedia.org/wiki/Southern_blothttp://en.wikipedia.org/wiki/Northern_blothttp://en.wikipedia.org/wiki/Northern_blothttp://en.wikipedia.org/wiki/Northern_blothttp://en.wikipedia.org/wiki/DNA_sequencinghttp://en.wikipedia.org/wiki/DNA_sequencinghttp://en.wikipedia.org/wiki/DNA_sequencinghttp://en.wikipedia.org/wiki/DNA_cloninghttp://en.wikipedia.org/wiki/DNA_cloninghttp://en.wikipedia.org/wiki/DNA_cloninghttp://en.wikipedia.org/wiki/Sequence-tagged_sitehttp://en.wikipedia.org/wiki/Sequence-tagged_sitehttp://en.wikipedia.org/wiki/Human_Genome_Projecthttp://en.wikipedia.org/wiki/Human_Genome_Projecthttp://en.wikipedia.org/wiki/Human_Genome_Projecthttp://en.wikipedia.org/wiki/Human_Genome_Projecthttp://en.wikipedia.org/wiki/Phylogenyhttp://en.wikipedia.org/wiki/Phylogenyhttp://en.wikipedia.org/wiki/Phylogenyhttp://en.wikipedia.org/wiki/Ancient_DNAhttp://en.wikipedia.org/wiki/Ancient_DNAhttp://en.wikipedia.org/wiki/Ancient_DNAhttp://en.wikipedia.org/wiki/Neanderthalhttp://en.wikipedia.org/wiki/Neanderthalhttp://en.wikipedia.org/wiki/Neanderthalhttp://en.wikipedia.org/wiki/Mammothhttp://en.wikipedia.org/wiki/Mammothhttp://en.wikipedia.org/wiki/Mammothhttp://en.wikipedia.org/wiki/Gene_expressionhttp://en.wikipedia.org/wiki/Gene_expressionhttp://en.wikipedia.org/wiki/Gene_expressionhttp://en.wikipedia.org/wiki/Q-PCRhttp://en.wikipedia.org/wiki/Q-PCRhttp://en.wikipedia.org/wiki/Q-PCRhttp://en.wikipedia.org/wiki/Q-PCRhttp://en.wikipedia.org/wiki/Applications_of_PCR#cite_note-4http://en.wikipedia.org/wiki/Applications_of_PCR#cite_note-4http://en.wikipedia.org/wiki/Genetic_linkagehttp://en.wikipedia.org/wiki/Genetic_linkagehttp://en.wikipedia.org/wiki/Genetic_linkagehttp://en.wikipedia.org/wiki/Chromosomal_crossoverhttp://en.wikipedia.org/wiki/Chromosomal_crossoverhttp://en.wikipedia.org/wiki/Chromosomal_crossoverhttp://en.wikipedia.org/wiki/Meiosishttp://en.wikipedia.org/wiki/Meiosishttp://en.wikipedia.org/wiki/Meiosishttp://en.wikipedia.org/wiki/Meiosishttp://en.wikipedia.org/wiki/Chromosomal_crossoverhttp://en.wikipedia.org/wiki/Genetic_linkagehttp://en.wikipedia.org/wiki/Applications_of_PCR#cite_note-4http://en.wikipedia.org/wiki/Q-PCRhttp://en.wikipedia.org/wiki/Q-PCRhttp://en.wikipedia.org/wiki/Gene_expressionhttp://en.wikipedia.org/wiki/Mammothhttp://en.wikipedia.org/wiki/Neanderthalhttp://en.wikipedia.org/wiki/Ancient_DNAhttp://en.wikipedia.org/wiki/Phylogenyhttp://en.wikipedia.org/wiki/Human_Genome_Projecthttp://en.wikipedia.org/wiki/Human_Genome_Projecthttp://en.wikipedia.org/wiki/Sequence-tagged_sitehttp://en.wikipedia.org/wiki/DNA_cloninghttp://en.wikipedia.org/wiki/DNA_sequencinghttp://en.wikipedia.org/wiki/Northern_blothttp://en.wikipedia.org/wiki/Southern_blothttp://en.wikipedia.org/wiki/Hybridization_probehttp://en.wikipedia.org/wiki/Nucleic_acid_hybridization -
7/29/2019 M.sc.Micro. R DNA Tech.unit 5
4/41
M.Sc.Microbiology R- DNA Technology Unit 5
4
without having to wait (or pay for) the long and laborious processes of fertilization,
embryogenesis, etc.
=======================================================================
DNA sequencing
DNA sequencing is the process of determining the precise order ofnucleotideswithin
aDNAmolecule. It includes any method or technology that is used to determine the order of the
four basesadenine,guanine,cytosine, andthyminein a strand of DNA. The advent of rapid
DNA sequencing methods has greatly accelerated biological and medical research and
discovery.
Knowledge of DNA sequences has become indispensable for basic biological research, and in
numerous applied fields such as diagnostic, biotechnology,forensic biology, and
biologicalsystematics. The rapid speed of sequencing attained with modern DNA sequencing
technology has been instrumental in the sequencing of complete DNA sequences,
orgenomesof numerous types and species of life, including thehuman genomeand other
complete DNA sequences of many animal, plant, andmicrobialspecies.
The first DNA sequences were obtained in the early 1970s by academic researchers using
laborious methods based ontwo-dimensional chromatography. Following the developmentoffluorescence-based sequencing methods withautomated analysis,[1]DNA sequencing has
become easier and orders of magnitude faster.
Use of sequencing
DNA sequencing may be used to determine the sequence of individual genes, larger genetic
regions (i.e. clusters of genes oroperons), full chromosomes or entire genomes. Depending on
the methods used, sequencing may provide the order of nucleotides in DNA orRNAisolated
from cells of animals, plants, bacteria,archaea, or virtually any other source of genetic
information. The resulting sequences may be used by researchers inmolecular
biologyorgeneticsto further scientific progress or may be used by medical personnel to make
treatment decisions or aid ingenetic counseling.
http://en.wikipedia.org/wiki/Nucleotideshttp://en.wikipedia.org/wiki/Nucleotideshttp://en.wikipedia.org/wiki/Nucleotideshttp://en.wikipedia.org/wiki/DNAhttp://en.wikipedia.org/wiki/DNAhttp://en.wikipedia.org/wiki/DNAhttp://en.wikipedia.org/wiki/Adeninehttp://en.wikipedia.org/wiki/Adeninehttp://en.wikipedia.org/wiki/Guaninehttp://en.wikipedia.org/wiki/Guaninehttp://en.wikipedia.org/wiki/Guaninehttp://en.wikipedia.org/wiki/Cytosinehttp://en.wikipedia.org/wiki/Cytosinehttp://en.wikipedia.org/wiki/Cytosinehttp://en.wikipedia.org/wiki/Thyminehttp://en.wikipedia.org/wiki/Thyminehttp://en.wikipedia.org/wiki/Biotechnologyhttp://en.wikipedia.org/wiki/Biotechnologyhttp://en.wikipedia.org/wiki/Forensic_biologyhttp://en.wikipedia.org/wiki/Forensic_biologyhttp://en.wikipedia.org/wiki/Forensic_biologyhttp://en.wikipedia.org/wiki/Systematicshttp://en.wikipedia.org/wiki/Systematicshttp://en.wikipedia.org/wiki/Systematicshttp://en.wikipedia.org/wiki/Genomeshttp://en.wikipedia.org/wiki/Genomeshttp://en.wikipedia.org/wiki/Genomeshttp://en.wikipedia.org/wiki/Human_genomehttp://en.wikipedia.org/wiki/Human_genomehttp://en.wikipedia.org/wiki/Human_genomehttp://en.wikipedia.org/wiki/Microbehttp://en.wikipedia.org/wiki/Microbehttp://en.wikipedia.org/wiki/Microbehttp://en.wikipedia.org/wiki/Two-dimensional_chromatographyhttp://en.wikipedia.org/wiki/Two-dimensional_chromatographyhttp://en.wikipedia.org/wiki/Two-dimensional_chromatographyhttp://en.wikipedia.org/wiki/Fluorescencehttp://en.wikipedia.org/wiki/Fluorescencehttp://en.wikipedia.org/wiki/Fluorescencehttp://en.wikipedia.org/wiki/DNA_sequencerhttp://en.wikipedia.org/wiki/DNA_sequencerhttp://en.wikipedia.org/wiki/DNA_sequencing#cite_note-olsvik1993-1http://en.wikipedia.org/wiki/DNA_sequencing#cite_note-olsvik1993-1http://en.wikipedia.org/wiki/DNA_sequencing#cite_note-olsvik1993-1http://en.wikipedia.org/wiki/Genehttp://en.wikipedia.org/wiki/Genehttp://en.wikipedia.org/wiki/Genehttp://en.wikipedia.org/wiki/Operonshttp://en.wikipedia.org/wiki/Operonshttp://en.wikipedia.org/wiki/Operonshttp://en.wikipedia.org/wiki/RNAhttp://en.wikipedia.org/wiki/RNAhttp://en.wikipedia.org/wiki/RNAhttp://en.wikipedia.org/wiki/Archaeahttp://en.wikipedia.org/wiki/Archaeahttp://en.wikipedia.org/wiki/Archaeahttp://en.wikipedia.org/wiki/Molecular_biologyhttp://en.wikipedia.org/wiki/Molecular_biologyhttp://en.wikipedia.org/wiki/Molecular_biologyhttp://en.wikipedia.org/wiki/Molecular_biologyhttp://en.wikipedia.org/wiki/Geneticshttp://en.wikipedia.org/wiki/Geneticshttp://en.wikipedia.org/wiki/Geneticshttp://en.wikipedia.org/wiki/Genetic_counselinghttp://en.wikipedia.org/wiki/Genetic_counselinghttp://en.wikipedia.org/wiki/Genetic_counselinghttp://en.wikipedia.org/wiki/Genetic_counselinghttp://en.wikipedia.org/wiki/Geneticshttp://en.wikipedia.org/wiki/Molecular_biologyhttp://en.wikipedia.org/wiki/Molecular_biologyhttp://en.wikipedia.org/wiki/Archaeahttp://en.wikipedia.org/wiki/RNAhttp://en.wikipedia.org/wiki/Operonshttp://en.wikipedia.org/wiki/Genehttp://en.wikipedia.org/wiki/DNA_sequencing#cite_note-olsvik1993-1http://en.wikipedia.org/wiki/DNA_sequencerhttp://en.wikipedia.org/wiki/Fluorescencehttp://en.wikipedia.org/wiki/Two-dimensional_chromatographyhttp://en.wikipedia.org/wiki/Microbehttp://en.wikipedia.org/wiki/Human_genomehttp://en.wikipedia.org/wiki/Genomeshttp://en.wikipedia.org/wiki/Systematicshttp://en.wikipedia.org/wiki/Forensic_biologyhttp://en.wikipedia.org/wiki/Biotechnologyhttp://en.wikipedia.org/wiki/Thyminehttp://en.wikipedia.org/wiki/Cytosinehttp://en.wikipedia.org/wiki/Guaninehttp://en.wikipedia.org/wiki/Adeninehttp://en.wikipedia.org/wiki/DNAhttp://en.wikipedia.org/wiki/Nucleotides -
7/29/2019 M.sc.Micro. R DNA Tech.unit 5
5/41
M.Sc.Microbiology R- DNA Technology Unit 5
5
History
Though the structure of DNA was established as adouble helixin 1953,]several decades would
pass before fragments of DNA could be reliably analyzed for their sequence in the laboratory.
RNA sequencing was one of the earliest forms of nucleotide sequencing. The major landmark ofRNA sequencing is the sequence of the first complete gene and the complete genome
ofBacteriophage MS2, identified and published byWalter Fiersand his coworkers at
theUniversity of Ghent(Ghent,Belgium), in 1972 and 1976.
Several notable advancements in DNA sequencing were made during the 1970s. Frederick
Sangerdeveloped rapid DNA sequencing methods at theMRC Centre,Cambridge, UK and
published a method for "DNA sequencing with chain-terminating inhibitors" in 1977.Walter
GilbertandAllan MaxamatHarvardalso developed sequencing methods, including one for
"DNA sequencing by chemical degradation". In 1973, Gilbert and Maxam reported the sequence
of 24 basepairs using a method known as wandering-spot analysis.[ Advancements in
sequencing were aided by the concurrent development ofrecombinant DNAtechnology,
allowing DNA samples to be isolated from sources other than viruses.
The first full DNA genome to be sequenced was that ofbacteriophage X174in 1977Medical
Research Councilscientists deciphered the complete DNA sequence of theEpstein-Barr virusin
1984, finding it to be 170 thousand base-pairs long.
Leroy E. Hood's laboratory at theCalifornia Institute of Technologyand Smith announced the
first semi-automated DNA sequencing machine in 1986. This was followed byAppliedBiosystems' marketing of the first fully automated sequencing machine, the ABI 370, in 1987. By
1990, the U.S.National Institutes of Health(NIH) had begun large-scale sequencing trials
onMycoplasma capricolum,Escherichia coli,Caenorhabditis elegans, andSaccharomyces
cerevisiaeat a cost of US$0.75 per base. Meanwhile, sequencing of humancDNAsequences
calledexpressed sequence tagsbegan inCraig Venter's lab, an attempt to capture the coding
fraction of thehuman genome.
In 1995, Venter,Hamilton Smith, and colleagues atThe Institute for Genomic Research(TIGR)
published the first complete genome of a free-living organism, the bacteriumHaemophilus
influenzae. The circular chromosome contains 1,830,137 bases and its publication in the journal
Science]marked the first published use of whole-genome shotgun sequencing, eliminating the
need for initial mapping efforts. By 2001, shotgun sequencing methods had been used to
produce a draft sequence of the human genome.
http://en.wikipedia.org/wiki/DNA_double_helixhttp://en.wikipedia.org/wiki/DNA_double_helixhttp://en.wikipedia.org/wiki/DNA_double_helixhttp://en.wikipedia.org/wiki/DNA_sequencing#cite_note-pmid13168976-3http://en.wikipedia.org/wiki/DNA_sequencing#cite_note-pmid13168976-3http://en.wikipedia.org/wiki/DNA_sequencing#cite_note-pmid13168976-3http://en.wikipedia.org/wiki/Bacteriophage_MS2http://en.wikipedia.org/wiki/Bacteriophage_MS2http://en.wikipedia.org/wiki/Bacteriophage_MS2http://en.wikipedia.org/wiki/Walter_Fiershttp://en.wikipedia.org/wiki/Walter_Fiershttp://en.wikipedia.org/wiki/Walter_Fiershttp://en.wikipedia.org/wiki/University_of_Ghenthttp://en.wikipedia.org/wiki/University_of_Ghenthttp://en.wikipedia.org/wiki/University_of_Ghenthttp://en.wikipedia.org/wiki/Ghenthttp://en.wikipedia.org/wiki/Ghenthttp://en.wikipedia.org/wiki/Ghenthttp://en.wikipedia.org/wiki/Belgiumhttp://en.wikipedia.org/wiki/Belgiumhttp://en.wikipedia.org/wiki/Belgiumhttp://en.wikipedia.org/wiki/Frederick_Sangerhttp://en.wikipedia.org/wiki/Frederick_Sangerhttp://en.wikipedia.org/wiki/Frederick_Sangerhttp://en.wikipedia.org/wiki/Frederick_Sangerhttp://en.wikipedia.org/wiki/Medical_Research_Council_(United_Kingdom)http://en.wikipedia.org/wiki/Medical_Research_Council_(United_Kingdom)http://en.wikipedia.org/wiki/Medical_Research_Council_(United_Kingdom)http://en.wikipedia.org/wiki/Cambridgehttp://en.wikipedia.org/wiki/Cambridgehttp://en.wikipedia.org/wiki/Cambridgehttp://en.wikipedia.org/wiki/Walter_Gilberthttp://en.wikipedia.org/wiki/Walter_Gilberthttp://en.wikipedia.org/wiki/Walter_Gilberthttp://en.wikipedia.org/wiki/Walter_Gilberthttp://en.wikipedia.org/wiki/Allan_Maxamhttp://en.wikipedia.org/wiki/Allan_Maxamhttp://en.wikipedia.org/wiki/Allan_Maxamhttp://en.wikipedia.org/wiki/Harvard_Universityhttp://en.wikipedia.org/wiki/Harvard_Universityhttp://en.wikipedia.org/wiki/Harvard_Universityhttp://en.wikipedia.org/wiki/DNA_sequencing#cite_note-9http://en.wikipedia.org/wiki/DNA_sequencing#cite_note-9http://en.wikipedia.org/wiki/DNA_sequencing#cite_note-9http://en.wikipedia.org/wiki/Recombinant_DNAhttp://en.wikipedia.org/wiki/Recombinant_DNAhttp://en.wikipedia.org/wiki/Recombinant_DNAhttp://en.wikipedia.org/wiki/Bacteriophage_%CF%86X174http://en.wikipedia.org/wiki/Bacteriophage_%CF%86X174http://en.wikipedia.org/wiki/Bacteriophage_%CF%86X174http://en.wikipedia.org/wiki/Medical_Research_Council_(UK)http://en.wikipedia.org/wiki/Medical_Research_Council_(UK)http://en.wikipedia.org/wiki/Medical_Research_Council_(UK)http://en.wikipedia.org/wiki/Medical_Research_Council_(UK)http://en.wikipedia.org/wiki/Epstein-Barr_virushttp://en.wikipedia.org/wiki/Epstein-Barr_virushttp://en.wikipedia.org/wiki/Leroy_E._Hoodhttp://en.wikipedia.org/wiki/Leroy_E._Hoodhttp://en.wikipedia.org/wiki/California_Institute_of_Technologyhttp://en.wikipedia.org/wiki/California_Institute_of_Technologyhttp://en.wikipedia.org/wiki/California_Institute_of_Technologyhttp://en.wikipedia.org/wiki/Applied_Biosystemshttp://en.wikipedia.org/wiki/Applied_Biosystemshttp://en.wikipedia.org/wiki/Applied_Biosystemshttp://en.wikipedia.org/wiki/Applied_Biosystemshttp://en.wikipedia.org/wiki/National_Institutes_of_Healthhttp://en.wikipedia.org/wiki/National_Institutes_of_Healthhttp://en.wikipedia.org/wiki/National_Institutes_of_Healthhttp://en.wikipedia.org/wiki/Mycoplasma_capricolumhttp://en.wikipedia.org/wiki/Mycoplasma_capricolumhttp://en.wikipedia.org/wiki/Mycoplasma_capricolumhttp://en.wikipedia.org/wiki/Escherichia_colihttp://en.wikipedia.org/wiki/Escherichia_colihttp://en.wikipedia.org/wiki/Escherichia_colihttp://en.wikipedia.org/wiki/Caenorhabditis_eleganshttp://en.wikipedia.org/wiki/Caenorhabditis_eleganshttp://en.wikipedia.org/wiki/Caenorhabditis_eleganshttp://en.wikipedia.org/wiki/Saccharomyces_cerevisiaehttp://en.wikipedia.org/wiki/Saccharomyces_cerevisiaehttp://en.wikipedia.org/wiki/Saccharomyces_cerevisiaehttp://en.wikipedia.org/wiki/Saccharomyces_cerevisiaehttp://en.wikipedia.org/wiki/CDNAhttp://en.wikipedia.org/wiki/CDNAhttp://en.wikipedia.org/wiki/CDNAhttp://en.wikipedia.org/wiki/Expressed_sequence_taghttp://en.wikipedia.org/wiki/Expressed_sequence_taghttp://en.wikipedia.org/wiki/Expressed_sequence_taghttp://en.wikipedia.org/wiki/Craig_Venterhttp://en.wikipedia.org/wiki/Craig_Venterhttp://en.wikipedia.org/wiki/Craig_Venterhttp://en.wikipedia.org/wiki/Human_genomehttp://en.wikipedia.org/wiki/Human_genomehttp://en.wikipedia.org/wiki/Human_genomehttp://en.wikipedia.org/wiki/Hamilton_O._Smithhttp://en.wikipedia.org/wiki/Hamilton_O._Smithhttp://en.wikipedia.org/wiki/Hamilton_O._Smithhttp://en.wikipedia.org/wiki/The_Institute_for_Genomic_Researchhttp://en.wikipedia.org/wiki/The_Institute_for_Genomic_Researchhttp://en.wikipedia.org/wiki/The_Institute_for_Genomic_Researchhttp://en.wikipedia.org/wiki/Haemophilus_influenzaehttp://en.wikipedia.org/wiki/Haemophilus_influenzaehttp://en.wikipedia.org/wiki/Haemophilus_influenzaehttp://en.wikipedia.org/wiki/Haemophilus_influenzaehttp://en.wikipedia.org/wiki/DNA_sequencing#cite_note-12http://en.wikipedia.org/wiki/DNA_sequencing#cite_note-12http://en.wikipedia.org/wiki/DNA_sequencing#cite_note-12http://en.wikipedia.org/wiki/DNA_sequencing#cite_note-12http://en.wikipedia.org/wiki/Haemophilus_influenzaehttp://en.wikipedia.org/wiki/Haemophilus_influenzaehttp://en.wikipedia.org/wiki/The_Institute_for_Genomic_Researchhttp://en.wikipedia.org/wiki/Hamilton_O._Smithhttp://en.wikipedia.org/wiki/Human_genomehttp://en.wikipedia.org/wiki/Craig_Venterhttp://en.wikipedia.org/wiki/Expressed_sequence_taghttp://en.wikipedia.org/wiki/CDNAhttp://en.wikipedia.org/wiki/Saccharomyces_cerevisiaehttp://en.wikipedia.org/wiki/Saccharomyces_cerevisiaehttp://en.wikipedia.org/wiki/Caenorhabditis_eleganshttp://en.wikipedia.org/wiki/Escherichia_colihttp://en.wikipedia.org/wiki/Mycoplasma_capricolumhttp://en.wikipedia.org/wiki/National_Institutes_of_Healthhttp://en.wikipedia.org/wiki/Applied_Biosystemshttp://en.wikipedia.org/wiki/Applied_Biosystemshttp://en.wikipedia.org/wiki/California_Institute_of_Technologyhttp://en.wikipedia.org/wiki/Leroy_E._Hoodhttp://en.wikipedia.org/wiki/Epstein-Barr_virushttp://en.wikipedia.org/wiki/Medical_Research_Council_(UK)http://en.wikipedia.org/wiki/Medical_Research_Council_(UK)http://en.wikipedia.org/wiki/Bacteriophage_%CF%86X174http://en.wikipedia.org/wiki/Recombinant_DNAhttp://en.wikipedia.org/wiki/DNA_sequencing#cite_note-9http://en.wikipedia.org/wiki/Harvard_Universityhttp://en.wikipedia.org/wiki/Allan_Maxamhttp://en.wikipedia.org/wiki/Walter_Gilberthttp://en.wikipedia.org/wiki/Walter_Gilberthttp://en.wikipedia.org/wiki/Cambridgehttp://en.wikipedia.org/wiki/Medical_Research_Council_(United_Kingdom)http://en.wikipedia.org/wiki/Frederick_Sangerhttp://en.wikipedia.org/wiki/Frederick_Sangerhttp://en.wikipedia.org/wiki/Belgiumhttp://en.wikipedia.org/wiki/Ghenthttp://en.wikipedia.org/wiki/University_of_Ghenthttp://en.wikipedia.org/wiki/Walter_Fiershttp://en.wikipedia.org/wiki/Bacteriophage_MS2http://en.wikipedia.org/wiki/DNA_sequencing#cite_note-pmid13168976-3http://en.wikipedia.org/wiki/DNA_double_helix -
7/29/2019 M.sc.Micro. R DNA Tech.unit 5
6/41
M.Sc.Microbiology R- DNA Technology Unit 5
6
Several new methods for DNA sequencing were developed in the mid to late 1990s. These
techniques comprise the first of the "next-generation" sequencing methods. In 1996, Pl
Nyrnand his studentMostafa Ronaghiat the Royal Institute of Technology
inStockholmpublished their method ofpyrosequencing
A year later, Pascal Mayer and Laurent Farinelli submitted patents to the World Intellectual
Property Organization describing DNA colony sequencing. Lynx Therapeutics published and
marketed "Massively parallel signature sequencing", or MPSS, in 2000. This method
incorporated a parallelized, adapter/ligation-mediated, bead-based sequencing technology and
served as the first commercially available "next-generation" sequencing method, though
noDNA sequencerswere sold to independent laboratories. In 2004,454 Life
Sciencesmarketed a parallelized version of pyrosequencing. The first version of their machine
reduced sequencing costs 6-fold compared to automated Sanger sequencing, and was the
second of the new generation of sequencing technologies, after MPSS.
The large quantities of data produced by DNA sequencing have also required development of
new methods and programs for sequence analysis. Phil Green and Brent Ewing of the
University of Washington described theirphred quality scorefor sequencer data analysis in
1998.
Basic methods
Maxam-Gilbert sequencing
Allan MaxamandWalter Gilbertpublished a DNA sequencing method in 1977 based onchemical modification of DNA and subsequent cleavage at specific bases.[7]Also known as
chemical sequencing, this method allowed purified samples of double-stranded DNA to be used
without further cloning. This method's use of radioactive labeling and its technical complexity
discouraged extensive use after refinements in the Sanger methods had been made.
Maxam-Gilbert sequencing requires radioactive labeling at one 5' end of the DNA and
purification of the DNA fragment to be sequenced. Chemical treatment then generates breaks at
a small proportion of one or two of the four nucleotide bases in each of four reactions (G, A+G,
C, C+T). The concentration of the modifying chemicals is controlled to introduce on average onemodification per DNA molecule. Thus a series of labeled fragments is generated, from the
radiolabeled end to the first "cut" site in each molecule. The fragments in the four reactions are
electrophoresed side by side in denaturing acrylamide gels for size separation. To visualize the
fragments, the gel is exposed to X-ray film for autoradiography, yielding a series of dark bands
each corresponding to a radiolabeled DNA fragment, from which the sequence may be inferred.
http://en.wikipedia.org/wiki/P%C3%A5l_Nyr%C3%A9nhttp://en.wikipedia.org/wiki/P%C3%A5l_Nyr%C3%A9nhttp://en.wikipedia.org/wiki/P%C3%A5l_Nyr%C3%A9nhttp://en.wikipedia.org/wiki/P%C3%A5l_Nyr%C3%A9nhttp://en.wikipedia.org/wiki/Mostafa_Ronaghihttp://en.wikipedia.org/wiki/Mostafa_Ronaghihttp://en.wikipedia.org/wiki/Mostafa_Ronaghihttp://en.wikipedia.org/wiki/Stockholmhttp://en.wikipedia.org/wiki/Stockholmhttp://en.wikipedia.org/wiki/Stockholmhttp://en.wikipedia.org/wiki/Pyrosequencinghttp://en.wikipedia.org/wiki/Pyrosequencinghttp://en.wikipedia.org/wiki/Pyrosequencinghttp://en.wikipedia.org/wiki/Massively_parallel_signature_sequencinghttp://en.wikipedia.org/wiki/Massively_parallel_signature_sequencinghttp://en.wikipedia.org/wiki/Massively_parallel_signature_sequencinghttp://en.wikipedia.org/wiki/DNA_sequencershttp://en.wikipedia.org/wiki/DNA_sequencershttp://en.wikipedia.org/wiki/DNA_sequencershttp://en.wikipedia.org/wiki/454_Life_Scienceshttp://en.wikipedia.org/wiki/454_Life_Scienceshttp://en.wikipedia.org/wiki/454_Life_Scienceshttp://en.wikipedia.org/wiki/454_Life_Scienceshttp://en.wikipedia.org/wiki/Phred_quality_scorehttp://en.wikipedia.org/wiki/Phred_quality_scorehttp://en.wikipedia.org/wiki/Phred_quality_scorehttp://en.wikipedia.org/wiki/Allan_Maxamhttp://en.wikipedia.org/wiki/Allan_Maxamhttp://en.wikipedia.org/wiki/Walter_Gilberthttp://en.wikipedia.org/wiki/Walter_Gilberthttp://en.wikipedia.org/wiki/Walter_Gilberthttp://en.wikipedia.org/wiki/DNA_sequencing#cite_note-Maxam77-7http://en.wikipedia.org/wiki/DNA_sequencing#cite_note-Maxam77-7http://en.wikipedia.org/wiki/DNA_sequencing#cite_note-Maxam77-7http://en.wikipedia.org/wiki/DNA_sequencing#cite_note-Maxam77-7http://en.wikipedia.org/wiki/Walter_Gilberthttp://en.wikipedia.org/wiki/Allan_Maxamhttp://en.wikipedia.org/wiki/Phred_quality_scorehttp://en.wikipedia.org/wiki/454_Life_Scienceshttp://en.wikipedia.org/wiki/454_Life_Scienceshttp://en.wikipedia.org/wiki/DNA_sequencershttp://en.wikipedia.org/wiki/Massively_parallel_signature_sequencinghttp://en.wikipedia.org/wiki/Pyrosequencinghttp://en.wikipedia.org/wiki/Stockholmhttp://en.wikipedia.org/wiki/Mostafa_Ronaghihttp://en.wikipedia.org/wiki/P%C3%A5l_Nyr%C3%A9nhttp://en.wikipedia.org/wiki/P%C3%A5l_Nyr%C3%A9n -
7/29/2019 M.sc.Micro. R DNA Tech.unit 5
7/41
M.Sc.Microbiology R- DNA Technology Unit 5
7
Chain-termination methods
Thechain-termination methoddeveloped by Frederick Sanger and coworkers in 1977 soon
became the method of choice, owing to its relative ease and reliability. When invented, the
chain-terminator method used fewer toxic chemicals and lower amounts of radioactivity than the
Maxam and Gilbert method. Because of its comparative ease, the Sanger method was soon
automated and was the method used in the first generation ofDNA sequencers.
Sanger sequencing is the method which prevailed from the 80's until the mid-2000's. Over that
period, great advances were made in the technique, such as fluorescent labelling, capillary
electrophoresis, and general automation. These developments allowed much more efficient
sequencing, leading to lower costs. The Sanger method, in mass production form, is the
technology which produced thefirst human genomein 2001, ushering in the age ofgenomics.
However, later in the decade, radically different approaches reached the market, bringing the
cost per genome down from $100 million in 2001 to $10,000 in 2011.
Advanced methods and de novo sequencing
http://en.wikipedia.org/wiki/Sanger_sequencinghttp://en.wikipedia.org/wiki/Sanger_sequencinghttp://en.wikipedia.org/wiki/Sanger_sequencinghttp://en.wikipedia.org/wiki/DNA_sequencerhttp://en.wikipedia.org/wiki/DNA_sequencerhttp://en.wikipedia.org/wiki/DNA_sequencerhttp://en.wikipedia.org/wiki/Human_Genome_Projecthttp://en.wikipedia.org/wiki/Human_Genome_Projecthttp://en.wikipedia.org/wiki/Human_Genome_Projecthttp://en.wikipedia.org/wiki/Genomicshttp://en.wikipedia.org/wiki/Genomicshttp://en.wikipedia.org/wiki/Genomicshttp://en.wikipedia.org/wiki/File:DNA_Sequencing_gDNA_libraries.jpghttp://en.wikipedia.org/wiki/Genomicshttp://en.wikipedia.org/wiki/Human_Genome_Projecthttp://en.wikipedia.org/wiki/DNA_sequencerhttp://en.wikipedia.org/wiki/Sanger_sequencing -
7/29/2019 M.sc.Micro. R DNA Tech.unit 5
8/41
M.Sc.Microbiology R- DNA Technology Unit 5
8
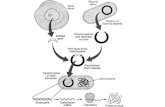
Genomic DNA is fragmented into random pieces and cloned as a bacterial library. DNA from
individual bacterial clones is sequenced and the sequence is assembled by using overlapping
DNA regions.(click to expand)
Large-scale sequencing often aims at sequencing very long DNA pieces, such as
wholechromosomes, although large-scale sequencing can also be used to generate very large
numbers of short sequences, such as found inphage display. For longer targets such as
chromosomes, common approaches consist of cutting (withrestriction enzymes) or shearing
(with mechanical forces) large DNA fragments into shorter DNA fragments. The fragmented
DNA may then beclonedinto aDNA vectorand amplified in a bacterial host such
asEscherichia coli. Short DNA fragments purified from individual bacterial colonies are
individually sequenced andassembled electronicallyinto one long, contiguous sequence.
The term "de novo sequencing" specifically refers to methods used to determine the sequence
of DNA with no previously known sequence. De novo translates from Latin as "from the
beginning". Gaps in the assembled sequence may be filled byprimer walking. The different
strategies have different tradeoffs in speed and accuracy;shotgun methodsare often used for
sequencing large genomes, but its assembly is complex and difficult, particularly withsequence
repeatsoften causing gaps in genome assembly.
Most sequencing approaches use an in vitro cloning step to amplify individual DNA molecules,
because their molecular detection methods are not sensitive enough for single molecule
sequencing. Emulsion PCR isolates individual DNA molecules along with primer-coated beads
in aqueous droplets within an oil phase. Apolymerase chain reaction(PCR) then coats each
bead with clonal copies of the DNA molecule followed by immobilization for later sequencing.
Emulsion PCR is used in the methods developed by Marguilis et al. (commercialized by454 Life
Sciences), Shendure and Porreca et al. (also known as "Polony sequencing") andSOLiD
sequencing, (developed byAgencourt, laterApplied Biosystems, nowLife Technologies).
Shotgun sequencing
Shotgun sequencing is a sequencing method designed for analysis of DNA sequences longer
than 1000 base pairs, up to and including entire chromosomes. This method requires the target
DNA to be broken into random fragments. After sequencing individual fragments, the sequences
can be reassembled on the basis of their overlapping regions.
Bridge PCR
Another method forin vitroclonal amplification is bridge PCR, in which fragments are amplified
upon primers attached to a solid surface and form "DNA colonies" or "DNA clusters". This
http://en.wikipedia.org/wiki/Chromosomehttp://en.wikipedia.org/wiki/Chromosomehttp://en.wikipedia.org/wiki/Chromosomehttp://en.wikipedia.org/wiki/Phage_displayhttp://en.wikipedia.org/wiki/Phage_displayhttp://en.wikipedia.org/wiki/Phage_displayhttp://en.wikipedia.org/wiki/Restriction_enzymehttp://en.wikipedia.org/wiki/Restriction_enzymehttp://en.wikipedia.org/wiki/Restriction_enzymehttp://en.wikipedia.org/wiki/Clone_(genetics)http://en.wikipedia.org/wiki/Clone_(genetics)http://en.wikipedia.org/wiki/Clone_(genetics)http://en.wikipedia.org/wiki/Vector_DNAhttp://en.wikipedia.org/wiki/Vector_DNAhttp://en.wikipedia.org/wiki/Vector_DNAhttp://en.wikipedia.org/wiki/Escherichia_colihttp://en.wikipedia.org/wiki/Escherichia_colihttp://en.wikipedia.org/wiki/Escherichia_colihttp://en.wikipedia.org/wiki/Sequence_assemblyhttp://en.wikipedia.org/wiki/Sequence_assemblyhttp://en.wikipedia.org/wiki/Sequence_assemblyhttp://en.wikipedia.org/wiki/Primer_walkinghttp://en.wikipedia.org/wiki/Primer_walkinghttp://en.wikipedia.org/wiki/Primer_walkinghttp://en.wikipedia.org/wiki/Shotgun_sequencinghttp://en.wikipedia.org/wiki/Shotgun_sequencinghttp://en.wikipedia.org/wiki/Shotgun_sequencinghttp://en.wikipedia.org/wiki/Microsatellite_(genetics)http://en.wikipedia.org/wiki/Microsatellite_(genetics)http://en.wikipedia.org/wiki/Microsatellite_(genetics)http://en.wikipedia.org/wiki/Microsatellite_(genetics)http://en.wikipedia.org/wiki/Polymerase_chain_reactionhttp://en.wikipedia.org/wiki/Polymerase_chain_reactionhttp://en.wikipedia.org/wiki/Polymerase_chain_reactionhttp://en.wikipedia.org/wiki/454_Life_Scienceshttp://en.wikipedia.org/wiki/454_Life_Scienceshttp://en.wikipedia.org/wiki/454_Life_Scienceshttp://en.wikipedia.org/wiki/454_Life_Scienceshttp://en.wikipedia.org/wiki/Polony_(biology)http://en.wikipedia.org/wiki/Polony_(biology)http://en.wikipedia.org/wiki/Polony_(biology)http://en.wikipedia.org/wiki/ABI_Solid_Sequencinghttp://en.wikipedia.org/wiki/ABI_Solid_Sequencinghttp://en.wikipedia.org/wiki/ABI_Solid_Sequencinghttp://en.wikipedia.org/wiki/ABI_Solid_Sequencinghttp://en.wikipedia.org/wiki/Agencourthttp://en.wikipedia.org/wiki/Agencourthttp://en.wikipedia.org/wiki/Agencourthttp://en.wikipedia.org/wiki/Applied_Biosystemshttp://en.wikipedia.org/wiki/Applied_Biosystemshttp://en.wikipedia.org/wiki/Applied_Biosystemshttp://en.wikipedia.org/wiki/Life_Technologieshttp://en.wikipedia.org/wiki/Life_Technologieshttp://en.wikipedia.org/wiki/Life_Technologieshttp://en.wikipedia.org/wiki/In_vitrohttp://en.wikipedia.org/wiki/In_vitrohttp://en.wikipedia.org/wiki/In_vitrohttp://en.wikipedia.org/wiki/DNA_sequencing#subsection_Illumina_.28Solexa.29_sequencinghttp://en.wikipedia.org/wiki/DNA_sequencing#subsection_Illumina_.28Solexa.29_sequencinghttp://en.wikipedia.org/wiki/DNA_sequencing#subsection_Illumina_.28Solexa.29_sequencinghttp://en.wikipedia.org/wiki/DNA_sequencing#subsection_Illumina_.28Solexa.29_sequencinghttp://en.wikipedia.org/wiki/In_vitrohttp://en.wikipedia.org/wiki/Life_Technologieshttp://en.wikipedia.org/wiki/Applied_Biosystemshttp://en.wikipedia.org/wiki/Agencourthttp://en.wikipedia.org/wiki/ABI_Solid_Sequencinghttp://en.wikipedia.org/wiki/ABI_Solid_Sequencinghttp://en.wikipedia.org/wiki/Polony_(biology)http://en.wikipedia.org/wiki/454_Life_Scienceshttp://en.wikipedia.org/wiki/454_Life_Scienceshttp://en.wikipedia.org/wiki/Polymerase_chain_reactionhttp://en.wikipedia.org/wiki/Microsatellite_(genetics)http://en.wikipedia.org/wiki/Microsatellite_(genetics)http://en.wikipedia.org/wiki/Shotgun_sequencinghttp://en.wikipedia.org/wiki/Primer_walkinghttp://en.wikipedia.org/wiki/Sequence_assemblyhttp://en.wikipedia.org/wiki/Escherichia_colihttp://en.wikipedia.org/wiki/Vector_DNAhttp://en.wikipedia.org/wiki/Clone_(genetics)http://en.wikipedia.org/wiki/Restriction_enzymehttp://en.wikipedia.org/wiki/Phage_displayhttp://en.wikipedia.org/wiki/Chromosome -
7/29/2019 M.sc.Micro. R DNA Tech.unit 5
9/41
M.Sc.Microbiology R- DNA Technology Unit 5
9
method is used in theIlluminaGenome Analyzersequencers. Single-molecule methods, such
as that developed byStephen Quake's laboratory (later commercialized byHelicos) are an
exception: they use bright fluorophores and laser excitation to detect base addition events from
individual DNA molecules fixed to a surface, eliminating the need for molecular amplification.
Next-generation methods
The high demand for low-cost sequencing has driven the development of high-throughput
sequencing (or next-generation sequencing) technologies thatparallelizethe sequencing
process, producing thousands or millions of sequences concurrently. High-throughput
sequencing technologies are intended to lower the cost of DNA sequencing beyond what is
possible with standard dye-terminator methods.]In ultra-high-throughput sequencing as many
as 500,000 sequencing-by-synthesis operations may be run in parallel.
Multiple, fragmented sequence reads must be assembled together on the basis of theiroverlapping areas.
Massively parallel signature sequencing (MPSS)
The first of the next-generation sequencing technologies,massively parallel signature
sequencing(or MPSS), was developed in the 1990s at Lynx Therapeutics, a company founded
in 1992 bySydney BrennerandSam Eletr. MPSS was a bead-based method that used a
complex approach of adapter ligation followed by adapter decoding, reading the sequence in
increments of four nucleotides. This method made it susceptible to sequence-specific bias or
loss of specific sequences. Because the technology was so complex, MPSS was only
performed 'in-house' by Lynx Therapeutics and no DNA sequencing machines were sold to
independent laboratories. Lynx Therapeutics merged with Solexa (later acquired byIllumina) in
2004, leading to the development of sequencing-by-synthesis, a more simple approach
acquired fromManteia Predictive Medicine, which rendered MPSS obsolete. However, the
essential properties of the MPSS output were typical of later "next-generation" data types,
including hundreds of thousands of short DNA sequences. In the case of MPSS, these were
typically used for sequencingcDNAfor measurements ofgene expressionlevels.
Polony sequencing
ThePolony sequencingmethod, developed in the laboratory ofGeorge M. Churchat Harvard,
was among the first next-generation sequencing systems and was used to sequence a full
genome in 2005. It combined an in vitro paired-tag library with emulsion PCR, an automated
microscope, and ligation-based sequencing chemistry to sequence an E. coligenome at an
accuracy of >99.9999% and a cost approximately 1/9 that of Sanger sequencing.[45]The
http://en.wikipedia.org/wiki/Illumina_(company)http://en.wikipedia.org/wiki/Illumina_(company)http://en.wikipedia.org/wiki/Illumina_(company)http://en.wikipedia.org/wiki/DNA_sequencing#subsection_Illumina_.28Solexa.29_sequencinghttp://en.wikipedia.org/wiki/DNA_sequencing#subsection_Illumina_.28Solexa.29_sequencinghttp://en.wikipedia.org/wiki/DNA_sequencing#subsection_Illumina_.28Solexa.29_sequencinghttp://en.wikipedia.org/wiki/Stephen_Quakehttp://en.wikipedia.org/wiki/Stephen_Quakehttp://en.wikipedia.org/wiki/Stephen_Quakehttp://en.wikipedia.org/wiki/Helicos_Bioscienceshttp://en.wikipedia.org/wiki/Helicos_Bioscienceshttp://en.wikipedia.org/wiki/Helicos_Bioscienceshttp://en.wikipedia.org/wiki/Multiplex_(assay)http://en.wikipedia.org/wiki/Multiplex_(assay)http://en.wikipedia.org/wiki/Multiplex_(assay)http://en.wikipedia.org/wiki/DNA_sequencing#cite_note-pmid18165802-20http://en.wikipedia.org/wiki/DNA_sequencing#cite_note-pmid18165802-20http://en.wikipedia.org/wiki/DNA_sequencing#cite_note-pmid18165802-20http://en.wikipedia.org/wiki/Massively_parallel_signature_sequencinghttp://en.wikipedia.org/wiki/Massively_parallel_signature_sequencinghttp://en.wikipedia.org/wiki/Massively_parallel_signature_sequencinghttp://en.wikipedia.org/wiki/Massively_parallel_signature_sequencinghttp://en.wikipedia.org/wiki/Sydney_Brennerhttp://en.wikipedia.org/wiki/Sydney_Brennerhttp://en.wikipedia.org/wiki/Sydney_Brennerhttp://en.wikipedia.org/wiki/Applied_Biosystems#Historyhttp://en.wikipedia.org/wiki/Applied_Biosystems#Historyhttp://en.wikipedia.org/wiki/Applied_Biosystems#Historyhttp://en.wikipedia.org/wiki/Illumina_(company)http://en.wikipedia.org/wiki/Illumina_(company)http://en.wikipedia.org/wiki/Manteia_Predictive_Medicinehttp://en.wikipedia.org/wiki/Manteia_Predictive_Medicinehttp://en.wikipedia.org/wiki/Manteia_Predictive_Medicinehttp://en.wikipedia.org/wiki/CDNAhttp://en.wikipedia.org/wiki/CDNAhttp://en.wikipedia.org/wiki/CDNAhttp://en.wikipedia.org/wiki/Gene_expressionhttp://en.wikipedia.org/wiki/Gene_expressionhttp://en.wikipedia.org/wiki/Gene_expressionhttp://en.wikipedia.org/wiki/Polony_sequencinghttp://en.wikipedia.org/wiki/Polony_sequencinghttp://en.wikipedia.org/wiki/Polony_sequencinghttp://en.wikipedia.org/wiki/George_M._Churchhttp://en.wikipedia.org/wiki/George_M._Churchhttp://en.wikipedia.org/wiki/George_M._Churchhttp://en.wikipedia.org/wiki/DNA_sequencing#cite_note-45http://en.wikipedia.org/wiki/DNA_sequencing#cite_note-45http://en.wikipedia.org/wiki/DNA_sequencing#cite_note-45http://en.wikipedia.org/wiki/DNA_sequencing#cite_note-45http://en.wikipedia.org/wiki/George_M._Churchhttp://en.wikipedia.org/wiki/Polony_sequencinghttp://en.wikipedia.org/wiki/Gene_expressionhttp://en.wikipedia.org/wiki/CDNAhttp://en.wikipedia.org/wiki/Manteia_Predictive_Medicinehttp://en.wikipedia.org/wiki/Illumina_(company)http://en.wikipedia.org/wiki/Applied_Biosystems#Historyhttp://en.wikipedia.org/wiki/Sydney_Brennerhttp://en.wikipedia.org/wiki/Massively_parallel_signature_sequencinghttp://en.wikipedia.org/wiki/Massively_parallel_signature_sequencinghttp://en.wikipedia.org/wiki/DNA_sequencing#cite_note-pmid18165802-20http://en.wikipedia.org/wiki/Multiplex_(assay)http://en.wikipedia.org/wiki/Helicos_Bioscienceshttp://en.wikipedia.org/wiki/Stephen_Quakehttp://en.wikipedia.org/wiki/DNA_sequencing#subsection_Illumina_.28Solexa.29_sequencinghttp://en.wikipedia.org/wiki/Illumina_(company) -
7/29/2019 M.sc.Micro. R DNA Tech.unit 5
10/41
M.Sc.Microbiology R- DNA Technology Unit 5
10
technology was licensed to Agencourt Biosciences, subsequently spun out into Agencourt
Personal Genomics, and eventually incorporated into the Applied Biosystems SOLiD platform,
which is now owned byLife Technologies.
454 pyrosequencing
A parallelized version ofpyrosequencingwas developed by454 Life Sciences, which has since
been acquired byRoche Diagnostics. The method amplifies DNA inside water droplets in an oil
solution (emulsion PCR), with each droplet containing a single DNA template attached to a
single primer-coated bead that then forms a clonal colony. The sequencing machine contains
manypicoliter-volume wells each containing a single bead and sequencing enzymes.
Pyrosequencing usesluciferaseto generate light for detection of the individual nucleotides
added to the nascent DNA, and the combined data are used to generate sequence read-
outs. This technology provides intermediate read length and price per base compared to Sanger
sequencing on one end and Solexa and SOLiD on the other.
Illumina (Solexa) sequencing
Solexa, now part ofIllumina, developed a sequencing method based on reversible dye-
terminators technology, and engineered polymerases, that it developed internally. The
terminated chemistry was developed internally at Solexa and the concept of the Solexa system
was invented by Balasubramanian and Klennerman from Cambridge University's chemistry
department. In 2004, Solexa acquired the companyManteia Predictive Medicinein order to gain
a massivelly parallel sequencing technology based on "DNA Clusters", which involves the clonal
amplification of DNA on a surface. The cluster technology was co-acquired with Lynx
Therapeutics of California. Solexa Ltd. later merged with Lynx to form Solexa Inc.
In this method, DNA molecules and primers are first attached on a slide and amplified
withpolymeraseso that local clonal DNA colonies, later coined "DNA clusters", are formed. To
determine the sequence, four types of reversible terminator bases (RT-bases) are added and
non-incorporated nucleotides are washed away. A camera takes images of thefluorescently
labelednucleotides, then the dye, along with the terminal 3' blocker, is chemically removed from
the DNA, allowing for the next cycle to begin. Unlike pyrosequencing, the DNA chains are
extended one nucleotide at a time and image acquisition can be performed at a delayed
moment, allowing for very large arrays of DNA colonies to be captured by sequential images
taken from a single camera.
Decoupling the enzymatic reaction and the image capture allows for optimal throughput and
theoretically unlimited sequencing capacity. With an optimal configuration, the ultimately
reachable instrument throughput is thus dictated solely by the analog-to-digital conversion rate
http://en.wikipedia.org/wiki/Life_Technologieshttp://en.wikipedia.org/wiki/Life_Technologieshttp://en.wikipedia.org/wiki/Life_Technologieshttp://en.wikipedia.org/wiki/Pyrosequencinghttp://en.wikipedia.org/wiki/Pyrosequencinghttp://en.wikipedia.org/wiki/Pyrosequencinghttp://en.wikipedia.org/wiki/454_Life_Scienceshttp://en.wikipedia.org/wiki/454_Life_Scienceshttp://en.wikipedia.org/wiki/454_Life_Scienceshttp://en.wikipedia.org/wiki/Roche_Diagnosticshttp://en.wikipedia.org/wiki/Roche_Diagnosticshttp://en.wikipedia.org/wiki/Roche_Diagnosticshttp://en.wikipedia.org/wiki/Picoliterhttp://en.wikipedia.org/wiki/Picoliterhttp://en.wikipedia.org/wiki/Picoliterhttp://en.wikipedia.org/wiki/Luciferasehttp://en.wikipedia.org/wiki/Luciferasehttp://en.wikipedia.org/wiki/Luciferasehttp://en.wikipedia.org/wiki/Solexahttp://en.wikipedia.org/wiki/Solexahttp://en.wikipedia.org/wiki/Illumina_(company)http://en.wikipedia.org/wiki/Illumina_(company)http://en.wikipedia.org/wiki/Illumina_(company)http://en.wikipedia.org/wiki/Manteia_Predictive_Medicinehttp://en.wikipedia.org/wiki/Manteia_Predictive_Medicinehttp://en.wikipedia.org/wiki/Manteia_Predictive_Medicinehttp://en.wikipedia.org/wiki/Polymerasehttp://en.wikipedia.org/wiki/Polymerasehttp://en.wikipedia.org/wiki/Polymerasehttp://en.wikipedia.org/wiki/Fluorescent_labelinghttp://en.wikipedia.org/wiki/Fluorescent_labelinghttp://en.wikipedia.org/wiki/Fluorescent_labelinghttp://en.wikipedia.org/wiki/Fluorescent_labelinghttp://en.wikipedia.org/wiki/Fluorescent_labelinghttp://en.wikipedia.org/wiki/Fluorescent_labelinghttp://en.wikipedia.org/wiki/Polymerasehttp://en.wikipedia.org/wiki/Manteia_Predictive_Medicinehttp://en.wikipedia.org/wiki/Illumina_(company)http://en.wikipedia.org/wiki/Solexahttp://en.wikipedia.org/wiki/Luciferasehttp://en.wikipedia.org/wiki/Picoliterhttp://en.wikipedia.org/wiki/Roche_Diagnosticshttp://en.wikipedia.org/wiki/454_Life_Scienceshttp://en.wikipedia.org/wiki/Pyrosequencinghttp://en.wikipedia.org/wiki/Life_Technologies -
7/29/2019 M.sc.Micro. R DNA Tech.unit 5
11/41
M.Sc.Microbiology R- DNA Technology Unit 5
11
of the camera, multiplied by the number of cameras and divided by the number of pixels per
DNA colony required for visualizing them optimally (approximately 10 pixels/colony). In 2012,
with cameras operating at more than 10 MHz A/D conversion rates and available optics, fluidics
and enzymatics, throughput can be multiples of 1 million nucleotides/second, corresponding
roughly to 1 human genome equivalent at 1xcoverageper hour per instrument, and 1 human
genome re-sequenced (at approx. 30x) per day per instrument (equipped with a single camera).
SOLiD sequencing
Library preparation for the SOLiD platform
Applied Biosystems' (now aLife Technologiesbrand) SOLiD technology employssequencing
by ligation. Here, a pool of all possible oligonucleotides of a fixed length are labeled according
to the sequenced position. Oligonucleotides are annealed and ligated; the preferential ligation
byDNA ligasefor matching sequences results in a signal informative of the nucleotide at that
position. Before sequencing, the DNA is amplified by emulsion PCR. The resulting beads, each
containing single copies of the same DNA molecule, are deposited on a glass slide. The result
is sequences of quantities and lengths comparable to Illumina sequencing.
Ion semiconductor sequencing[
Ion Torrent Systems Inc. (now owned byLife Technologies) developed a system based on
using standard sequencing chemistry, but with a novel, semiconductor based detection system.
This method of sequencing is based on the detection ofhydrogen ionsthat are released during
thepolymerisationofDNA, as opposed to the optical methods used in other sequencing
http://en.wikipedia.org/wiki/Read_depthhttp://en.wikipedia.org/wiki/Read_depthhttp://en.wikipedia.org/wiki/Read_depthhttp://en.wikipedia.org/wiki/Applied_Biosystemshttp://en.wikipedia.org/wiki/Applied_Biosystemshttp://en.wikipedia.org/wiki/Life_Technologieshttp://en.wikipedia.org/wiki/Life_Technologieshttp://en.wikipedia.org/wiki/Life_Technologieshttp://en.wikipedia.org/wiki/Sequencing_by_ligationhttp://en.wikipedia.org/wiki/Sequencing_by_ligationhttp://en.wikipedia.org/wiki/Sequencing_by_ligationhttp://en.wikipedia.org/wiki/Sequencing_by_ligationhttp://en.wikipedia.org/wiki/DNA_ligasehttp://en.wikipedia.org/wiki/DNA_ligasehttp://en.wikipedia.org/wiki/DNA_ligasehttp://en.wikipedia.org/wiki/Life_Technologieshttp://en.wikipedia.org/wiki/Life_Technologieshttp://en.wikipedia.org/wiki/Life_Technologieshttp://en.wikipedia.org/wiki/Hydrogen_ionhttp://en.wikipedia.org/wiki/Hydrogen_ionhttp://en.wikipedia.org/wiki/Hydrogen_ionhttp://en.wikipedia.org/wiki/DNA_polymerasehttp://en.wikipedia.org/wiki/DNA_polymerasehttp://en.wikipedia.org/wiki/DNA_polymerasehttp://en.wikipedia.org/wiki/DNAhttp://en.wikipedia.org/wiki/DNAhttp://en.wikipedia.org/wiki/DNAhttp://en.wikipedia.org/wiki/File:Library_preparation_for_the_SOLiD_platform.svghttp://en.wikipedia.org/wiki/DNAhttp://en.wikipedia.org/wiki/DNA_polymerasehttp://en.wikipedia.org/wiki/Hydrogen_ionhttp://en.wikipedia.org/wiki/Life_Technologieshttp://en.wikipedia.org/wiki/DNA_ligasehttp://en.wikipedia.org/wiki/Sequencing_by_ligationhttp://en.wikipedia.org/wiki/Sequencing_by_ligationhttp://en.wikipedia.org/wiki/Life_Technologieshttp://en.wikipedia.org/wiki/Applied_Biosystemshttp://en.wikipedia.org/wiki/Read_depth -
7/29/2019 M.sc.Micro. R DNA Tech.unit 5
12/41
M.Sc.Microbiology R- DNA Technology Unit 5
12
systems. A microwell containing a template DNA strand to be sequenced is flooded with a
single type ofnucleotide. If the introduced nucleotide iscomplementaryto the leading template
nucleotide it is incorporated into the growing complementary strand. This causes the release of
a hydrogen ion that triggers a hypersensitive ion sensor, which indicates that a reaction has
occurred. Ifhomopolymerrepeats are present in the template sequence multiple nucleotides will
be incorporated in a single cycle. This leads to a corresponding number of released hydrogens
and a proportionally higher electronic signal.
Sequencing of the TAGGCT template with IonTorrent, PacBioRS and GridION
DNA nanoball sequencing
DNA nanoball sequencingis a type ofhigh throughput sequencingtechnology used to
determine the entiregenomic sequenceof an organism. The companyComplete
Genomicsuses this technology to sequence samples submitted by independent researchers.
The method usesrolling circle replicationto amplify small fragments of genomic DNA into DNA
nanoballs. Unchained sequencing by ligation is then used to determine the nucleotide
sequence. This method of DNA sequencing allows large numbers of DNA nanoballs to be
sequenced per run and at lowreagentcosts compared to other next generation sequencing
platforms. However, only short sequences of DNA are determined from each DNA nanoball
which makes mapping the short reads to areference genomedifficult. This technology has been
used for multiple genome sequencing projects and is scheduled to be used for more.
Heliscope single molecule sequencing
Heliscope sequencing is a method of single-molecule sequencing developed byHelicos
Biosciences. It uses DNA fragments with added poly-A tail adapters which are attached to the
http://en.wikipedia.org/wiki/Nucleotidehttp://en.wikipedia.org/wiki/Nucleotidehttp://en.wikipedia.org/wiki/Nucleotidehttp://en.wikipedia.org/wiki/Complementarity_(molecular_biology)http://en.wikipedia.org/wiki/Complementarity_(molecular_biology)http://en.wikipedia.org/wiki/Complementarity_(molecular_biology)http://en.wikipedia.org/wiki/Homopolymerhttp://en.wikipedia.org/wiki/Homopolymerhttp://en.wikipedia.org/wiki/Homopolymerhttp://en.wikipedia.org/wiki/DNA_nanoball_sequencinghttp://en.wikipedia.org/wiki/DNA_nanoball_sequencinghttp://en.wikipedia.org/wiki/High_throughput_sequencinghttp://en.wikipedia.org/wiki/High_throughput_sequencinghttp://en.wikipedia.org/wiki/High_throughput_sequencinghttp://en.wikipedia.org/wiki/Genomic_sequencehttp://en.wikipedia.org/wiki/Genomic_sequencehttp://en.wikipedia.org/wiki/Genomic_sequencehttp://en.wikipedia.org/wiki/Complete_Genomicshttp://en.wikipedia.org/wiki/Complete_Genomicshttp://en.wikipedia.org/wiki/Complete_Genomicshttp://en.wikipedia.org/wiki/Complete_Genomicshttp://en.wikipedia.org/wiki/Rolling_circle_replicationhttp://en.wikipedia.org/wiki/Rolling_circle_replicationhttp://en.wikipedia.org/wiki/Rolling_circle_replicationhttp://en.wikipedia.org/wiki/Reagenthttp://en.wikipedia.org/wiki/Reagenthttp://en.wikipedia.org/wiki/Reagenthttp://en.wikipedia.org/wiki/Reference_genomehttp://en.wikipedia.org/wiki/Reference_genomehttp://en.wikipedia.org/wiki/Reference_genomehttp://en.wikipedia.org/wiki/Helicos_Bioscienceshttp://en.wikipedia.org/wiki/Helicos_Bioscienceshttp://en.wikipedia.org/wiki/Helicos_Bioscienceshttp://en.wikipedia.org/wiki/Helicos_Bioscienceshttp://en.wikipedia.org/wiki/File:From_second_to_fourth-generation_sequencing,_illustration_on_TAGGCT_template.svghttp://en.wikipedia.org/wiki/Helicos_Bioscienceshttp://en.wikipedia.org/wiki/Helicos_Bioscienceshttp://en.wikipedia.org/wiki/Reference_genomehttp://en.wikipedia.org/wiki/Reagenthttp://en.wikipedia.org/wiki/Rolling_circle_replicationhttp://en.wikipedia.org/wiki/Complete_Genomicshttp://en.wikipedia.org/wiki/Complete_Genomicshttp://en.wikipedia.org/wiki/Genomic_sequencehttp://en.wikipedia.org/wiki/High_throughput_sequencinghttp://en.wikipedia.org/wiki/DNA_nanoball_sequencinghttp://en.wikipedia.org/wiki/Homopolymerhttp://en.wikipedia.org/wiki/Complementarity_(molecular_biology)http://en.wikipedia.org/wiki/Nucleotide -
7/29/2019 M.sc.Micro. R DNA Tech.unit 5
13/41
M.Sc.Microbiology R- DNA Technology Unit 5
13
flow cell surface. The next steps involve extension-based sequencing with cyclic washes of the
flow cell with fluorescently labeled nucleotides (one nucleotide type at a time, as with the
Sanger method). The reads are performed by the Heliscope sequencer. The reads are short, up
to 55 bases per run, but recent improvements allow for more accurate reads of stretches of one
type of nucleotides
This sequencing method and equipment were used to sequence the genome of theM13
bacteriophage.
Single molecule real time (SMRT) sequencing[
SMRT sequencing is based on the sequencing by synthesis approach. The DNA is synthesized
in zero-mode wave-guides (ZMWs) small well-like containers with the capturing tools located
at the bottom of the well. The sequencing is performed with use of unmodified polymerase
(attached to the ZMW bottom) and fluorescently labelled nucleotides flowing freely in the
solution. The wells are constructed in a way that only the fluorescence occurring by the bottom
of the well is detected. The fluorescent label is detached from the nucleotide at its incorporation
into the DNA strand, leaving an unmodified DNA strand. According toPacific Biosciences, the
SMRT technology developer, this methodology allows detection of nucleotide modifications
(such as cytosine methylation). This happens through the observation of polymerase kinetics.
This approach allows reads of up to 15,000 nucleotides, with mean read lengths of 2.5 to 2.9
kilobases.
Methods in development
DNA sequencing methods currently under development include labeling the DNA polymerase,
reading the sequence as a DNA strand transits throughnanopores, and microscopy-based
techniques, such asatomic force microscopyortransmission electron microscopythat are used
to identify the positions of individual nucleotides within long DNA fragments (>5,000 bp) by
nucleotide labeling with heavier elements (e.g., halogens) for visual detection and
recording. Third generation technologies aim to increase throughput and decrease the time to
result and cost by eliminating the need for excessive reagents and harnessing the processivity
of DNA polymerase.
Nanopore DNA sequencing
This method is based on the readout of electrical signals occurring at nucleotides passing by
alpha-hemolysinpores covalently bound withcyclodextrin. The DNA passing through the
nanopore changes its ion current. This change is dependent on the shape, size and length of
the DNA sequence. Each type of the nucleotide blocks the ion flow through the pore for a
http://en.wikipedia.org/wiki/M13_bacteriophagehttp://en.wikipedia.org/wiki/M13_bacteriophagehttp://en.wikipedia.org/wiki/M13_bacteriophagehttp://en.wikipedia.org/wiki/M13_bacteriophagehttp://en.wikipedia.org/wiki/Pacific_Bioscienceshttp://en.wikipedia.org/wiki/Pacific_Bioscienceshttp://en.wikipedia.org/wiki/Pacific_Bioscienceshttp://en.wikipedia.org/wiki/Nanopore_sequencinghttp://en.wikipedia.org/wiki/Nanopore_sequencinghttp://en.wikipedia.org/wiki/Nanopore_sequencinghttp://en.wikipedia.org/wiki/Atomic_force_microscopehttp://en.wikipedia.org/wiki/Atomic_force_microscopehttp://en.wikipedia.org/wiki/Atomic_force_microscopehttp://en.wikipedia.org/wiki/Transmission_electron_microscopy_DNA_sequencinghttp://en.wikipedia.org/wiki/Transmission_electron_microscopy_DNA_sequencinghttp://en.wikipedia.org/wiki/Transmission_electron_microscopy_DNA_sequencinghttp://en.wikipedia.org/wiki/Hemolysinhttp://en.wikipedia.org/wiki/Hemolysinhttp://en.wikipedia.org/wiki/Hemolysinhttp://en.wikipedia.org/wiki/Cyclodextrinhttp://en.wikipedia.org/wiki/Cyclodextrinhttp://en.wikipedia.org/wiki/Cyclodextrinhttp://en.wikipedia.org/wiki/Cyclodextrinhttp://en.wikipedia.org/wiki/Hemolysinhttp://en.wikipedia.org/wiki/Transmission_electron_microscopy_DNA_sequencinghttp://en.wikipedia.org/wiki/Atomic_force_microscopehttp://en.wikipedia.org/wiki/Nanopore_sequencinghttp://en.wikipedia.org/wiki/Pacific_Bioscienceshttp://en.wikipedia.org/wiki/M13_bacteriophagehttp://en.wikipedia.org/wiki/M13_bacteriophage -
7/29/2019 M.sc.Micro. R DNA Tech.unit 5
14/41
M.Sc.Microbiology R- DNA Technology Unit 5
14
different period of time. The method has a potential of development as it does not require
modified nucleotides, however single nucleotide resolution is not yet available.
Sequencing by hybridization
Sequencing by hybridizationis a non-enzymatic method that uses aDNA microarray. A singlepool of DNA whose sequence is to be determined is fluorescently labeled and hybridized to an
array containing known sequences. Strong hybridization signals from a given spot on the array
identifies its sequence in the DNA being sequenced.
Sequencing with mass spectrometry
Mass spectrometrymay be used to determine DNA sequences. Matrix-assisted laser desorption
ionization time-of-flight mass spectrometry, orMALDI-TOF MS, has specifically been
investigated as an alternative method to gel electrophoresis for visualizing DNA fragments. With
this method, DNA fragments generated by chain-termination sequencing reactions arecompared by mass rather than by size. The mass of each nucleotide is different from the others
and this difference is detectable by mass spectrometry. Single-nucleotide mutations in a
fragment can be more easily detected with MS than by gel electrophoresis alone. MALDI-TOF
MS can more easily detect differences between RNA fragments, so researchers may indirectly
sequence DNA with MS-based methods by converting it to RNA first.
The higher resolution of DNA fragments permitted by MS-based methods is of special interest to
researchers in forensic science, as they may wish to findsingle-nucleotide polymorphismsin
human DNA samples to identify individuals. These samples may be highly degraded so forensicresearchers often prefermitochondrial DNAfor its higher stability and applications for lineage
studies. MS-based sequencing methods have been used to compare the sequences of human
mitochondrial DNA from samples in aFederal Bureau of Investigationdatabase and from bones
found in mass graves of World War I soldiers.
Early chain-termination and TOF MS methods demonstrated read lengths of up to 100 base
pairs. Researchers have been unable to exceed this average read size; like chain-termination
sequencing alone, MS-based DNA sequencing may not be suitable for large de
novo sequencing projects. Even so, a recent study did use the short sequence reads and mass
spectroscopy to compare single-nucleotide polymorphisms in pathogenicStreptococcusstrains.
Microfluidic Sanger sequencing
In microfluidic Sanger sequencing the entire thermocycling amplification of DNA fragments as
well as their separation by electrophoresis is done on a single glass wafer (approximately 10 cm
in diameter) thus reducing the reagent usage as well as cost .[70]In some instances researchers
http://en.wikipedia.org/wiki/Sequencing_by_hybridizationhttp://en.wikipedia.org/wiki/Sequencing_by_hybridizationhttp://en.wikipedia.org/wiki/DNA_microarrayhttp://en.wikipedia.org/wiki/DNA_microarrayhttp://en.wikipedia.org/wiki/DNA_microarrayhttp://en.wikipedia.org/wiki/Mass_spectrometryhttp://en.wikipedia.org/wiki/Mass_spectrometryhttp://en.wikipedia.org/wiki/Matrix-assisted_laser_desorption/ionizationhttp://en.wikipedia.org/wiki/Matrix-assisted_laser_desorption/ionizationhttp://en.wikipedia.org/wiki/Matrix-assisted_laser_desorption/ionizationhttp://en.wikipedia.org/wiki/Single-nucleotide_polymorphismshttp://en.wikipedia.org/wiki/Single-nucleotide_polymorphismshttp://en.wikipedia.org/wiki/Single-nucleotide_polymorphismshttp://en.wikipedia.org/wiki/Mitochondrial_DNAhttp://en.wikipedia.org/wiki/Mitochondrial_DNAhttp://en.wikipedia.org/wiki/Mitochondrial_DNAhttp://en.wikipedia.org/wiki/Federal_Bureau_of_Investigationhttp://en.wikipedia.org/wiki/Federal_Bureau_of_Investigationhttp://en.wikipedia.org/wiki/Federal_Bureau_of_Investigationhttp://en.wikipedia.org/wiki/Streptococcushttp://en.wikipedia.org/wiki/Streptococcushttp://en.wikipedia.org/wiki/Streptococcushttp://en.wikipedia.org/wiki/DNA_sequencing#cite_note-70http://en.wikipedia.org/wiki/DNA_sequencing#cite_note-70http://en.wikipedia.org/wiki/DNA_sequencing#cite_note-70http://en.wikipedia.org/wiki/DNA_sequencing#cite_note-70http://en.wikipedia.org/wiki/Streptococcushttp://en.wikipedia.org/wiki/Federal_Bureau_of_Investigationhttp://en.wikipedia.org/wiki/Mitochondrial_DNAhttp://en.wikipedia.org/wiki/Single-nucleotide_polymorphismshttp://en.wikipedia.org/wiki/Matrix-assisted_laser_desorption/ionizationhttp://en.wikipedia.org/wiki/Mass_spectrometryhttp://en.wikipedia.org/wiki/DNA_microarrayhttp://en.wikipedia.org/wiki/Sequencing_by_hybridization -
7/29/2019 M.sc.Micro. R DNA Tech.unit 5
15/41
M.Sc.Microbiology R- DNA Technology Unit 5
15
have shown that they can increase the throughput of conventional sequencing through the use
of microchips. Research will still need to be done in order to make this use of technology
effective.
Microscopy-based techniques
This approach directly visualizes the sequence of DNA molecules using electron microscopy.
The first identification of DNA base pairs within intact DNA molecules by enzymatically
incorporating modified bases, which contain atoms of increased atomic number, direct
visualization and identification of individually labeled bases within a synthetic 3,272 base-pair
DNA molecule and a 7,249 base-pair viral genome has been demonstrated.
RNAP sequencing
This method is based on use ofRNA polymerase(RNAP), which is attached to
apolystyrenebead. One end of DNA to be sequenced is attached to another bead, with bothbeads being placed in optical traps. RNAP motion during transcription brings the beads in closer
and their relative distance changes, which can then be recorded at a single nucleotide
resolution. The sequence is deduced based on the four readouts with lowered concentrations of
each of the four nucleotide types, similarly to the Sanger method.
In vitrovirus high-throughput sequencing
A method has been developed to analyze full sets ofprotein interactionsusing a combination of
454 pyrosequencing and an in vitro virusmRNA displaymethod. Specifically, this method
covalently links proteins of interest to the mRNAs encoding them, then detects the mRNApieces using reverse transcriptionPCRs. The mRNA may then be amplified and sequenced.
The combined method was titled IVV-HiTSeq and can be performed under cell-free conditions,
though its results may not be representative ofin vivo conditions.
Development initiatives
In October 2006, theX Prize Foundationestablished an initiative to promote the development
offull genome sequencingtechnologies, called theArchon X Prize, intending to award $10
million to "the first Team that can build a device and use it to sequence 100 human genomes
within 10 days or less, with an accuracy of no more than one error in every 100,000 bases
sequenced, with sequences accurately covering at least 98% of the genome, and at a recurring
cost of no more than $10,000 (US) per genome."
http://en.wikipedia.org/wiki/RNA_polymerasehttp://en.wikipedia.org/wiki/RNA_polymerasehttp://en.wikipedia.org/wiki/RNA_polymerasehttp://en.wikipedia.org/wiki/Polystyrenehttp://en.wikipedia.org/wiki/Polystyrenehttp://en.wikipedia.org/wiki/Polystyrenehttp://en.wikipedia.org/wiki/Interactomehttp://en.wikipedia.org/wiki/Interactomehttp://en.wikipedia.org/wiki/Interactomehttp://en.wikipedia.org/wiki/MRNA_displayhttp://en.wikipedia.org/wiki/MRNA_displayhttp://en.wikipedia.org/wiki/MRNA_displayhttp://en.wikipedia.org/wiki/Polymerase_chain_reactionhttp://en.wikipedia.org/wiki/Polymerase_chain_reactionhttp://en.wikipedia.org/wiki/Polymerase_chain_reactionhttp://en.wikipedia.org/wiki/X_Prize_Foundationhttp://en.wikipedia.org/wiki/X_Prize_Foundationhttp://en.wikipedia.org/wiki/X_Prize_Foundationhttp://en.wikipedia.org/wiki/Full_genome_sequencinghttp://en.wikipedia.org/wiki/Full_genome_sequencinghttp://en.wikipedia.org/wiki/Full_genome_sequencinghttp://en.wikipedia.org/wiki/Archon_X_Prizehttp://en.wikipedia.org/wiki/Archon_X_Prizehttp://en.wikipedia.org/wiki/Archon_X_Prizehttp://en.wikipedia.org/wiki/Archon_X_Prizehttp://en.wikipedia.org/wiki/Full_genome_sequencinghttp://en.wikipedia.org/wiki/X_Prize_Foundationhttp://en.wikipedia.org/wiki/Polymerase_chain_reactionhttp://en.wikipedia.org/wiki/MRNA_displayhttp://en.wikipedia.org/wiki/Interactomehttp://en.wikipedia.org/wiki/Polystyrenehttp://en.wikipedia.org/wiki/RNA_polymerase -
7/29/2019 M.sc.Micro. R DNA Tech.unit 5
16/41
M.Sc.Microbiology R- DNA Technology Unit 5
16
Each year theNational Human Genome Research Institute, or NHGRI, promotes grants for new
research and developments ingenomics. 2010 grants and 2011 candidates include continuing
work in microfluidic, polony and base-heavy sequencing methodologies.
Chemical DNA sequencing of DNA
The field ofbiotechnologyis one of constant change. The rapid growth anddevelopment of cutting-edge research is dependent on the innovation and creativity of scientists and their
ability to see the potential in a basic molecular technique and apply it to new processes. The advent
ofPCRopened up many doors in genetic research, including a means of DNA analysis and identificationof different genes based on theirDNAsequences. DNA sequencing is also dependent on our ability tousegel electrophoresisto separate strands of DNA that differ in size by as little as one base pair.
In the late 1970's, two DNA sequencing techniques for longer DNA molecules were invented. These werethe Sanger (or dideoxy) method and theMaxam-Gilbert (chemical cleavage) method. The Maxam-Gilbert method is based on nucleotide-specfic cleavage by chemicals and is best used to sequence
oligonucleotides (short nucleotide polymers, usually smaller than 50 base-pairs in length). The Sangermethod is more commonly used because it has been proven technically easier to apply, and, with theadvent of PCR and automation of the technique, is easily applied to long strands of DNA including some
entire genes. This technique is based on chain termination by dideoxy nucleotides during PCR elongationreactions.
In the Sanger method, the DNA strand to be analyzed is used as a template and DNA polymerase is used,in a PCR reaction, to generate complimentary strands using primers. Four different PCR reactionmixtures are prepared, each containing a certain percentage of dideoxynucleoside triphosphate (ddNTP)analogs to one of the four nucleotides (ATP, CTP, GTP or TTP). Synthesis of the new DNA strandcontinues until one of these analogs is incorporated, at which time the strand is prematurely truncated.
Each PCR reaction will end up containing a mixture of different lengths of DNA strands, all ending withthe nucleotide that was dideoxy labeled for that reaction. Gel electrophoresis is then used to separate thestrands of the four reactions, in four separate lanes, and determine the sequence of the original template
based on what lengths of strands end with what nucleotide.
In the automated Sanger reaction, primers are used that are labeled with four different colouredfluorescent tags. PCR reactions, in the presence of the different dideoxy nucleotides, are performed asdescribed above. However, next, the four reaction mixtures are then combined and applied to a single laneof a gel. The colour of each fragment is detected using a laser beam and the information is collected by acomputer which generates chromatograms showing peaks for each colour, from which the template DNAsequence can be determined.
http://en.wikipedia.org/wiki/National_Human_Genome_Research_Institutehttp://en.wikipedia.org/wiki/National_Human_Genome_Research_Institutehttp://en.wikipedia.org/wiki/National_Human_Genome_Research_Institutehttp://en.wikipedia.org/wiki/Genomicshttp://en.wikipedia.org/wiki/Genomicshttp://en.wikipedia.org/wiki/Genomicshttp://biotech.about.com/od/pcr/a/od/whatisbiotechnology/a/whatisbiotech.htmhttp://biotech.about.com/od/pcr/a/od/whatisbiotechnology/a/whatisbiotech.htmhttp://biotech.about.com/od/pcr/a/od/whatisbiotechnology/a/whatisbiotech.htmhttp://biotech.about.com/od/pcr/a/od/pcr/a/PCRtheory.htmhttp://biotech.about.com/od/pcr/a/od/pcr/a/PCRtheory.htmhttp://biotech.about.com/od/pcr/a/od/pcr/a/PCRtheory.htmhttp://biotech.about.com/od/glossary/g/What-Is-Dna.htmhttp://biotech.about.com/od/glossary/g/What-Is-Dna.htmhttp://biotech.about.com/od/glossary/g/What-Is-Dna.htmhttp://biotech.about.com/od/protocols/ht/DNAgel.htmhttp://biotech.about.com/od/protocols/ht/DNAgel.htmhttp://biotech.about.com/od/protocols/ht/DNAgel.htmhttp://biotech.about.com/od/protocols/ht/DNAgel.htmhttp://biotech.about.com/od/glossary/g/What-Is-Dna.htmhttp://biotech.about.com/od/pcr/a/od/pcr/a/PCRtheory.htmhttp://biotech.about.com/od/pcr/a/od/whatisbiotechnology/a/whatisbiotech.htmhttp://en.wikipedia.org/wiki/Genomicshttp://en.wikipedia.org/wiki/National_Human_Genome_Research_Institute -
7/29/2019 M.sc.Micro. R DNA Tech.unit 5
17/41
M.Sc.Microbiology R- DNA Technology Unit 5
17
Typically, the automated sequencing method is only accurate for sequences up to a maximum of about700-800 base-pairs in length. However, it is possible to obtain full sequences of larger genes and, in fact,
whole genomes, using step-wise methods such as Primer Walking and Shotgun sequencing.
In Primer Walking, a workable portion of a larger gene is sequenced using the Sanger method. New
primers are generated from a reliable segment of the sequence, and used to continue sequencing theportion of the gene that was out of range of the original reactions.
Shotgun sequencing entails randomly cutting the DNA segment of interest into more appropriate
(manageable) sized fragments, sequencing each fragment, and arranging the pieces based on overlappingsequences. This technique has been made easier by the application of computer software for arr
=====================================================================================
RNA-Seq
RNA-seq, also called "Whole Transcriptome Shotgun Sequencing" ("WTSS"), refers to the use
ofhigh-throughput sequencingtechnologies to sequencecDNAin order to get information about
a sample'sRNAcontent. The technique has been adopted in studies of diseases like cancer.
With deep coverage and base-level resolution,next-generation sequencingprovides information
on differential expression of genes, including gene alleles and differently spliced transcripts;
non-coding RNAs; post-transcriptional mutations or editing; and gene fusions.
Introduction
The introduction ofnext-generation sequencingorhigh-throughput sequencingtechnologies
opened new doors into the field ofDNA sequencing, however as understanding of these
technologies becomes more widespread and new tools are developed, new innovative ways of
applying these technologies are being created.
Givenhigh-throughput sequencingtechnologies' low requirements of nucleotide sequence
product, together with its deep coverage and base-scale resolution, its use has expanded to the
field oftranscriptomics.]
Transcriptomics is an area of research characterizing the RNAtranscribed from a particulargenomeunder investigation.
Although transcriptomes are more dynamic than genomicDNA, these molecules provide direct
access togene regulationand protein information. Sequencing transcriptomes is not a new
idea, however: various methods have been developed previously to directly determine cDNA
sequences, based mostly around traditional (and more expensive)Sanger sequencing, while
http://en.wikipedia.org/wiki/High-throughput_sequencinghttp://en.wikipedia.org/wiki/High-throughput_sequencinghttp://en.wikipedia.org/wiki/High-throughput_sequencinghttp://en.wikipedia.org/wiki/CDNAhttp://en.wikipedia.org/wiki/CDNAhttp://en.wikipedia.org/wiki/CDNAhttp://en.wikipedia.org/wiki/RNAhttp://en.wikipedia.org/wiki/RNAhttp://en.wikipedia.org/wiki/RNAhttp://en.wikipedia.org/wiki/Cancerhttp://en.wikipedia.org/wiki/Cancerhttp://en.wikipedia.org/wiki/Cancerhttp://en.wikipedia.org/wiki/Next-generation_sequencinghttp://en.wikipedia.org/wiki/Next-generation_sequencinghttp://en.wikipedia.org/wiki/Next-generation_sequencinghttp://en.wikipedia.org/wiki/Next-generation_sequencinghttp://en.wikipedia.org/wiki/Next-generation_sequencinghttp://en.wikipedia.org/wiki/Next-generation_sequencinghttp://en.wikipedia.org/wiki/High-throughput_sequencinghttp://en.wikipedia.org/wiki/High-throughput_sequencinghttp://en.wikipedia.org/wiki/High-throughput_sequencinghttp://en.wikipedia.org/wiki/DNA_sequencinghttp://en.wikipedia.org/wiki/DNA_sequencinghttp://en.wikipedia.org/wiki/DNA_sequencinghttp://en.wikipedia.org/wiki/High-throughput_sequencinghttp://en.wikipedia.org/wiki/High-throughput_sequencinghttp://en.wikipedia.org/wiki/High-throughput_sequencinghttp://en.wikipedia.org/wiki/Transcriptomicshttp://en.wikipedia.org/wiki/Transcriptomicshttp://en.wikipedia.org/wiki/RNA-Seq#cite_note-wang2009-4http://en.wikipedia.org/wiki/RNA-Seq#cite_note-wang2009-4http://en.wikipedia.org/wiki/RNA-Seq#cite_note-wang2009-4http://en.wikipedia.org/wiki/Genomehttp://en.wikipedia.org/wiki/Genomehttp://en.wikipedia.org/wiki/Genomehttp://en.wikipedia.org/wiki/DNAhttp://en.wikipedia.org/wiki/DNAhttp://en.wikipedia.org/wiki/DNAhttp://en.wikipedia.org/wiki/Gene_regulationhttp://en.wikipedia.org/wiki/Gene_regulationhttp://en.wikipedia.org/wiki/Gene_regulationhttp://en.wikipedia.org/wiki/Sanger_sequencinghttp://en.wikipedia.org/wiki/Sanger_sequencinghttp://en.wikipedia.org/wiki/Sanger_sequencinghttp://en.wikipedia.org/wiki/Sanger_sequencinghttp://en.wikipedia.org/wiki/Gene_regulationhttp://en.wikipedia.org/wiki/DNAhttp://en.wikipedia.org/wiki/Genomehttp://en.wikipedia.org/wiki/RNA-Seq#cite_note-wang2009-4http://en.wikipedia.org/wiki/Transcriptomicshttp://en.wikipedia.org/wiki/High-throughput_sequencinghttp://en.wikipedia.org/wiki/DNA_sequencinghttp://en.wikipedia.org/wiki/High-throughput_sequencinghttp://en.wikipedia.org/wiki/Next-generation_sequencinghttp://en.wikipedia.org/wiki/Next-generation_sequencinghttp://en.wikipedia.org/wiki/Cancerhttp://en.wikipedia.org/wiki/RNAhttp://en.wikipedia.org/wiki/CDNAhttp://en.wikipedia.org/wiki/High-throughput_sequencing -
7/29/2019 M.sc.Micro. R DNA Tech.unit 5
18/41
M.Sc.Microbiology R- DNA Technology Unit 5
18
others include methodologies such asSeri





![BT 0502 - M.Tech r-DNA Technology lab manual[1] 0502 - M...BT 0502 - M.Tech r-DNA Technology lab manual IInd Semester Department of Biotechnology School of Bioengineering BT 0502 r-DNA](https://static.fdocuments.in/doc/165x107/5abf928b7f8b9aa3088e3e9f/bt-0502-mtech-r-dna-technology-lab-manual1-0502-mbt-0502-mtech-r-dna.jpg)