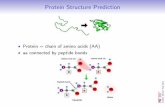
Molecular Dynamics: Review. Molecular Simulations NMR or X-ray structure refinements Protein...
-
Upload
steven-vize -
Category
Documents
-
view
227 -
download
7
Transcript of Molecular Dynamics: Review. Molecular Simulations NMR or X-ray structure refinements Protein...

Molecular Dynamics: Review

Molecular Simulations
• NMR or X-ray structure refinements
• Protein structure prediction• Protein folding kinetics and
mechanics• Conformational dynamics• Global optimization• DNA/RNA simulations• Membrane proteins/lipid
layers simulations

Molecular Dynamics
From Lecture 6 (Robert):• MD is our approximation to how molecules explore theirpotential energy surface in the real world• – The atoms are “heated” by giving them a distribution ofvelocities corresponding to temperature we wish to simulate• – The wiggling and jiggling of the atoms is then obtained byintegrating the Newtonian laws of motion• – This gives us the Ei's of all states “i” occupied at thattemperature as long as we simulate long enough

I. Force Fields

Force Fields: Typical Energy Functions
20
20
12 6
1( )
2
1( )
2
[1 cos( )]2
( )
[ ]
rbonds
angles
n
torsions
improper
i j
elec ij
ij ij
LJ ij ij
U k r r
k
Vn
V improper torsion
q q
r
A B
r r
Bond stretches
Angle bending
Torsional rotation
Improper torsion (sp2)
Electrostatic interaction
Lennard-Jones interaction

Bonding Terms: bond stretch
• Most often Harmonic
20 )(
2
1rrkV r
bondsbond
Harmonic Potential
bond length
Vbond
r0

Bonding Terms: angle bending
• Most often Harmonic
• CHARMM force field’s Urey-Bradley angle term:
20 )(
2
1 kVangles
angle
Harmonic Potential
angle
Vangle
0
20 )(
2
1sskV UB
UBUB
This UB term is only found in CHARMM force field to optimize the fit to vibrational spectra. s: the 1,3-distance.
Mackerell et al. J. Phys. Chem. B 102, 3586, 1998

Bonding Terms: Torsions
• Torsion energy: rotation about a bond (dihedral angles)
)]cos(1[2
nV
Utorsions
ntorsion
Vn: force constant n: periodicity of the angle ( determines how many peaks and wells in the potential, often from 1-6 ) : phase of the angle (often 0º or 180º)
i
lj
k
i-j-k-l

Bonding Terms: Improper Torsions
• Improper torsion is not a regular torsion angle. It is used to describe the energy of out-of-plane motions. It is often necessary for planar groups, such as sp2 hybridized carbons in carbonyl groups and in aromatic rings, because the normal torsion terms described above is not sufficient to maintain the planarity (w~0).
)]1802cos(1[22
improperimproper
VU
20 )(
2
improper
wimproper
kU
or
j
li
k
i-j-k-l

Non-bonded Terms
• Electrostatic interactions (Coulomb’s Law)
• Lennard-Jones interactions
ji ij
jielec r
qqV
41
ji ij
ij
ij
ijijLJ
rrV 6
6
12
12
4
Coulomb Potential
pair distance
Vele
c
LJ Potential
pair distance r/sigma
VL
J
~1/r

II. Solvation Models

Solvation Models
• Explicit solvent models– Fixed charge models: SPC, SPC/E, TIP3P,
TIP4P, TIP5P, ST2,…– Polarizable water models: TIP4P/FQ, POL5,
MCDHO,…
• Implicit Solvent models– Poisson-Boltzman solver (Delphi, Honig)– Generalized Born Model (Still)– Karplus’ EEF1 model – Benoit Roux’s Spherical Solvent Boundary
Potential (SSBP)

Explicit Water modelsSPC, SPC/E, TIPnP, POL5

Water Model Geometries

Water Model Parameters• SPC, SPC/E (Berendsen)• TIP3P, TIP4P, TIP5P (Jorgensen)• TIP4P/FQ, POL5 (Berne)

Implicit Solvent ModelsPBF, GB

Continuum Solvent Model
continuum solvent=80
=1-4protein

III. Molecular Dynamics

Molecular Dynamics
• Solve Newton’s equation for a molecular system:
amF

Integrator: Verlet Algorithm
)()()()(2)( 42 tOtatttrtrttr
)()(2
1)()()( 32 tOtatttvtrttr
)()(2
1)()()( 32 tOtatttvtrttr
Start with {r(t), v(t)}, integrate it to {r(t+t), v(t+t)}:
{r(t), v(t)}
{r(t+t), v(t+t)}
The new position at t+t:
Similarly, the old position at t-t:
(1)
(2)
Add (1) and (2):
Thus the velocity at t is:
(3)
)())()((2
1)()( 2tOttrttr
ttrtv
(4)

Typical MD Flowchart
Program MYMD simple MD program
call init initialization t = 0 do while (t .lt. tmax) MD loop call force (x, f, en) calculate the force call integrate (x, f, en) integrate equation of motion t = t + delt call sample sample averages enddo stop end

Periodic Boundary ConditionsMinimum Image
Central simulationbox
rc

One MD example
Determining voltage threshold for translocation of dsDNA through Si3N4 pores
To establish the threshold field required to drive dsDNA through a 2.0 nanometer diameter pore. The 3.9 V path caused the partial unzipping of the DNA strands prior to reaching the center of the membrane.
http://www.ks.uiuc.edu/Research/nanopore/

Historical Perspective on MD

The Next Generation in MD
• Current longest MD simulations: microsecond vs. time scale of many biologically interesting phenomena is millisecond
• Anton, Desmond• Scientific advances &
Drug Discovery
Faculty in Computer Science Department at Columbia University, till1986
D. E. Shaw & Co., Inc., founded in 1988
1994, pointed by President Clinton, President's Council of Advisors on Science and Technology

Acknowledgement
• Powerpoint slices from Ruhong Zhou