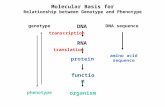
Molecular Basis for Relationship between Genotype and Phenotype
description
Transcript of Molecular Basis for Relationship between Genotype and Phenotype

Molecular Basis forRelationship between Genotype and Phenotype
DNA
RNA
protein
genotype
function
organismphenotype
DNA sequence
amino acidsequence
transcription
translation

Vectors for Larger DNA Inserts
Fosmids:
Hybrid between phage DNA and plasmid DNA - can carry inserts 35-kb to 45-kb
PAC:
P1 Artificial Chromosome (derivative of bacteriophage P1) - can carry inserts 80-kb to 100-kb
BAC:
Bacterial Artificial Chromosome (derivative of F plasmid) - can carry inserts 150-kb to 300 kb
YAC:
Yeast Artificial Chromosome - can carry inserts larger than 300-kb

Modes of delivering recombinant DNA into bacterial cells
(a) Plasmid DNA is introduced into host cell by transformation.
(b) Fosmids are introduced in phage heads by transduction. Once inside, they replicate as large plasmids.
(c) Phage vectors are introduced by infection.

Molecular Basis forRelationship between Genotype and Phenotype
DNA
RNA
protein
genotype
function
organismphenotype
DNA sequence
amino acidsequence
transcription
translation

Four Ribonucleotides Found in RNA


Complementarity and Asymmetry in RNA Synthesis
Only one strand of DNA is used as template for RNA synthesis.
Template strand is antiparallel to RNA transcript.
RNA bases are complementary to bases of template DNA.

RNA Polymerase
RNA polymerase adds ribonucleotides in 5’ to 3’ direction.
RNA polymerase in E. coli consists of 4 different subunits (see model below).
A single type of RNA polymerase transcribes RNA in prokaryotes.
recognizes the promoter. Holoenzyme is needed for correct initiation of transcription.

Promoters signal transcription in prokaryotes.
Promoter Sequences in E. coli

subunit positions RNA polymerase for correct initiation.
Upon initiation of transcription, subunit dissociates.
Transcription Initiation in Prokaryotes

Elongation
RNA polymerase adds ribonucleotides in 5’ to 3’ direction.
RNA polymerase catalyzes the following reaction:
NTP + (NMP)n (NMP)n+1 + PPiDNA
Mg++RNA polymerase

Termination
Termination of transcription occurs beyond the coding sequence of a gene. This region is 3’ untranslated region (3’ UTR), which is recognized by RNA polymerase.

Termination
RNA polymerase recognizes signals for chain termination.
(1) Intrinsic: Termination site on template DNA consists of GC-rich sequences followed by A’s. Intra-molecular hydrogen bonding causes formation of hairpin loop.
In E. coli, this structure signals release of RNA polymerase, thus terminating transcription.
(2) rho factor (hexameric protein) dependent: These termination signals do not produce hairpin loops. rho binds to RNA at rut site. rho pulls RNA away from RNA polymerase.
rut site

Colinearity of Gene and Protein
DNA
RNA
protein
genotype
function
organismphenotype
DNA sequence
amino acidsequence
transcription
translation

There are three stop (termination) codons. They are often called nonsense codons.
Genetic Code is degenerate. Some amino acids are encoded by more than one codon.
Genetic Code
Genetic Code is nonoverlapping.
A codon (three bases or triplet) encodes an amino acid.
Genetic Code is read continuously from a fixed starting point.
There is a start codon (AUG).

Molecular Basis forRelationship between Genotype and Phenotype
DNA
RNA
protein
genotype
function
organismphenotype
DNA sequence
amino acidsequence
transcription
translation

Eukaryotic RNA
Three RNA Polymerases
RNA Polymerase
IIIIII
Synthesis of
rRNA (except 5S rRNA)mRNA*, some snRNA
tRNA, some snRNA, 5S rRNA
* eukaryotic RNA is monocistronic prokaryotic RNA can be polycistronic

Eukaryotic RNA
Primary transcript (pre-mRNA) must be processed into mature mRNA.
1. Cap at 5’ end (7-methylguanosine)2. Addition of poly(A) tail3. Splicing of RNA transcript
Many proteins must assemble at promoter before transcription.
General transcription factors (GTF’s) bind before RNA polymerase II, while other proteins bind after RNA polymerase II binds.
Chromatin structure affects gene expression (gene transcription) in eukaryotes.

Prokaryotic and Eukaryotic Transcription and Translation Compared

TATA binding protein (TBP), part of TFIID complex, must bind to promoter before other GTFs and RNA polymerase II can form preinitiation complex (PIC).
Phosphorylation of carboxyl tail domain (CTD), the protein tail of subunit of RNA polymerase II, allows separation of RNA polymerase II from GTFs to start transcription.
Transcription Initiation in Eukaryotes

State of phosphorylation of CTD determines the type of proteins that can associate with the CTD (thus defining cotranscriptional process).
5’ end of pre-mRNA is capped with 7-methylguanosine. This protects the transcript from degradation; capping is also necessary for translation of mature mRNA.
Cotranscriptional Processing of RNA

Cotranscriptional Processing
3’ end of the transcript typically contains AAUAAA or AUUAAA.
This sequence is recognized by an enzyme that cleaves the newly synthesized transcript ~20 nucleotides downstream.
At the 3’ end, a poly(A) tail consisting of 150 - 200 adenine nucleotides is added.
Polyadenylation is another characteristic of transcription in eukaryotes.

Different mRNA can be produced; different -tropomyosin can be produced.Alternative splicing is a mechanism for gene regulation. Gene product can be differentin different cell types and at different stages of development.
Complex Patterns of Eukaryotic RNA Splicing

Intron Splicing: Conserved Sequences
exons - coding sequences introns - noncoding sequences
Small nuclear ribonucleoprotein particles (snRNPs) recognize consensus splice junction sequence of GU/AG.
snRNPs are complexes of protein and small nuclear RNA (snRNA). Several snRNPs comprise a spliceosome.
Spliceosome directs the removal of introns and joining of exons.

One end of conserved sequence attaches to conserved adenine in the intron.
The “lariat” is released and adjacent exons are joined.
Spliceosome interacts with CTD and attaches to pre-mRNA.
snRNAs in spliceosomes direct alignment of the splice sites.
Spliceosome Assembly and Function

Reactions in Exon Splicing

These self-splicing introns are an example of RNA that can catalyze a reaction.
RNA molecules can act somewhat like enzymes (ribozymes).
In the protozoan Tetrahymena, the primary transcript of an rRNA can excise a 413-nucleotide intron from itself.
Self-Splicing Reaction