Noise and Fluctuations: Twentieth International Conference on Noise and Fluctuations
Modeling fluctuations in the force-extension single-molecule experiments Alexander Vologodskii New...
-
Upload
basil-preston -
Category
Documents
-
view
218 -
download
0
Transcript of Modeling fluctuations in the force-extension single-molecule experiments Alexander Vologodskii New...

Modeling fluctuations in the force-extension single-molecule experiments
Alexander Vologodskii
New York University

Diagram of the force-extension experiment
F

0.0 0.2 0.4 0.6 0.8 1.00
20
40
60
80- Experimental data of Smith et al.
chainWorm-like
Extension, <x>/L
The force-extension dependence for DNA is well studied

The force has entropic nature

From: J. W. Shaevitz1, E. A. Abbondanzieri, R. Landick & S. M. Block. Backtracking by single RNA polymerase molecules observed at near-base-pair resolution. Nature, 426, 684-687.
The entropic force from extended DNA molecule is used in many single-molecule experiments

DNA model for Brownian dynamics simulations
The intersegment interaction is specified by the Debye-Hückel potential
Segments are stretchable
Virtual beads of a certain diameter placed at chain vertices specify hydrodynamic interaction with solution and between the beads
Discrete wormlike chain with some modifications:

Dynamics of the chain is described by the Langevin equations
midvi
dt ij
j v j Fi ij
j f j
ij
where
is a configuration-dependent friction tensor
is a force acting on beadFi i
represents the randomly fluctuating force resulting from the thermal motion of the surrounding fluid
ijj f j
is the mass of bead imi

How accurate is Brownian dynamics simulation of DNA properties?
Comparison of measured and simulated diffusion coefficients of knots along stretched DNA moleculeshows that simulation is quite accurate

Tying knots by optical tweezers

Experimental measurement of knot diffusion
X. R. Bao, H.J. Lee and S.R. Quake, Phys. Rev. Lett., 91, 265506 (2003)

Brownian dynamics simulation of knot diffusion
Typical simulated conformations of knotted model chains

Comparison of the measured and computed diffusion coefficients of knots
Knot type
Computed diffusion
coefficient, m2/s
Measured diffusion
coefficient, m2/s
8.6 ± 1
12.5 ± 0.5
6.0 ± 1
7.9 ± 0.3
2.5 ± 0.3
4.8 ± 0.2

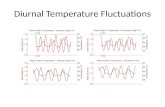
Simulated values of the force fluctuate strongly
Time, ns
0.0 0.2 0.4 0.6 0.8 1.0
For
ce,
Fb/
kT
-10
-5
0
5
10
Time, ns
0.0 0.2 0.4 0.6 0.8 1.0
x/L
0.79
0.80
0.81
0.82
0.83
0.84

Time, ns
0 20 40 60 80 100
For
ce,
pn
-20
-10
0
10
The force fluctuations do not depend on its average value
Each point is the averaging over 1 ns

The force averaging does not occur over 0.1 s
Each point is the averaging over 100 ns
Time, s
0 2 4 6 8 10
For
ce,
pn
-4
-2
0
2
4dt = 400 psdt = 4 ps

Time, ms
0.0 0.2 0.4 0.6 0.8 1.0
For
ce,
pn
-1.0
-0.5
0.0dt = 400 ps
The force averaging does not occur over 10 s

Time, ms
0 20 40 60 80 100
For
ce,
Fb/
kT
-0.7
-0.6
-0.5
-0.4
-0.3
A good averaging of the force is achieved by averaging over 1 ms

Fluctuations of the force do not depend on DNA length
DNA length, bp
1000 10000
x, p
n0
20
40
60
80
f = 0.5 pn
f = 2.5 pn
f = 8.3 pn
DNA length, bp
1000 10000
f, pn
0
1
2
3
4
5
f = 0.1 pn
f = 0.5 pn
f = 2.5 pnf = 8.3 pn

Presence of a protein-induced bend decreases DNA extension
Can the extension measurement be used to determine the bend angle?

Simulated values of the extension reduction resultingfrom DNA bending by angle

Time
Ext
ensi
on
No bound protein No bound proteinOne protein is bound
Large fluctuations of the extension and a finite time of the protein-bound state create a problem

DNA length, bp
0 1000 2000 3000
, n
m
0
50
100
150F = 0.1 pn
F = 1 pn
The variations of the extensions are large

Extension of a single DNA molecule by force
ForceForce
These are actual proportions for 1500 bp DNA

time, ms
0 20 40 60 80
Ext
ensi
on,
nm
0
100
200
300
400L = 500 nmF = 0.1 pn
No beadBead radius 500 nm
Fluctuations of DNA extension averaged over 0.4 ms

Fluctuations of DNA extension averaged over 40 ms
Time, ms
200 400 600 800
Ext
ensi
on, n
m
200
400L = 500 nmF = 0.1 pn
No beadBead radius is 500 nm

Interval of averaging, ms
0.01 0.1 1 10 100 1000
Ext
ensi
on v
aria
tion,
nm
10
100
L = 500 nmF = 0.1 pn
With bead R = 500 nmNo bead
What averaging interval do we need?

The work was supported by NIH


Displacement of unknotted part of the model chain eliminates the chain length restriction