Midline Signaling and Evolution of the Forebrain in Chordates: A ...
Transcript of Midline Signaling and Evolution of the Forebrain in Chordates: A ...

SYMPOSIUM
Midline Signaling and Evolution of the Forebrain in Chordates:A Focus on the Lamprey Hedgehog CaseSylvie Retaux1 and Shungo Kano
NeD-UPR3294, CNRS, Institut Alfred Fessard, avenue de la Terrasse, 91198 Gif-sur-Yvette, France
From the symposium ‘‘Insights of Early Chordate Genomics: Endocrinology and Development in Amphioxus, Tunicates
and Lampreys’’ presented at the annual meeting of the Society for Integrative and Comparative Biology, January 3–7,
2010, at Seattle, Washington.
1E-mail: [email protected]
Synopsis Lampreys are agnathans (vertebrates without jaws). They occupy a key phylogenetic position in the emergence
of novelties and in the diversification of morphology at the dawn of vertebrates. We have used lampreys to investigate the
possibility that embryonic midline signaling systems have been a driving force for the evolution of the forebrain in
vertebrates. We have focused on Sonic Hedgehog/Hedgehog (Shh/Hh) signaling. In this article, we first review and sum-
marize our recent work on the comparative analysis of embryonic expression patterns for Shh/Hh, together with Fgf8
(fibroblast growth factor 8) and Wnt (wingless-Int) pathway components, in the embryonic lamprey forebrain.
Comparison with nonvertebrate chordates on one hand, and jawed vertebrates on the other hand, shows that these
morphogens/growth factors acquired new expression domains in the most rostral part of the neural tube in lampreys
compared to nonvertebrate chordates, and in jawed vertebrates compared to lampreys. These data are consistent with the
idea that changes in Shh, Fgf8 or Wnt signaling in the course of evolution have been instrumental for the emergence and
diversification of the telencephalon, a part of the forebrain that is unique to vertebrates. We have then used comparative
genomics on Shh/Hh loci to identify commonalities and differences in noncoding regulatory sequences across species and
phyla. Conserved noncoding elements (CNEs) can be detected in lamprey Hh introns, even though they display unique
structural features and need adjustments of parameters used for in silico alignments to be detected, because of
lamprey-specific properties of the genome. The data also show conservation of a ventral midline enhancer located in
Shh/Hh intron 2 of all chordates, the very species which possess a notochord and a floor plate, but not in earlier emerged
deuterostomes or protostomes. These findings exemplify how the Shh/Hh locus is one of the best loci to study genome
evolution with regards to developmental events.
The telencephalon is a vertebratenovelty
The vertebrate forebrain is an amazingly sophisticat-
ed part of the nervous system. It is composed of the
telencephalon (including the pallium or cortex and
the basal ganglia) and the diencephalon (including
the thalamus and hypothalamus). During embryo-
genesis, the forebrain is generated from the
anterior-most part of the neural plate and neural
tube through complex morphogenetic events
(Fig. 1). Meanwhile, waves of tightly controlled pro-
liferation, neurogenesis, specification, migration, and
axonal outgrowth generate thousands of types of
neurones that are cytoarchitectonically organized
and topographically interconnected (for reviews see
Wilson and Rubenstein, 2000; Guillemot, 2005).
Among vertebrates, the forebrain is also the part of
the brain that has undergone the most dramatic
morphological and anatomical diversification, this
being particularly true when considering the most
rostral and dorsal part, the telencephalon. Would
anyone have thought, for example, that the mamma-
lian cerebral cortex, with its well-known organization
into six layers, could be homologous with the everted
dorsal pallium of teleost fishes?
Ten to fifteen years of ‘‘evo-devo’’ studies using
comparative expression-pattern analysis for develop-
mental genes have shown that the forebrain, includ-
ing the telencephalon of various vertebrate species,
Integrative and Comparative Biology, volume 50, number 1, pp. 98–109
doi:10.1093/icb/icq032
Advanced Access publication May 11, 2010
� The Author 2010. Published by Oxford University Press on behalf of the Society for Integrative and Comparative Biology. All rights reserved.
For permissions please email: [email protected] from https://academic.oup.com/icb/article-abstract/50/1/98/733550by gueston 09 April 2018

Fig. 1 Development and evolution of signaling systems in the chordate forebrain. (A) Summary of the development of the forebrain in
a generic jawed vertebrate. Left: at neural plate stage, a fate map of the anterior neural plate shows a region destined to become the
telencephalon (tel, pink and green domains) and diencephalon (di, blue domains). The anterior neural ridge (ANR) and notochord (red)
are indicated. Middle: after closure of the neural tube, the forebrain develops under the influence of signaling molecules secreted from
the ventral side (Shh, red), the dorsal midline (Wnts and Bmps, brown) and the rostral pole (Fgf, blue). They generate a field of
organization in the adjacent neuroepithelium and control growth and morphogenesis. Right: regionalization of the forebrain as a
consequence of the concerted action of signaling centers. The telencephalon (tel) is further subdivided into the subpallial regions
(pink shades) and pallial areas (green shades) and the diencephalon (di) is subdivided into several tranverse domains including the
thalamus (thal) and the hypothalamus (hyp), mes, mesencephalon; is, isthmus; cb, cerebellum; met, metencephalon. (B) Evolution of the
expression of Hedgehog, Fgf8 and SFPRs in chordates. A simplified phylogenetic tree of chordates shows drawings of embryonic
expression of Shh/Hh (red), Fgf8 (blue), and SFRP1/5 (green) in representative species. The drawing of amphioxus is compiled from data
from Lin et al. (2009), Onai et al. (2009), Shimeld (1999), and Shimeld et al. (2007). The drawing of Ciona is compiled from data from
the aniseed database (Imai et al. 2002; Takatori et al. 2002). The drawing of the lamprey is from Guerin et al. (2009) and Osorio et al.
(2005). The typical gnathostome is represented by a zebrafish brain and is compiled from the ZFIN database. fp, floorplate; hb,
hindbrain; hypoth, hypothalamus; mhb, mid-hindbrain boundary; no, notochord; p, pineal gland; pcp, prechordal plate; tel, telencephalon;
zli, zona limitans intrathalamica.
Hedgehog and chordate brain evolution 99
Downloaded from https://academic.oup.com/icb/article-abstract/50/1/98/733550by gueston 09 April 2018

are specified and built according to shared genetic
mechanisms (e.g., Puelles et al. 2000; Bachy et al.
2002). In contrast, the sister group of vertebrates,
the urochordates, represented by ascidians (Delsuc
et al. 2006), together with the earlier-emerged
group of cephalochordates, represented by amphiox-
us, do not possess a telencephalon. The anterior tip
of their central nervous system is composed of a
sensory vesicle (ascidians) or a cerebral vesicle
(amphioxus). When comparing the patterning of
the anterior neural tube among the three groups
of chordates, one observes similarities that are
likely inherited from their common ancestor. For
example Otx expression is a hallmark of the anterior
neural tube in all chordates (Wada et al. 1998). In
the ascidian, Ciona intestinalis, detailed patterning
analysis has led to the proposal that the anterior
ventral neural tube corresponds to the vertebrate
presumptive hypothalamus (Moret et al. 2005), i.e.,
a diencephalic, but not telencephalic, region of the
brain.
The telencephalon appears to be a vertebrate
novelty (or synapomorphy). In the course of our
studies, we have used the evolutionary developmental
approach to examine the possible genetic mecha-
nisms that may lie at the origin of the vertebrate
forebrain. To this end, we have used lampreys as
model animals, because they occupy a very crucial
phylogenetic position in the chordate tree, i.e., they
belong to agnathans or cyclostomes, the sister group
of jawed vertebrates (Osorio and Retaux 2008).
Any character that is shared by lampreys and other
vertebrates but not by urochordates or cephalochor-
dates is therefore potentially relevant for the descrip-
tion of the common ancestor of all vertebrates.
Hence, we may be able to at least have a better pic-
ture of what has happened �500-million years ago
(mya), when this common ancestor experienced the
emergence and swelling of a part of the nervous
system at the tip of its dorsal (alar) neural tube: a
telencephalon.
Midline signaling as a driver offorebrain evolution
During embryogenesis, the brain develops under the
influence of several signaling centers, sometimes
called secondary organizers (Fig. 1A). These centers
secrete diffusible molecules with morphogen proper-
ties, which control growth and patterning and gen-
erate a field of organization in the adjacent
neuroepithelium [recently reviewed by Vieira et al.
(2010)]. The morphogen gradients are later translat-
ed into discrete neuroepithelial domains or divisions
that correspond to the future functional units of the
mature brain. Therefore, any change in the time of
appearance, in the strength, or in the exact location
of such secondary organizers has the potential of
significantly influencing the size or shape or pattern-
ing, in other words the neuroanatomy, of the brain
region they ‘‘organize’’.
A number of the signaling centers mentioned
above are located at a midline position respective
to the neural tube. Ventrally, the notochord and pre-
chordal plate (of mesodermal origin) and the in-
duced floor plate of the neural tube secrete Sonic
Hedgehog (Shh) (Echelard et al. 1993). At
mid-embryogenesis, new areas of Shh expression
appear in the forebrain, always emanating from the
ventral midline: the zona limitans intrathalamica (zli)
and the ventral telencephalon (Kiecker and Lumsden
2004; Scholpp et al. 2006; Vieira and Martinez 2006).
Dorsally, the roof plate of the neural tube secretes
molecules of the wingless-int (Wnt) (Lee and Jessell
1999; Muroyama et al. 2002) and bone morphoge-
netic protein (Bmp) families (Liem et al. 1997). Wnt
ligands are also secreted from the mid-diencephalon.
Finally, at the rostral tip of the neural tube, the an-
terior neural ridge (ANR) and later the rostral telen-
cephalon produce fibroblast growth factor 8 (Fgf8,
also expressed at the mid-hindbrain organizer)
(Shimamura and Rubenstein 1997; Rubenstein et al.
1998).
In this article, we first review and summarize our
recent work on the comparative analysis of embry-
onic expression patterns for Shh/Hh, together with
Fgf8 and Wnt pathway components, in the embryon-
ic lamprey forebrain. We then use comparative ge-
nomics on Shh/Hh loci to investigate whether
changes in noncoding regulatory sequences can ac-
count for the observed changes in Shh/Hh expression
across species. Our findings exemplify how the
Shh/Hh locus is an ideal locus to study the evolution
of the genome with regards to developmental events.
Hedgehog, Fgf8, and Wnt pathwayexpression patterns in lampreys
We have carried out a medium-scale in situ hybrid-
ization screen for genes expressed in the forebrain of
lamprey embryos (Guerin et al. 2009). Genes in-
volved in proliferation, stemcellness, neurogenesis,
and regional patterning showed globally highly sim-
ilar expression when compared to their gnathostome
counterparts, suggesting that the basic mechanisms
governing these events are shared among all verte-
brates. Yet strikingly, we observed major differences
100 S. Retaux and S. Kano
Downloaded from https://academic.oup.com/icb/article-abstract/50/1/98/733550by gueston 09 April 2018

when considering the midline signaling systems
(summarized in Fig. 1B):
(1) The two lamprey Hedgehogs (Hh) (Osorio et al.
2005; J.-H. Xiao and Sylvie Retaux, unpublished
data) were expressed in the notochord and floor
plate, in the zli (for only one of the lamprey’s
Hhs), and in the hypothalamus. None of them
was expressed, even at a late amnocoete larval
stage, in the ventral telencephalon.
(2) Lamprey Fgf8 was classically expressed at the
mid-hindbrain boundary together with some ad-
ditional expression areas in the diencephalon,
also present in jawed vertebrates. Yet it was
not expressed at the telencephalic tip before a
late larval stage.
(3) Lamprey Wnt pathway components such as Wnt
ligands, their frizzled receptors, and their SFRP
inhibitors were also studied. Major differences
were observed for the SFRPs: lamprey SFRP1/5
expression was restricted to the hypothalamus,
whereas gnathostome orthologs expression also
expands into the telencephalon.
The differences in expression of the Hh, Fgf8 and
SFRP signaling pathways components in lampreys
and jawed vertebrates summarized above concern
mainly the telencephalon, whereas other expression
domains (diencephalic or mesencephalic) are con-
served (Guerin et al. 2009) (Fig. 1B). These differ-
ences are interesting to discuss in regard to the
peculiar anatomy of the lamprey telencephalon and
to the evolution of the telencephalon in vertebrates.
The lack of Hh signal in the telencephalon, togeth-
er with the absence of expression of Nkx2.1 in the
same area (Murakami et al. 2001; Osorio et al. 2005),
correlates with the absence of a pallidal division in
the ventral telencephalon of lampreys (Weigle and
Northcutt 1999). In a sense, the mouse Nkx2.1�/�
(knock-out) phenotype mimics the lampreys’ condi-
tion, with a ventral telencephalon that is almost ex-
clusively composed of a striatal division (Sussel et al.
1999).
There is heterochrony of Fgf8 expression in lam-
preys and it is tempting to relate the lack of Fgf8
signal at the rostral tip of the telencephalon during
the first 10 days of development to the extremely
slow growth of the telencephalon in embryonic lam-
preys. Indeed, whereas the gnathostome telencephalic
vesicles undergo considerable growth and swelling
during the early phases of embryogenesis, lamprey
telencephalon remains very tiny and starts growing
significantly only in the larval stages (Villar-Cheda
et al. 2006). It is probable that other Fgfs, such as
Fgf3, which has a complementary role to Fgf8 in fish
(Walshe and Mason 2003), are expressed in the lam-
prey telencephalon. Moreover, as important and re-
ciprocal interactions occur between the ventral (Shh)
and rostral (Fgf8) signaling centers and shape the
gnathostome telencephalon (e.g., Kuschel et al.
2003), it is probable that the absence of the former
and the late appearance of the latter influence the
patterning and morphogenesis of this region in lam-
preys considerably. These differences are also intrigu-
ing and raise questions on how the reciprocal
transcriptional regulations these pathways exert on
each other are controlled in lampreys.
Inhibition of the Wnt pathway by SFRP1/5 does
not occur in the lamprey telencephalon. This is also
surprising because in jawed vertebrates SFRP1 and
SFRP5 (and the fish-specific Tlc) (Tendeng and
Houart 2006) are in major part responsible for the
inhibition of Wnt signals emanating from the dien-
cephalon, and serve in determining the fate of the
anterior telencephalon (Houart et al. 2002). As lam-
preys do have a telencephalon, albeit very small, it is
highly probable that other factors involved in inhi-
bition of the Wnt signal are present in the embryonic
lamprey’s forebrain. They remain to be discovered,
and, once again, will likely highlight the distinctive
manner in which lampreys regulate and control the
specification and growth of their anterior neural
tube.
The hypotheses driven by the comparative ap-
proach await functional validation. Within this
framework, and in an attempt to mimic the gnathos-
tome situation in the lamprey telencephalon, we have
injected mammalian Shh protein into the basal
telencephalon of stage-24 lamprey embryos.
Unfortunately, this did not result in the ectopic in-
duction of Nkx2.1 in the subpallium (J. Osorio and
S.R., unpublished data), nor in any particular phe-
notype. More experiments interfering with the mid-
line signaling pathways in lampreys are needed to
understand the interactions between these pathways
and the effects they produce.
The data reported above would also strongly ben-
efit from a similar analysis in hagfish, the only other
extant agnathan/cyclostome representative. The diffi-
culty in obtaining hagfish embryos has long ham-
pered the study of embryology in this species, but
the recent success reported in breeding these animals
has generated hope for future years (Ota et al. 2007).
It will be crucial to know whether hagfish embryos
‘‘behave’’ like lamprey embryos in terms of telence-
phalic transcriptome and associated gene regulatory
networks. For example, if hagfish do not express Shh
in their ventral telencephalon, it will further reinforce
Hedgehog and chordate brain evolution 101
Downloaded from https://academic.oup.com/icb/article-abstract/50/1/98/733550by gueston 09 April 2018

the idea that Hh/Shh signaling has been recruited to
the telencephalon in the ancestor of jawed verte-
brates. Alternatively, if hagfish embryos do express
Shh in their telencephalon, it will substantiate a
loss of Hh expression during lamprey evolution,
and will suggest that the recruitment of Hh in the
basal telencephalon happened in the common ances-
tor of all vertebrates.
Comparing midline signaling in basal chordates
A larger and general view can be obtained by further
comparing conditions between vertebrate and non-
vertebrate chordates. A search of the literature and ex-
pression databases (http://crfb.univ-mrs.fr/aniseed/)
for ascidians and amphioxus for reported midline
components resulted in the schematic summary in
Fig. 1B. In these two nonvertebrate chordate repre-
sentatives, Hh expression is certainly ventral (in the
ventral nerve cord and notochord in amphioxus
only) but it is restricted to the posterior nervous
system, in regions that are comparable to the hind-
brain/spinal cord patterning domains of vertebrates
(Shimeld 1999; Takatori et al. 2002). In addition, in
Ciona, Hh1-expressing cells are found at the tip of
the sensory vesicle. Later on, Hh2 is partially
co-localized with Nkx2.1 (Ristoratore et al. 1999;
Islam et al. 2010). In amphioxus the expression of
Fgf8 spreads throughout the anterior ectoderm, like
Otx (Meulemans and Bronner-Fraser 2007; Holland
2009). In Ciona, two or three Fgf8-expressing cells
were reported in a position corresponding to the
junction between the sensory vesicle and the posteri-
or nerve cord (Ikuta and Saiga 2007), thus
pre-figuring a possible anterior–posterior brain
boundary, which is exemplified by the vertebrate
mid-hindbrain boundary (Imai et al. 2009). Finally,
SFRP1/5 in Ciona is not expressed in the anterior
sensory vesicle, while amphioxus SFRP1/5 has not
been identified to date.
When comparing these organizers across chor-
dates, it appears that they become localized more
and more anteriorly in the neural tube as the subdi-
visions of the forebrain become larger and more nu-
merous, and as the telencephalon emerges and
expands (Fig. 1B). This is particularly true for
Shh/Hh and Fgf8 which have complex expression
patterns composed of multiple expression areas.
This tendency correlates well with the two main
roles of these signaling centers and the molecules
they produce: they organize and control the growth
of the neuroepithelium. As such, they are ideally
placed to function as drivers of brain evolution,
both for the emergence of novelties and for the di-
versification of structures.
Transcriptional regulation of Shh/Hhmidline signaling
As a next step in the understanding of the molecular
mechanisms of the evolution of the forebrain, we
studied transcriptional regulation of the genes
coding for morphogen factors. Indeed, subtle differ-
ences in expression domains or in the timing of tran-
script expression mostly originate in noncoding
regulatory sequences. As a case study, we choose
the Shh/Hh gene/locus.
The regulatory logics of Shh transcription has
been the focus of many studies in gnathostomes.
Using phylogenetic footprinting, it is possible to
align and compare Shh loci, and to detect stretches
of highly conserved sequences between species
(Goode et al. 2003, 2005; Jeong et al. 2006; Ertzer
et al. 2007; see also Strahle and Rastegar, 2008 for
review). Highly conserved sequences often corre-
spond to exon-coding fragments, but also reside in
noncoding intronic sequences or in 50- or 30-gene
regions (Fig. 2A). The fact that these noncoding se-
quences are highly conserved—sometimes even
better than coding sequences themselves—suggests
that they are under strong selective pressure, and
that they contain crucial regulatory elements, includ-
ing binding sites for transcription factor (Vavouri
and Lehner, 2009).
In the case of gnathostome Shh genes, the align-
ment and VISTA visualization of conserved noncod-
ing elements (CNEs) can be easily performed
(Fig. 2A, lines 1–4; see also Appendix 1). A
number of CNEs have been previously identified by
this method, and have been functionally tested for
enhancer activity in the mouse or the zebrafish
(Jeong et al. 2006; Ertzer et al. 2007). Hence, the
regulation of Shh expression appears highly modular,
with specific enhancers governing expression in the
notochord, the floor plate, the zli, the hypothalamus,
and the telencephalon. Of note, this feature of the
Shh locus is highly favorable in the context of our
hypothesis of Hh signaling as a driver of brain evo-
lution: the modular nature of cis-regulatory elements
and the pleiotropy of gene products are among the
‘‘evolutionary toolkit’’ that allows for selective
spatio-temporal changes of expression patterns, and
thus of morphological changes (Carroll, 2008).
Interestingly, recent studies have shown that this
modular nature of enhancers also applies to the
Fgf8 locus (Komisarczuk et al. 2009).
102 S. Retaux and S. Kano
Downloaded from https://academic.oup.com/icb/article-abstract/50/1/98/733550by gueston 09 April 2018

Fig. 2 Comparison of Shh/Hh loci across vertebrates. (A) VISTA alignment and search for CNEs in several vertebrates (mouse, chick,
zebrafish, medaka, and lamprey) using the Fugu Shh locus as baseline. The zebrafish intronic elements previously described (ar-A and
ar-B in intron 1 and ar-C in intron 2 (Muller et al. 1999; Ertzer et al. 2007) were added to identify the conserved peaks. In this and the
following plots, gray shading indicates more than 75% sequence identity. The exons (E1–E3) appear as rectangles (gray peaks on the
plots), and non-coding conserved sequences appear as ovals (black peaks on the plots). ar-X stands for ‘activating region’ X. (B) VISTA
alignment and search for CNEs in zebrafish and lamprey. When using the Fugu Shh locus as baseline, the ar-C element in intron 2
appears as a single peak of conservation (black arrowhead). (C) VISTA alignment and search for CNEs in zebrafish and Fugu, using the
lamprey Hhb locus as baseline. The ar-C element in intron 2 appears dispersed into four peaks (black arrows). (D) Local alignment
showing lamprey, zebrafish and Fugu ar-C nucleotide sequences, illustrating the dispersed nature of lamprey ar-C. The four
sub-elements ar-C1 to ar-C4 identified in zebrafish by Hadzhiev et al. (2007) are dispersed along intron 2 in a manner that corresponds
to the four conservation peaks detected in C (first peak: beginning of ar-C; second peak: C1; third peak: C2; fourth peak: end of
C2 þ C3 þ C4). Note that ar-C4 in Fugu also contains an insertion. The full alignment can be seen in Supplementary Fig. 1.
Hedgehog and chordate brain evolution 103
Downloaded from https://academic.oup.com/icb/article-abstract/50/1/98/733550by gueston 09 April 2018

Hh regulatory logics in lampreys
Through in silico searches in the sea lamprey
Petromyzon marinus (Pm) preliminary genome assem-
bly (http://pre.ensembl.org/Petromyzon_marinus/
Info/Index), and through cosmid library screening
and sequencing in the river lamprey Lampetra fluvia-
tilis (Lf), we have obtained genomic sequences for
two distinct Hh genes in the two lamprey species.
We have called them Hha and Hhb (S. Kano et al.
unpublished data). They are both Shh-like in terms of
molecular phylogeny and expression patterns, and
therefore allowed us to perform a comparative geno-
mic analysis with their gnathostome orthologous
Shh locus. We have recently analyzed and reported
CNEs in lamprey Hhs by comparisons based on the
coelacanth Shh locus (S. Kano et al. unpublished
data). Here we present the same approach for
Lampetra Hhb (LfHhb), and use the Fugu genome
as baseline for comparison, i.e., complete genome
that allows more systematic comparisons and that
has proven very helpful in the past (Goode et al.
2003, 2005).
Using the Fugu Shh locus as baseline, previously
identified intronic motifs ar-A and ar-C were easily
identified among gnathostomes (Fig. 2A). Other
motifs, ar-B in intron 1 and ar-D in the 50-upstream
region, were less conserved, although a few conserved
blocks can be detected among teleost species for the
ar-B motif, originally identified in zebrafish (Muller
et al. 1999; Ertzer et al. 2007). When compared with
gnathostome loci, almost no significant CNEs are
found at first glance in the Lampetra fluviatilis
LfHhb locus, besides the three exons (Fig. 2A, fifth
line). However, the use of a more limited number of
species’ loci for comparison allowed the detection of
conserved blocks and significant CNEs in lamprey
(Fig. 3B). More specifically, these include ar-C [the
‘‘midline’’ enhancer described in mice and zebrafish
(Jeong et al. 2006; Ertzer et al. 2007; Hadzhiev et al.
2007)] and ar-D (an additional floorplate enhancer
also active both in mice and zebrafish) (Jeong et al.
2006; Ertzer et al. 2007).
When mapped onto the Fugu Shh locus, the
ar-C element emerges as a single conservation peak
Fig. 3 comparison of Shh/Hh loci among deuterostomes. (A) Each Shh/Hh locus was independently compared with Fugu and zebrafish
because global alignments performed together disturb proper alignment, as mentioned above (data not shown). The presentation is the
same as in Fig. 2. (B) Local alignment of the ar-C motif shared among chordates. The conserved sequence shared among chordates
corresponds to the ar-C1 sub-element. The position of a consensus FoxA2 binding site is indicated.
104 S. Retaux and S. Kano
Downloaded from https://academic.oup.com/icb/article-abstract/50/1/98/733550by gueston 09 April 2018

(black arrowhead in Fig. 2B). However, when the
reverse in silico experiment is done, i.e., mapping
the Fugu or the zebrafish loci onto the lamprey
LfHhb used as baseline, ar-C emerges as four indi-
vidual peaks in intron 2 (arrows in Fig. 2C). This
demonstrates that ar-C is split into several conserved
subelements or blocks in the lamprey locus. These
conserved blocks can be readily observed on local
alignments (Fig. 2D and Supplementary Fig. 1). In
particular, a �1 kb-long insertion occurs between
ar-C1 and the rest of ar-C. The lamprey C1
sub-element was further validated as a midline en-
hancer responsible for notochord and floor-plate ex-
pression after injections in zebrafish embryos
(S. Kano et al. unpublished data). These findings
show and confirm that functional CNEs are found
in the lamprey genome, as first reported for Hox
genes (Carr et al. 1998; Irvine et al. 2002) and
more recently in a survey of 13 gene loci (McEwen
et al. 2009).
These data also raise two issues. (1) The first is an
alignment issue when comparing the diverged lam-
prey genome with other vertebrate genomes. When
carefully checking local alignments, the lamprey
genome appears to contain lamprey-specific inser-
tions in the introns. In fact, it was difficult to tune
alignments when both Hha and Hhb loci were in-
cluded together with vertebrates because the two
lamprey paralogs evolved independently, and the
paralog-specific insertions disturb alignments
(better results are also obtained on the Hha locus
when it is analyzed independently from Hhb; not
shown). This may well explain why fewer CNEs
were found in lamprey genes than in other verte-
brates (McEwen et al. 2009). (2) The second is a
window-size issue for MLAGAN and VISTA analysis.
We have used a 20-bp window size and a 70% iden-
tity cut-off in our analysis whereas McEwen et al.
(2009) have used a window size of 40 bp and a
cut-off of 65% identity. Using these parameters, we
were able to dissect out a �30 bp lamprey ar-C1.
Thus, a narrow window size is advisable when sur-
veying CNEs in the lamprey genome.
Survey of CNEs shared amongchordates and deuterostomes inShh/Hh loci
The lamprey CNEs are likely to be essential for Hh
expression. As CNEs frequently contain regulatory
elements (Vavouri et al. 2007; Vavouri and Lehner,
2009), and as Hh genes are expressed at least in part
in similar domains in vertebrate and nonvertebrate
chordates, we have pursued a similar comparative
genomics approach on other chordates and deutero-
stomes. Of note, some CNEs were recently identified
in amphioxus and showed enhancer activity in zeb-
rafish (Hufton et al. 2009).
We compared Hh loci between fugu and three
other gnathostomes (zebrafish and two amniotes
species: chick and mouse), a jawless vertebrate
(lamprey), two nonvertebrate chordates (ascidians
and amphioxus), a hemichordate (acorn worm), an
echinoderm (sea urchin), and a protostome (fly)
(Fig. 3A). Conservation peaks were detected even
though the degree of conservation was moderate in
most cases (see also Fig. 4). Interestingly, the amphi-
oxus Hh locus displayed highest conservation with
fish genomes (all the intronic peaks are colored in
pink). Putative amphioxus ar-A, ar-B, and ar-C
motifs showed significant conservation but were
shorter than vertebrate CNEs. It is reasonable to
think that the important conservation found between
amphioxus and vertebrates reflects the fact that am-
phioxus’ body plan and Hh expression pattern re-
semble those of vertebrates (see also Holland et al.
2008; Holland, 2009).
We found that the intron 2 ar-C motif is well
shared only among chordates (Fig. 3A). Further, it
was possible to identify a ‘‘core’’ ar-C1 motif, pro-
posed by Hadzhiev et al. (2007) to be the major
functional sub-element of ar-C, in all the chordate
Hh loci examined (Fig. 3B). This raises the
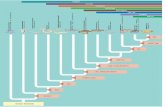
Fig. 4 Hh CNEs in bilaterians. A simplified tree of bilaterians
highlights the bilaterian origin of Hh CNEs (dotted line), and the
probable emergence and fixation of a midline ar-C motif in
chordates. The shaded grey points to the interesting phylogenetic
position of lampreys which possess a telencephalon and whose
genome has undergone whole-genome duplications (WGD;
Kuraku et al., 2008).
Hedgehog and chordate brain evolution 105
Downloaded from https://academic.oup.com/icb/article-abstract/50/1/98/733550by gueston 09 April 2018

interesting possibility that ar-C is a key motif for the
chordate lineage (see also Fig. 4). Indeed, ar-C is
notably responsible for notochord and floorplate ex-
pression in these animals which all possess a noto-
chord and a floor plate as a defining feature, e.g., in
mice (Jeong et al. 2006), in zebrafish (Hadzhiev et al.
2007), and in lamprey (S. Kano et al. unpublished
data). In addition, it is worth mentioning that the
ar-C1 element contains a consensus FoxA2 transcrip-
tion factor binding site (Fig. 3B). This is interesting
in regards of the described implication of this factor,
expressed in the floor plate, in the regulation of Shh
expression at the ventral midline (for review, Placzek
and Briscoe, 2005).
Surprisingly, our analysis also revealed conserved
peaks in the introns of Saccoglossus, Strongylocentrus
and Drosophila, although Hh expression in these an-
imals is not comparable to chordates. The evolution
of Hh genes has been the focus of much interest, and
their origins have been traced to the first bilaterians
(Adamska et al. 2007). Our data on ‘‘signatures’’ re-
maining on every Shh/Hh intron suggest that the Hh
introns have been maintained since bilaterians ap-
peared, and further highlight this evolutionary sce-
nario. They are also in line with the idea that intron
gain is an extremely rare event in vertebrate evolu-
tion (Loh et al. 2008), so that most introns are of
ancestral origin (Fig. 4).
The primary function of ar-C is up-regulation at
the midline. This element was apparently recruited
into the midline of the tail (i.e., notochord and
floorplate) in early chordates. Additional regulatory
elements must have further emerged to drive Shh/Hh
expression in new domains—such as the forebrain
Shh expression domains observed in jawed verte-
brates. Of note, repressor elements also appeared,
such as zebrafish ar-C2 and ar-C4 which act as
floor-plate repressors (Hadzhiev et al. 2007). This
hypothesis needs further investigation on CNEs, sim-
ilar to those presented here.
Conclusions
Shh/Hh is known as one of the most powerful mor-
phogens during embryonic development. As it is par-
ticularly involved in controlling morphogenesis,
patterning and growth of the neural tube, we have
hypothesized that variations of its expression pattern
during embryogenesis may be an evolutionary driv-
ing force, causing changes in size and/or morpholo-
gy. Using lamprey as a key phylogenetically-placed
model animal, we have examined this possibility at
two levels: comparative expression patterns and com-
parative genomics.
A recent survey of 13 lamprey genes for CNEs by
McEwen et al. (2009) demonstrated that some CNEs
were indeed identified and validated; however, their
number was much less than expected. While a
large-scale analysis is now feasible and required in
the genome era, deep insight from some particular
cases are still essential. The Hedgehog locus is one of
best models to associate genomic evolutionary events
with developmental mechanisms, because it has been
deeply studied in several model animals, broadly
distributed phylogenetically. Here, we add a new
case study of lamprey and other deuterostome
Hedgehog loci.
Supplementary Data
Supplementary Data are available at ICB online.
Acknowledgments
We thank our colleagues, collaborators and friends
Jean-Stephane Joly, Sylvie Mazan, and Didier Casane,
and the other group members previously involved in
lamprey work Joana Osorio, Adele Guerin, and Jin-
Hua Xiao, for long-term and fertile scientific
interactions.
Funding
Work supported by a Groupement d’Interet
Scientifique (GIS) Genomique Marine and Agence
Nationale de la Recherche ANR-Neuro (MIDLINE)
(to S.R.). S.K. was supported by an ANR postdoc-
toral fellowship.
References
Adamska M, Matus DQ, Adamski M, Green K, Rokhsar DS,
Martindale MQ, Degnan BM. 2007. The evolutionary origin
of hedgehog proteins. Curr Biol 17:R836–7.
Bachy I, Berthon J, Retaux S. 2002. Defining pallial and sub-
pallial divisions in the developing Xenopus forebrain. Mech
Dev 117:163–72.
Brudno M, Do CB, Cooper GM, Kim MF, Davydov E,
Program NCS, Green ED, Sidow A, Batzoglou S. 2003.
LAGAN and Multi-LAGAN: Efficient tools for large-scale
multiple alignment of genomic DNA. Genome Res 13:
721–31.
Carr JL, Shashikant CS, Bailey WJ, Ruddle FH. 1998.
Molecular evolution of Hox gene regulation: Cloning and
transgenic analysis of the lamprey HoxQ8 gene. J Exp Zool
280:73–85.
Carroll SB. 2008. Evo-devo and an expanding evolutionary
synthesis: A genetic theory of morphological evolution.
Cell 134:25–36.
Delsuc F, Brinkmann H, Chourrout D, Philippe H. 2006.
Tunicates and not cephalochordates are the closest living
relatives of vertebrates. Nature 439:965–8.
106 S. Retaux and S. Kano
Downloaded from https://academic.oup.com/icb/article-abstract/50/1/98/733550by gueston 09 April 2018

Echelard Y, Epstein DJ, St-Jacques B, Shen L, Mohler J,
McMahon JA, McMahon AP. 1993. Sonic hedgehog, a
member of a family of putative signaling molecules, is
implicated in the regulation of CNS polarity. Cell 75:
1417–30.
Ertzer R, Muller F, Hadzhiev Y, Rathnam S, Fischer N,
Rastegar S, Strahle U. 2007. Cooperation of sonic hedgehog
enhancers in midline expression. Dev Biol 301:578–89.
Frazer KA, Pachter L, Poliakov A, Rubin EM, Dubchak I.
2004. VISTA: computational tools for comparative geno-
mics. Nucleic Acids Res 32:W273–9.
Goode DK, Snell P, Elgar G. 2003. Comparative analysis of
vertebrate Shh genes identifies novel conserved non-coding
sequence. Mamm Genome 14:192–201.
Goode DK, Snell P, Smith SF, Cooke JE, Elgar G. 2005.
Highly conserved regulatory elements around the SHH
gene may contribute to the maintenance of conserved syn-
teny across human chromosome 7q36.3. Genomics 86:
172–81.
Guerin A, d’Aubenton-Carafa Y, Marrakchi E, Da Silva C,
Wincker P, Mazan S, Retaux S. 2009. Neurodevelopment
genes in lampreys reveal trends for forebrain evolution in
craniates. PLoS One 4:e5374.
Guillemot F. 2005. Cellular and molecular control of neuro-
genesis in the mammalian telencephalon. Curr Opin Cell
Biol 17:639–47.
Hadzhiev Y, Lang M, Ertzer R, Meyer A, Strahle U, Muller F.
2007. Functional diversification of sonic hedgehog paralog
enhancers identified by phylogenomic reconstruction.
Genome Biol 8:R106.
Holland LZ. 2009. Chordate roots of the vertebrate nervous
system: Expanding the molecular toolkit. Nat Rev Neurosci
10:736–46.
Holland LZ, et al. 2008. The amphioxus genome illuminates
vertebrate origins and cephalochordate biology. Genome
Res 18:1100–11.
Houart C, Caneparo L, Heisenberg C, Barth K, Take-Uchi M,
Wilson S. 2002. Establishment of the telencephalon during
gastrulation by local antagonism of Wnt signaling. Neuron
35:255–65.
Hufton AL, Mathia S, Braun H, Georgi U, Lehrach H,
Vingron M, Poustka AJ, Panopoulou G. 2009. Deeply con-
served chordate noncoding sequences preserve genome syn-
teny but do not drive gene duplicate retention. Genome
Res 19:2036–51.
Ikuta T, Saiga H. 2007. Dynamic change in the expression of
developmental genes in the ascidian central nervous system:
revisit to the tripartite model and the origin of the mid-
brain-hindbrain boundary region. Dev Biol 312: 631–43.
Imai KS, Satoh N, Satou Y. 2002. Region specific gene expres-
sions in the central nervous system of the ascidian embryo.
Gene Expr Patterns 2:319–21.
Imai KS, Stolfi A, Levine M, Satou Y. 2009. Gene regulatory
networks underlying the compartmentalization of the
Ciona central nervous system. Development 136:285–93.
Irvine SQ, Carr JL, Bailey WJ, Kawasaki K, Shimizu N,
Amemiya CT, Ruddle FH. 2002. Genomic analysis of Hox
clusters in the sea lamprey Petromyzon marinus. J Exp
Zool 294:47–62.
Islam A, Moly P, Miyamot Y, Kusakabe T. 2010. Distinctive
expression patterns of Hedgehog pathway genes in the
ciona intestinalis larva: implications for a role of
Hedgehog signaling in postembryonic development and
chordate evolution. Zoolog Sci 27:84–90.
Jeong Y, El-Jaick K, Roessler E, Muenke M, Epstein DJ. 2006.
A functional screen for sonic hedgehog regulatory elements
across a 1 Mb interval identifies long-range ventral fore-
brain enhancers. Development 133:761–72.
Kiecker C, Lumsden A. 2004. Hedgehog signaling from the
ZLI regulates diencephalic regional identity. Nat Neurosci
7:1242–9.
Komisarczuk AZ, Kawakami K, Becker TS. 2009. Cis-regula-
tion and chromosomal rearrangement of the fgf8 locus after
the teleost/tetrapod split. Dev Biol 336:301–12.
Kuraku S, Meyer A, Kuratani S. 2008. Timing of genome
duplications relative to the origin of the vertebrates:
did cyclostomes diverge before, or after? Mol Biol Evol
26:47–59.
Kuschel S, Ruther U, Theil T. 2003. A disrupted balance
between Bmp/Wnt and Fgf signaling underlies the ventra-
lization of the Gli3 mutant telencephalon. Dev Biol 260:
484–95.
Lee KJ, Jessell TM. 1999. The specification of dorsal cell fates
in the vertebrate central nervous system. Annu Rev
Neurosci 22:261–94.
Liem KF Jr, Tremml G, Jessell TM. 1997. A role for the roof
plate and its resident TGFbeta-related proteins in neuronal
patterning in the dorsal spinal cord. Cell 91:127–38.
Lin Y, Cai Z, Huang S, Yang L, Wang C, Liu Z, Cao J, An Y,
Zhang H. 2009. Ptc, Smo, Sufu, and the Hedgehog signal-
ing pathway in amphioxus. Evol Dev 11:710–8.
Loh YH, Brenner S, Venkatesh B. 2008. Investigation
of loss and gain of introns in the compact genomes of
pufferfishes (Fugu and Tetraodon). Mol Biol Evol 25:
526–35.
Mayor C, Brudno M, Schwartz JR, Poliakov A, Rubin E M,
Frazer KA, Pachter LS, Dubchak I. 2000. VISTA:
Visualizing global DNA sequence alignments of arbitrary
length. Bioinformatics 16:1046–7.
McEwen GK, Goode DK, Parker HJ, Woolfe A, Callaway H,
Elgar G. 2009. Early evolution of conserved regulatory
sequences associated with development in vertebrates.
PLoS Genet 5:e1000762.
Meulemans D, Bronner-Fraser M. 2007. Insights from
amphioxus into the evolution of vertebrate cartilage.
PLoS One 2:e787.
Moret F, Christiaen L, Deyts C, Blin M, Vernier P, Joly JS.
2005. Regulatory gene expressions in the ascidian ventral
sensory vesicle: Evolutionary relationships with the verte-
brate hypothalamus. Dev Biol 277:567–79.
Muller F, Chang B, Albert S, Fischer N, Tora L, Strahle U.
1999. Intronic enhancers control expression of zebrafish
sonic hedgehog in floor plate and notochord.
Development 126:2103–16.
Hedgehog and chordate brain evolution 107
Downloaded from https://academic.oup.com/icb/article-abstract/50/1/98/733550by gueston 09 April 2018

Murakami Y, Ogasawara M, Sugahara F, Hirano S, Satoh N,
Kuratani S. 2001. Identification and expression of the lam-
prey Pax6 gene: Evolutionary origin of the segmented brain
of vertebrates. Development 128:3521–31.
Muroyama Y, Fujihara M, Ikeya M, Kondoh H, Takada S.
2002. Wnt signaling plays an essential role in neuronal
specification of the dorsal spinal cord. Genes Dev 16:
548–53.
Onai T, Lin HC, Schubert M, Koop D, Osborne PW,
Alvarez S, Alvarez R, Holland ND, Holland LZ. 2009.
Retinoic acid and Wnt/beta-catenin have complementary
roles in anterior/posterior patterning embryos of the basal
chordate amphioxus. Dev Biol 332:223–33.
Osorio J, Mazan S, Retaux S. 2005. Organisation of the lam-
prey (Lampetra fluviatilis) embryonic brain: Insights from
LIM-homeodomain, Pax and hedgehog genes. Dev Biol
288:100–12.
Osorio J, Retaux S. 2008. The lamprey in evolutionary studies.
Dev Genes Evol 218:221–35.
Ota KG, Kuraku S, Kuratani S. 2007. Hagfish embryology
with reference to the evolution of the neural crest. Nature
446:672–5.
Placzek M, Briscoe J. 2005. The floor plate: multiple cells,
multiple signals. Nat Rev Neurosci 6:230–40.
Puelles L, Kuwana E, Puelles E, Bulfone A, Shimamura K,
Keleher J, Smiga S, Rubenstein JL. 2000. Pallial and sub-
pallial derivatives in the embryonic chick and mouse tele-
ncephalon, traced by the expression of the genes Dlx-2,
Emx-1, Nkx-2.1, Pax-6, and Tbr-1. J Comp Neurol 424:
409–38.
Ristoratore F, Spagnuolo A, Aniello F, Branno M, Fabbrini F,
Di Lauro R. 1999. Expression and functional analysis of
Cititf1, an ascidian NK-2 class gene, suggest its role in
endoderm development. Development 126:5149–59.
Rubenstein JL, Shimamura K, Martinez S, Puelles L. 1998.
Regionalization of the prosencephalic neural plate. Annu
Rev Neurosci 21:445–77.
Scholpp S, Wolf O, Brand M, Lumsden A. 2006. Hedgehog
signalling from the zona limitans intrathalamica orches-
trates patterning of the zebrafish diencephalon.
Development 133:855–64.
Shimamura K, Rubenstein JL. 1997. Inductive interactions
direct early regionalization of the mouse forebrain.
Development 124:2709–18.
Shimeld SM. 1999. The evolution of the hedgehog gene family
in chordates: insights from amphioxus hedgehog. Dev
Genes Evol 209:40–7.
Shimeld SM, van den Heuvel M, Dawber R, Briscoe J. 2007.
An amphioxus Gli gene reveals conservation of midline
patterning and the evolution of hedgehog signalling diver-
sity in chordates. PLoS One 2:e864.
Strahle U, Rastegar S. 2008. Conserved non-coding
sequences and transcriptional regulation. Brain Res Bull
75:225–30.
Sussel L, Marin O, Kimura S, Rubenstein JL. 1999. Loss of
Nkx2.1 homeobox gene function results in a ventral to
dorsal molecular respecification within the basal
telencephalon: evidence for a transformation of the palli-
dum into the striatum. Development 126:3359–70.
Takatori N, Satou Y, Satoh N. 2002. Expression of hedgehog
genes in Ciona intestinalis embryos. Mech Dev 116:235–8.
Tendeng C, Houart C. 2006. Cloning and embryonic expres-
sion of five distinct sfrp genes in the zebrafish Danio rerio.
Gene Expr Patterns 6:761–71.
Vavouri T, Lehner B. 2009. Conserved noncoding elements
and the evolution of animal body plans. Bioessays 31:
727–35.
Vavouri T, Walter K, Gilks W, Lehner B, Elgar G. 2007.
Parallel evolution of conserved non-coding elements that
target a common set of developmental regulatory genes
from worms to humans. Genome Biol 8:R15.
Vieira C, Martinez S. 2006. Sonic hedgehog from the basal
plate and the zona limitans intrathalamica exhibits differ-
ential activity on diencephalic molecular regionalization
and nuclear structure. Neuroscience 143:129–40.
Vieira C, Pombero A, Garcia-Lopez R, Gimeno L,
Echevarria D, Martinez S. 2010. Molecular mechanisms
controlling brain development: an overview of neuroepithe-
lial secondary organizers. Int J Dev Biol 54:7–20.
Villar-Cheda B, Perez-Costas E, Melendez-Ferro M,
Abalo XM, Rodriguez-Munoz R, Anadon R, Rodicio MC.
2006. Cell proliferation in the forebrain and midbrain of
the sea lamprey. J Comp Neurol 494:986–1006.
Wada H, Saiga H, Satoh N, Holland PW. 1998. Tripartite
organization of the ancestral chordate brain and the anti-
quity of placodes: insights from ascidian Pax-2/5/8, Hox
and Otx genes. Development 125:1113–22.
Walshe J, Mason I. 2003. Unique and combinatorial functions
of Fgf3 and Fgf8 during zebrafish forebrain development.
Development 130:4337–49.
Weigle C, Northcutt RG. 1999. The chemoarchitecture of the
forebrain of lampreys: evolutionary implications by com-
parisons with gnathostomes. Eur J Morphol 37:122–5.
Wilson SW, Rubenstein JL. 2000. Induction and dorsoventral
patterning of the telencephalon. Neuron 28:641–51.
Appendix 1: Materials and methods
Genomic sequences for Shh/Hhloci comparison
MLAGAN (Brudno et al. 2003) and VISTA plots
(Mayor et al. 2000; Frazer et al. 2004) were applied
on genomic sequences with 2 kb of upstream and
3 kb of downstream sequences around Shh/hh loci
(except for the lamprey, 5 kb upstream). Most geno-
mic sequences for Shh/Hh loci were obtained from
the UCSC genome browser, except the acorn worm
(see below) and the lamprey sequence obtained by
ourselves [LfHhb accession number: FP929027]. Each
position of sequence and its assemble version is indi-
cated as follows: mouse (chr5: 28,780,380–28,795,641,
108 S. Retaux and S. Kano
Downloaded from https://academic.oup.com/icb/article-abstract/50/1/98/733550by gueston 09 April 2018

mm9), chicken (chr2: 8,021,868–8,036,924, galGal3),
zebrafish (chr7: 36,654,711–36,666,126, danRer5),
fugu (chrUn: 211,116,380–211,127,115, fr2), medaka
(chr20: 17,733,043–17,744,774, oryLat2), ascidian
(chr05q: 4,583,360–4,593,934, ci2), lancelet (chrUn:
399879050–399909584, braFlo1), sea urchin (scaf-
fold68424: 22314–56576, strPur2), fly (chr3R:
18950425–18970881, dm3). The genomic sequence,
scaffold44409, for Saccoglossus kowalevskii hedgehog
locus was obtained, using BLAST search with partial
cDNA sequence (DQ431035.1), from BCM-HGSC
web site, which is generated by Acorn Worm
Genome Sequencing Consortium (Dr R. Gibbs, per-
sonal communication). The genomic sequence was
manually annotated with aids of BlastX.
[Note: position of the Shh sequences, version
of assembly.] [Note #2: in the newest zebrafish
assembly (danRer6), two zebrafish Shha loci were
merged into a single one. The sequence used in
the present study is slightly different from the
newest one (in ver danRer6) with 99.4% of
similarity.]
Hedgehog and chordate brain evolution 109
Downloaded from https://academic.oup.com/icb/article-abstract/50/1/98/733550by gueston 09 April 2018