Loss of Function Mutations in NNT Are Associated with Left...
-
Upload
nguyenhanh -
Category
Documents
-
view
214 -
download
1
Transcript of Loss of Function Mutations in NNT Are Associated with Left...

DOI: 10.1161/CIRCGENETICS.115.001026
1
Loss of Function Mutations in NNT Are Associated with
Left Ventricular Noncompaction
Running title: Bainbridge et al.; Mutations in NNT are associated with LVNC
Matthew N. Bainbridge, PhD1,2*; Erica E. Davis, PhD3*; Wen-Yee Choi, PhD4;
Amy Dickson, BS4; Hugo R. Martinez, MD5; Min Wang, PhD1; Huyen Dinh, PhD1;
Donna Muzny, MS1; Ricardo Pignatelli, MD5; Nicholas Katsanis, PhD3; Eric Boerwinkle, PhD1;
Richard Gibbs, PhD1; John L. Jefferies, MD5
1Human Genome Sequencing Center, 5Department Pediatrics-Cardiology, Baylor College of Medicine; 2Codified Genomics, LLC, Houston, TX; 3Center for Human Disease Modeling, Duke
University Medical Center; 4Department of Cell Biology, Duke University, Durham, NC *contributed equally
Correspondence:
Richard A. Gibbs, PhD John Lynn Jefferies, MD
Human Genome Sequencing Center The Heart Institute, CCHMC
One Baylor Plaza, MS226 3333 Burnet Avenue
Houston, TX 77030 Cincinnati, OH 45229
Tel: (713) 798 6539 Tel: (513) 803-1675
Fax: (713) 798 5741 Fax: (513) 803-3315
E-mail: [email protected] E-mail: [email protected]
Journal Subject Codes: [108] Other myocardial biology
Donna Muzny, MS ; Ricardo Pignatelli, MD ; Nicholas Katsanis, PhD ; Eric Boerwwinkle, PhD ;
Richard Gibbs, PhD1; John L. Jefferies, MD5
1HHHuman Genonoommme SSSeqeqequeueuenncncininingg CeCeCennnterrr, 55Depapapartmmennnt PPPededediattrtricii s-CaCaCarddioololol gggy, BaBaBa lylororor CCColololleeeggge ooof f fMeMeMedidd cine; 2Codiiifiied GGGeeenommiciccs,s LLC, HoHoHousttoonnn, TX;X;X; 3CeCeCentntntererr fffor HHHuuuman DDDiseaeaeassse Modododeeliiinggg, DDDukkke
UnUnUniverrrsisisitytyty MMMeeediccaaal Centeteter;r;r; 4DeDeDeppparrtrtmemementntnt ooof f Ceeelllll BBBioioololologgygy,,, DDDuukukee e UnUU iiiveeersiiitytyty, , , Duuurhrhrhamamm, NNNC *contribibuteded equ lalllylyly
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

DOI: 10.1161/CIRCGENETICS.115.001026
2
Abstract:
Background - Left ventricular noncompaction (LVNC) is an autosomal dominant, genetically
heterogeneous cardiomyopathy with variable severity, which may co-occur with cardiac
hypertrophy.
Methods and Results - Here, we generated whole exome sequence (WES) data from multiple
members from five families with LVNC. In four out of five families, the candidate causative
mutation segregates with disease in known LVNC genes MYH7 and TPM1. Subsequent
sequencing of MYH7 in a larger LVNC cohort identified seven novel likely disease causing
variants. In the fifth family, we identified a frameshift mutation in NNT, a nuclear encoded
mitochondrial protein, not implicated previously in human cardiomyopathies. Resequencing of
NNT in additional LVNC families identified a second likely pathogenic missense allele.
Suppression of nnt in zebrafish caused early ventricular malformation and contractility defects,
likely driven by altered cardiomyocyte proliferation. In vivo complementation studies showed
that mutant human NNT failed to rescue nnt morpholino-induced heart dysfunction, indicating a
probable haploinsufficiency mechanism.
Conclusions - Together, our data expand the genetic spectrum of LVNC and demonstrate how
the intersection of WES with in vivo functional studies can accelerate the identification of genes
that drive human genetic disorders.
Key words: noncompaction cardiomyopathy, genetics, human, genomics, left ventricular noncompaction
y p g
Suppression of nnt in zebrafish t caused early ventricular malformation and contraaactcttilililitity y y dededefefefectctc s,
ikely driven by altered cardiomyocyte proliferation. In vivo complementation studidies showed
haaattt mmumuttantntt huhuhumamamannn NNT failed to rescue nnt morphphphooolino-induced heheheart dydydyssfunction, indicating a t
pprprobbbable haplooinininsuuufffficicicieieencncncyyy mememe hchchanananisismmm.
CoCoConncnclusions - Togggethererer, our r dadadata expxpxpannnddd theee gggenettticcc spsppececectrt umumum off f LLLVNCCC aaanddd dddeme onnnststtrarattte howww
he ini tetetersrsrseecectionn oofff WEWEWESSS wwith inn ivivivo funcncctititiononalal sstuttudididieses cccaanan aacccccelerratatatee thththe e idididene titiififificcacationn oofff gegeneness
hat drive human geneneneteteticicic dddisisisororordededersrsrs..
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

DOI: 10.1161/CIRCGENETICS.115.001026
3
Left ventricular noncompaction/hypertrabeculation (LVNC/LVHT) is a clinically distinct,
primary cardiomyopathy1. LVNC is characterized by a spongy, noncompacted myocardium layer
clearly distinct from the underlying, compacted myocardium2, as well as increased trabeculation
and deep intertrabecular recesses in the left ventricle3. LVNC has variable clinical severity,
ranging from benign to early-onset, severe heart failure and death4. LVNC may be first detected
prenatally or as late as 95 years of age, and appears to affect a greater proportion of males than
females5. Although readily diagnosable by echocardiogram6, the true prevalence is estimated to
be between 0.014% and 0.24%2,7.
Although LVNC may occur in isolated cases, familial LVNC is recognized as a genetic
disorder that is inherited in an autosomal dominant (AD) manner8. LVNC is associated
frequently with mitochondrial disorders and cardiac hypertrophy. It is most commonly attributed
to mutations in seven genes (TAZ, DTNA, LDB3, LMNA, SCN5A, MYH7 and MYBPC3), with
additional contributing variants reported in rare instances (ACTC1, TNNT2, MIB1, PRDM16, and
TPM1); all LVNC loci encode proteins involved in cellular energy, muscle development, ion
channel formation, or are components of the muscle filaments9–13. Despite these advances, many
of the causative LVNC genes have yet to be identified1. Mutations in MYH7 are the single most
common cause of LVNC, accounting for 8-13% of cases, with the remainder of the genes
reported to be mutated in rare cases10,12. The large number of causative loci, as well as overlap
with other cardiomyopathy genes, suggests that hypertrabeculation is a common compensatory
mechanism of impaired or injured myocardium1. To improve our understanding of the genetic
and molecular basis of this disorder we used whole exome sequencing14,15 (WES) to conduct an
unbiased investigation of the genetic basis of LVNC.
disorder that is inherited in an autosomal dominant (AD) manner8. LVNC is assooociciiatatatededed
frequentntlylyy witith mimitochondrial disorders and cardiaacc hypertrophy. It is mmoso t commonly attributed
ooo mmmutations iiin n sesesevevv n n n gegegenenenes s s (TATATAZ,Z,Z DDDTNTNTNA,AA LLLDBDBDB3, LLLMMMNA,A,A, SSSCNCNCN5AAA,,, MYMYMYH7H7H7 anaa d MYMYMYBPBPBPC3C33),),), wiwiwiththth
adadddidiititt onal contriibbuuutinggg vvvariannntststs repooortrr eeed in raaarrre inssstaaanccceseses (ACACACTC11(( , TNNTNTNT222, MIMIMIB1B , PRPRPRDMDMM161 , annnd
TPM1); all LVNC lococociii eencoddee e prpp oto eins involved ininin cellulararar eeeneerggy,y mumuuscscscle development, ion
chchanannenell foformrmatatioion,n,, oor r araree cocompmppononenentsts oof f ththee mumuscsclele ffililamamenentsts999–131313.. DeDespsppitite e ththesese e adadvavancnceses, , , mamanynyy
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

DOI: 10.1161/CIRCGENETICS.115.001026
4
Methods
IRB approval for this study was obtained from Baylor College of Medicine. All participants
provided informed consent prior to participating in this study.
DNA capture sequencing
Hybridization was conducted with a custom capture reagent15. Precapture libraries for SOLiD (2
ug) were hybridized in solution according to the NimbleGen Seqcap EZ protocol with minor
revisions. Specifically, hybridization enhancing oligos TrTA-A and SOLiD-B were used in the
hybridization reactions to block the common TrTA adaptor sequences for increased capture
efficiency. Following sequence capture, post-capture LM-PCR was performed using 12 cycles.
Capture libraries were quantified using PicoGreen (Cat. No. P7589) and their size distribution
was analyzed using the Agilent Bioanalyzer 2100 DNA Chip 7500 (Cat. No. 5067-1506).
Capture efficiency of each capture library was evaluated by performing a qPCR-based SYBR
Green assay (Applied Biosystems; Cat. No. 4368708 ) with built-in controls (RUNX2, PRKG1,
SMG1, and NLK). Capture library enrichment was estimated at 7 to 9-cycles over background by
qPCR. Captured libraries were processed further for sequencing, with approximately 6-12 Gbs of
sequence generated per capture library on SOLiDv4 instruments. Illumina library preparation
was conducted as described previously15.
Whole Exome Sequencing Analysis
Sequence data were aligned to the human genome (hg19) and variants were identified and
filtered for quality as described14,15. Briefly, read qualities were recalibrated with GATK and a
minimum quality score of 30 was required; the variant must have been present in at least 15% of
the reads that cover the position. In addition, prior to variant calling, reads with low (<11)
mapping qualities (MQ) (a value based on the ratio of the best alignment score to the second best
Capture libraries were quantified using PicoGreen (Cat. No. P7589) and their sizzze e dididiststtririribububutititionoo
was annala yzyzy edd usiingng the Agilent Bioanalyzer 2100 DND A Chip 7500 (Cat. . NoN . 5067-1506).
CCaCapppture efficicicienenncycycy of f f eaeaeachchch cccappptututurerere lllibibibrarararyryry wwwasasas evaaaluuuatededd bbby y y pepp rfffooormimimingngng aaa qqPCCRRR--bababaseses d d SYSYSYBRBRBR
GrGrGreeeeen assay (A(( pppplllied BBiosysssteemems; CCCata ... NNNo. 434343687770888 ) ) wiwiwitth buuuilt---innn contttroools ((RRRUNXXX22( ,, PPPRKKKG111,
SMG1, and NLK). CCCapapaptuture libbbrarararyryr enricchment waaasss ese timateteted d d atat 7 to 99KK -c-ccycycycles over background by
qPqPq CRCR.. CaCaptptp urureded llibibrararirieses wwereree prprp ococesessesed d fufurtrtheher r fofor r seseququq enencicingngg, , , wiwithth aapppppproroxiximamatetelylyy 66-112 2 GbGbss oo
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

DOI: 10.1161/CIRCGENETICS.115.001026
5
alignment score) were removed. Common variants were filtered based on minor allele frequency
and previous association with disease using previously established parameters16. Variant effect
on protein was established in both RefSeq and ENSEMBL (v78) gene sets. Variants were
prioritized based on phenotypic overlap, deleteriousness of the variant (e.g. truncating/splice
affecting and in silico predictions), and known mutational spectrum of the gene in unaffected
populations.
Capillary Sequencing
Validation and segregation of discovered variants, and sequencing of the NNT coding exons in
the expanded cohort was conducted on an ABI3730 capillary sequencer. Amplification and
sequencing primers were designed by an automated pipeline; variants were automatically called
using SNPDectector (version 3). Variants were subsequently validated visually.
Ion Torrent Sequencing
PCR primers were designed for each coding exon of MYH7 using primer3. Sixteen molecular
barcodes were generated and appended to each of the forward PCR primers. Each exon for each
subject was amplified separately, and then pooled into groups of 16 prior to sequencing on the
Ion Torrent PGM. Reads were mapped to the genome and variants called as described above for
the Illumina data.
Zebrafish embryo injections and live phenotypic assessment
We identified two zebrafish orthologs of human NNT protein, referred to as nnt-a and nnt-b on
D. rerio chromosomes 21 and 18 respectively using reciprocal BLAST. We then designed splice-
blocking morpholinos (MO)s against the splice donor site of exon 4 of each transcript (Gene
Tools). We injected 9ng of each MO into wild-type embryos at the one- to two-cell stage and
determined MO efficiency by RT-PCR of cDNA generated from whole embryos (n=25
equencing primers were designed by an automated pipeline; variants were autommmatataticici alalallylyly cccalalalllled
using g SNSNPDece tectctor (version 3). Variants were sububses quently validated vivisus ally.
ooon n n Torrent t SeSeSequququennncicicingngng
PCPCPCRRR primers weeereee desssigggned fofofor r eachhh codododing exxxon offf MYMYMYHHH7 uusinggg ppprimer3r3r3. SSiSixxtxteen momomoleeecuuular r 7
barcodes were geneerararateteted d and apapappepep ndedd to each of f f ththe forwwwararard PCR prprprimimimeree s. Each exon for each
uubjbjjecect t wawas s amamplplp ififieied d sesepapap raratetelylyy,,, anand d ththenen pppooooleled d inintoto gggroroupuppss ofof 1166 prprp ioior r toto sseqeqqueuencncining g g onon tthehe
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

DOI: 10.1161/CIRCGENETICS.115.001026
6
embryos/injection batch; Quantitect Reverse Transcription kit, Qiagen) harvested at 3 days post-
fertilization (dpf) in Trizol (Invitrogen). Specific targeting of each of nnt-a and nnt-b is
evidenced by the semi-quantitative decrease in correctly spliced transcript. Phenotype specificity
experiments were conducted by targeting hacl1, nav1, rab28, rbm28 and notch1a. We conducted
live embryo phenotypic scoring at 2, 3, or 5 dpf using either wild-type (EK/AB outcross); or
cmlc2:GFP 17, a myocardium-specific GFP transgene. MO dose response curves for individual
and combined suppression of nnt-a and nnt-b (3ng, 6ng, and 9ng MO) were generated with
EK/AB embryos at 3 dpf; n=50-100 embryos/injection, repeated three times with masked
scoring. For rescue experiments, we generated a full-length human NNT ORF construct by PCR
amplification from lymphocyte cDNA, cloning into a pCR8/GW vector (Invitrogen), and LR
recombinase-mediated cloning into the pCS2+ backbone. We conducted mutagenesis using the
QuikChange site directed mutagenesis kit (Agilent); all vectors were sequence confirmed.
Capped mRNA was in vitro transcribed using the mMessage mMachine kit (Ambion); 200 pg
RNA and 5 ng MOs were used for in vivo complementation experiments. To assess cardiac
heartbeat and morphology, we anesthetized 2 dpf cmlc2:GFP larvae with Tricaine, and
conducted live video imaging of ventral views using a Nikon AZ100 microscope and NIS
Elements AR software at 12x magnification (10 second videos at 8.77 frames/second; n=10
embryos/injection, replicate batches; see Supplementary Movies).
Zebrafish histology
We generated cardiomyocyte cell counts at 2 and 3 dpf using cmlc2:FUCCI18, a double
transgenic line enabling the detection of non-proliferating (cmlc2:mCherry-zCdt1, red) and
proliferating (cmlc2:Venus-hGeminin, green) cardiomyocytes. Larval batches were fixed
overnight in paraformaldehyde (PFA), and 50 -thick sections were imaged using a Zeiss LSM
amplification from lymphocyte cDNA, cloning into a pCR8/GW vector (Invitroggegen)n)n), ananand d d LRLRLR
ecombbininase-mmeddiai ted cloning into the pCS2+ bacckbkbone. We conducted d mum tagenesis using the
QQQuikikikChange sssititteee didd reeectctctededed mmmutagagagenenenesesesisiss kkkitii (((AgAgAgilii enttt);;; alll vevevectctctoroo s wwwererere seqeqequeueu nce e e cococonfnfnfiriri mememed.d.d.
CaCaCapppppped mRNA wawawas innn vvvitro tttraraanscribbbeddd uuusinngg tthe mmmMeMeessssssageee mMmMmMaccchine kkkit (((AmAmAmbionnn))); ; 2220000 pggg
RNA and 5 ng MOs s s wewew re useeed d d foff r in vivo compleeememm ntatiooon n n exexe perimeentntn sss.. To assess cardiac
hehearartbtbeaeat t anand d momorprpphoholologygygy, , , wewe aaneneststhehetitizezed d 22 dpdppf f cmcmlclc2:2:GFGFPP lalarvrvaae e wiwithth TTriricacainine,e,, aandnd
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

DOI: 10.1161/CIRCGENETICS.115.001026
7
700 confocal microscope and ZEN software at 20x magnification. Maximum intensity
projections were analyzed using Imaris Version 7.6 software; mCherry+ and Venus+ nuclei were
quantified using surface analysis (n=12 larvae/batch; repeated once). Red+green+ and red+green
nuclei were scored as non-proliferating cells, and red green+ nuclei were scored as proliferating.
Statistical Analyses
Mutation burden:
To test whether the frequency of rare variants in MYH7 differed between ARIC and our LVNC
cohort we used a two-tailed Fisher’s exact test. The Atherosclerosis Risk In Communities
(ARIC)19 cohort consists of 1,650 individuals who have undergone WES using the NimbleGen
capture reagent followed by Illumina sequencing15. On average, >99% of the coding bases in
MYH7 have at least 20x redundant coverage in this dataset.
In vivo complementation:
To test for differences in cardiac edema in 3 dpf morphant zebrafish larvae, we compared
qualitative scoring data (normal or affected) in a pairwise manner across embryo batches using
2 tests (n=46-15 embryos/batch; three biological replicates).
Contractile dysfunction and cardiomyocyte counts:
We used a student’s t-test to determine the statistical significance of differences in heart beat (2
dpf controls and morphants; n=10 larvae/injection batch; two biological replicates). Differences
in proliferating and non-proliferating cardiomyocytes were also determined using a two-tailed,
homoscedastic Student’s t-test (2 and 3 dpf controls and morphants; n=12 larvae/injection batch;
two biological replicates).
Results
We identified five families (A-E) with apparent hereditary LVNC (Supplementary Figure 1A-E).
capture reagent followed by Illumina sequencingff 15. On average, >99% of the coddidingngng bbbasasaseseses iiinn n
MYH7 hhava e ata leaeasts 20x redundant coverage in thiis s dad taset. 77
InInIn vvvivo complplplememmenene tataatititiononon:::
ToToTo tttesee t for differrrennncess innn cardididiaaca edeeemamama iiin 3 dpppf mmmorphahahantnnt zzzebbbrafffishh larvvvaeee, wwweee commpmpaaareeed
qualitative scoring dadadatatata (n( ormamamall l or affeccted) in a papap irwise mmmanana ner acroroossssss embryo batches using
2 tetetestststsss (((n=nn 464646-151515 eeembmbmbryryryososos/b/b/batatatchchch;;; thththrerereeee bibibiololologogogicicicalalal rrrepepeplililicacacatetetes)s)s)...
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

DOI: 10.1161/CIRCGENETICS.115.001026
8
Initial presentation of the index case for each family was secondary to either an abnormal
electrocardiogram or a screening echocardiogram that led to the diagnosis of LVNC. The
remaining family members in each pedigree underwent transthoracic echocardiography for
indicated screening. Varying degrees of coexisting myocardial involvement were identified,
including evidence of dilation of the left ventricle (LV) with depressed LV systolic function
(dilated phenotype); or increased left ventricular thickness and increased LV systolic function
(hypertrophic phenotype). One member of family C had severe dilated cardiomyopathy (DCM)
with associated LVNC that required mechanical assist and cardiac transplantation.
We generated WES data from multiple affected and unaffected members, when available,
from all families (Table 1). In total, 228 Gbp of sequence data were produced and aligned to the
human genome, producing an average depth of coverage of 79x (Supplementary Figure 1A-E).
Bioinformatic analyses were used to identify candidate causative alleles in each family. Variants
were prioritized for validation based on a) minor allele frequency (MAF); b) whether they were
predicted to alter protein function (either through a predicted change to mRNA splicing or
truncation/substitution of the amino acid sequence); c) previous association with LVNC or genes
with functional similarity to known LVNC genes; d) whether they were shared by all affected
individuals, but not present in unaffected individuals; and e) whether mutants in other organisms
had been reported with relevant heart pathologies. In all cases, the small size of the pedigree and
AD inheritance of the disease made it difficult to identify a single variant by segregation alone.
However, when a predicted pathogenic allele was identified in a known LVNC gene, it was
considered the most likely causative variant (Supplementary Table 1). We then segregated all
LVNC-associated variants reported here by capillary sequencing in all available members of the
pedigree.
from all families (Table 1). In total, 228 Gbp of sequence data were produced annddd alalaligiggnenened d d tototo tthehh
human n gegeg nomem , prprp oducing an average depth of coveverage of 79x (Supplemementary Figure 1A-E).
BBiBioioiinformatiiccc anananalala ysseseses wwwerereree usususededed ttto oo ididdenenentifyfyfy cccandiiidddate cccauauausasasativevee allllllelele esess iiin nn eachchch fffamamamili y.y.y. VVVararariaiaiantntn s
wewewereree prioritized fooor vaaaliddatiooon n bab sed d d ononn aaa) mmminnnor aaalleeleee fffrrrequuueeencyyy (MMMAFF); b)) wwwheh theerer ttheeeyy y weereee
predicted to alter prorooteteteinin functctctioioion (e( itheer througgh h a aa prp ediccteteted dd chc angeg ttooo mRmm NA splicing or
rrununcacatitionon/s/sububststititututioion n ofof tthehe aamiminono aacicid d seseququq enencece););); cc))) prprp evevioiousus aassssocociaiatitionon wwitith h LVLVNCNC oror gggeneneses
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

DOI: 10.1161/CIRCGENETICS.115.001026
9
In four out of five families we identified novel nonsynonymous changes in known LVNC
genes. In three families (B, D, E) we identified rare, protein changing mutations in MYH7,
encoding myosin heavy chain beta, p.S648L, p.Q163P and p.E700G respectively, none of which
have been previously reported but all of which were predicted to be pathogenic with multiple in
silico prediction algorithms (PolyPhen-2 HDIV/HVAR scores >0.9; Mutation Taster, Mutation
Assessor; Table 1). MYH7 consists of 40 exons, encoding a protein of 1,935 residues that is
present primarily in slow-type skeletal muscle and in the ventricles20. Mutations in MYH7 are
associated with other cardiomyopathies, including familial hypertrophic cardiomyopathy type 1
and dilated cardiomyopathy type 1S21. In family A, we discovered a p.D275H variant in another
known LVNC gene22,23, tropomyosin 1 (TPM1). TPM1 encodes an actin-binding tropomyosin, is
involved in the contractile system in both smooth and striated muscle24 and accounts for a small
proportion of LVNC cases.
In some cohorts, mutations in MYH7 are thought to account for a minority of LVNC
cases but were found in three of the five families we studied. We hypothesized that the difficulty
inherent in sequencing and interpreting data derived from a gene as large as MYH7 might lead to
under-reporting of its association with LVNC. The capture sequencing reagent used here
provides high, relatively even coverage of every coding exon in MYH7 and subsequently we
found 60% of our families had rare, protein changing variants in MYH7. We investigated the
prevalence of MYH7 mutations in a cohort of 49 individuals diagnosed with LVNC using a PCR-
amplicon high-throughput sequencing strategy. In total, we produced >350Mbp of sequence data
from the Ion Torrent sequencing platform, generating an average depth of coverage >200x across
all coding exons. Eight individuals harbored a single, rare (MAF <0.001), protein changing
mutation in MYH7 and one (L030) harbored two rare mutations (Supplementary Table 2). The
known LVNC gene22,23, tropomyosin 1 (TPM1). TPM1 encodes an actin-bindinggg troroopopoomymymyosososininin, is
nvolveded in thhe cocontn ractile system in both smooth anand striated muscle24 anand accounts for a small
prprproppportion of ff LVLVLVNCNCN cccasasaseseses.
In some cccooohorttts,,, muttatatititions innn MYMYMYH7 arrre thhhouugughththt ttto acacaccouuuntt t for aaa mmminnonorrrity offf LVLVLVNNNC77
cases but were foundndd iiin nn threee ofofof the five e families wwwee studieieed.d.d. WWe hyypopopothththesee ized that the difficulty
nnhehererentnt iin n seseququq enencicingngg aandnd iintntererprprp etetining g g dadatata ddereriviveded ffrorom m a a gegeg nene aass lalargrggee asas MYMYH7H7 mimighghg t t leleadad ttoo
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

DOI: 10.1161/CIRCGENETICS.115.001026
10
proportion (18.4%) of subjects with MYH7 mutations is modestly higher compared to similar
studies10,12 in which 7.9-13% of individuals harbored pathogenic mutations in MYH7. Although
we cannot be certain that all discovered variants are deleterious without functional testing, we
compared the frequency of rare, protein changing MYH7-mutations in LVNC to a cohort of
unselected individuals from the Atherosclerosis Risk In Communities (ARIC)19 cohort. We
found that only 2.5% of ARIC individuals had rare, protein changing mutations in MYH7, a 7.4
fold enrichment that, despite the relatively modest case sample size in our study, was nonetheless
significant (p<0.0001). Although the MYH7 sequencing data produced from the ARIC samples is
of high quality (>99% of coding bases at >20x coverage; see Methods), we cannot rule out the
possibility that different sequencing methods may contribute to this result. Even so, these
findings may imply that MYH7 plays a greater role in the development of LVNC either as the
sole cause of the disease, or possibly acting in concert with other genes. Of note, four (50%) of
the mutations we discovered in our LVNC cohort were reported previously25–28 and in one case
(p.L061P in L022) a mutation affecting the same amino acid (p.L061R) was reported21. This may
indicate that the set of pathogenic mutations in MYH7, at least in European populations, is
relatively well surveyed.
In family C, neither known LVNC genes, nor other genes associated with human
cardiomyopathy were found to contain rare, protein changing mutations. Of the 25 genes that
remained after analysis, the majority either had no known function or no shared function with
other known LVNC genes (e.g. related to the heart, musculature, or cellular energy). Cross-
referencing this gene set with human (OMIM) or mouse (MGI) phenotype databases reduced the
priority of most of these genes on the basis that loss of function mutations induced phenotypes
irrelevant to the cardiac defects in family C, with two exceptions: MAP1S and NNT
possibility that different sequencing methods may contribute to this result. Even sso,o,, tthehehesesese
findingsgsg mam y y y implplp y that MYH7 plays a greater rolee iin the development ofof LVNC either as the
oooleee cause of ff thththee e didd seeeasasase,e,e, ooor r r pooossssssibibiblylyly aaactctctinii g gg ininin connnceeert wwwititith h h ototo heer r r geeenenen s.s.s. OfOO nototteee, fofofoururu (5(5(50%0%0%) ) ) ofofof
hhhe e mmmutations weee dddiscooovvvered d innn our LLLVNVNNC cooohhhort wwwere e e rererepooorttted pppreeeviouuuslly25255–2228 and innn oonenee casssee
p.L061P in L022) a a a mumum tationnn aaaffff ece tingg the same amama ino acccididid ((p.p.L061R)R)R) wwwas reported21. This may
nndidicacatete tthahat t ththe e seset t ofof pppatathohogegeg ninic c mumutatatitionons s inin MYMYH7H7, , , atat lleaeastst iin n EuEuroropepep anan pppopoppululatatioionsns,,, isis
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

DOI: 10.1161/CIRCGENETICS.115.001026
11
(Supplementary Table 3). However, MAP1S contains numerous homozygous frameshift alleles
in control individuals, suggesting that its loss of function is tolerated in humans. In contrast,
NNT, encoding nicotinamide nucleotide transhydrogenase, remained a strong candidate for
several reasons. First, the mutation detected was a frameshift allele within the fourth coding exon
of the gene that introduces a premature codon that likely renders the transcript subject to
nonsense-mediated decay (Figure 1A). Second, we found a single heterozygous truncating or
stop mutation at any position of the coding sequence in 13,000 control chromosomes; this
frequency is consistent with the prevalence of LVNC and suggests that severe truncating
mutations at this locus are generally intolerable. Third, mouse Nnt mutants have revealed a direct
relevance of NNT to cardiac development and function: Nnt null29 mice have a mild
cardiovascular background phenotype including lower LV shortening, higher aortic ejection
time, and increased LV weight30. Moreover, when these mice are rendered null for a second
mitochondrial gene, MnSOD, they develop lethal prenatal cardiac-hypertrophy29. This is in
contrast to MnSOD-null mice on other (i.e. NNT normal) backgrounds which are long lived with
no signs of cardiac-hypertrophy29. Finally, all these observations are consistent with a critical
role for NNT in cellular health: NNT is an evolutionarily-conserved, nuclear-encoded
mitochondrial protein that is involved in cellular respiration31; it is responsible for coupling
hydride transfer between NAD(H) and NADP(+) to proton translocation across the inner
mitochondrial membrane (Figure 1B)32. In mitochondria, NADPH is used for the regeneration of
glutathione and theoredoxin, two important antioxidant molecules.
Despite the known cellular and in vivo roles of NNT relevant to heart development and
function, we were cautious in interpreting a pathogenic mutation in a single family. Therefore,
we expanded our genetic study and Sanger sequenced all coding exons of NNT in a cohort of 56
elevance of NNT to cardiac development and function: Nnt nullt 29 mice have a mmimildldld
cardiovavascs ulara bacackground phenotype including loowew r LV shortening, hhigiggher aortic ejection
iiimememe, and incrcrreaeaasesesed d LVLVLV wwweieieighghttt300.. MoMoM rerereovovoverrr, , whww enn thhheseee mmmicicice ee arrre e e reeendndnderererededed nulullll l fofofor r r a a a seeecococondndnd
mmimitooochondrial geeennne, MnMnMnSODDD,, tththey ddevee eeelooop leeethhhal pppreeenatatatalalal caarardiaccc-hhhyperrttrooophphhyyy29y . Thhhisss isss innn
contrast to MnSOD-D-nununull mice e e ononon oother ((i.e. NNT nnnoro mal)) bbbacaca kggroundsdss wwwhich are long lived with
nono ssigiggnsns oof f cacardrdiaiac-c hyhyypepep rtrtrorophphp yyy292929y .. FiFinanalllly,y,y, aallll tthehesese oobsbserervavatitiononss arare e coconsnsisistetentnt wwitith h a a crcrititicicalal
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

DOI: 10.1161/CIRCGENETICS.115.001026
12
unrelated individuals with LVNC (Supplementary Table 2). We identified a missense mutation
that affected an evolutionarily conserved residue (p.D277Y) in a single individual (L011), who
was diagnosed at infancy with LVNC and cyanotic congenital heart disease. This mutation was
predicted in silico to be pathogenic (PolyPhen-2: possibly damaging, score 0.686; Mutation
Taster: disease causing; Mutation Assessor: high impact) and was absent from all available
controls.
Cumulatively, the two families with NNT mutations and the previously-reported
phenotypes in mouse strengthened the candidacy of NNT in LVNC. Nonetheless, even though
the mutation in the second family was unique to the patient and was predicted to be deleterious,
we sought to test directly its effect on protein function. We therefore employed an in vivo
complementation strategy in a transient zebrafish model system, which we have used previously
to study the effect of missense mutations for a range of human phenotypes33,34 Using reciprocal
BLAST, we detected two NNT orthologs in the Danio rerio genome, nnt-a and nnt-b (83%
identical, 91% similar (a); and 75% identical, 85% similar (b), versus human respectively;
Supplementary Figure 2A), of which our in-house RNAseq data from the anterior structures of 5
dpf zebrafish larvae indicated that nnt-a has markedly increased expression when compared to
nnt-b (456 vs. 2 counts per million mapped reads; (GSE #63191)35. Individual injection of
morpholinos (MO) targeting either nnt-a or nnt-b and scoring blind to injection cocktail (n=50-
100 embryos per injection) resulted in reduced endogenous transcript (Supplementary Table 4;
Supplementary Figure 2B) and impaired heart function as evident by cardiac edema at 3-days
post fertilization (Figure 1C). Importantly, whereas individual MO injections gave rise to 30%-
40% affected embryos at the highest dose tested (9 ng) for nnt-a and nnt-b respectively,
combined suppression of nnt-a/b yielded 90% affected embryos (n=46-51 embryos/injection;
we sought to test directly its effect on protein function. We therefore employed aaann ininin vvvivivivooo
comppleemem ntatation n sts rategy in a transient zebrafish mmodo el system, which wwe have used previously
ooo stttudy the eeeffffffececectt t off mmmisisissesesensnn e e mumumutatatatitiononons ss fooor r r aa a rangggeee off hhhumumumanaa ppphhhenononotytytypepepesss33,34 UsUsUsinining g g reeeciciciprprprocococalala
BLBLBLAAAST, we deteeecttted tttwwwo NNNNTTT orthooolol gsgsgs in tttheee TT Daaannio rerererririo ggenomemem , nnttt-aaa aandndnd nnt-bbb (((83%3%3%
dentical, 91% similllararar ((a)a); annd d d 7577 % % idene tical,, 85%%% ssimilarrr (((b)b)b ,, versuss hhhumumuman respectively;
SuSupppppplelemementntarary y y FiFigugug rere 22AA),),), oof f whwhicich h oourur iinn-h-houousese RRNANAseseq q q dadatata ffrorom m ththe e ananteteririoror sstrtrucuctutureres s ofof 55
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

DOI: 10.1161/CIRCGENETICS.115.001026
13
(Supplementary Figure 2C); this may be the result of varying dose thresholds for each of the two
encoded zebrafish NNT proteins. Moreover, the cardiac edema phenotype of nnt morphants was
reminiscent of those observed for other morphants modeling known human LVNC genes such as
taz, ldb3, lmna, and prdm16 9,36–38and was consistent with the observed phenotype of our
positive experimental control (notch1a)39. Additionally, as a test of specificity of the assay,
suppression of four randomly selected genes harboring rare variants in LVNC Family C did not
produce a cardiac edema phenotype (Supplementary Figure 2A, B, D; Supplementary Table 4).
The edema phenotype was specific to the nnt-a/nnt-b MOs, since we were able to rescue
the MO-induced defects by co-injecting 200 pg of capped wild-type (WT) human NNT mRNA
(45% vs. 12% affected embryos for MO vs. WT rescue; p<0.0001; Figure 1D). Next, we
compared the rescue efficiency of NNT mRNA harboring the 277Tyr-encoding change found in
the LVNC patient to that of WT. Blind scoring of injected embryos showed that the mutant
rescue was significantly worse than WT (p=0.0027), and was ameliorated significantly from that
of MO (p<0.0001) (Figure 1D), indicating that the variant results in a partial loss of protein
function in this assay and is a likely hypomorph. We saw no significant cardiac defects in
embryos injected with either WT or p.D277Y message in the absence of MO.
To determine the cellular and morphological basis underlying the nnt-a/nnt-b MO-
induced cardiac edema, we utilized transgenic reporter zebrafish lines to refine the relevance of
the phenotype to LVNC in humans. Although humans and zebrafish display key differences in
cardiac structure (most notably, zebrafish do not have separate left-right circulation), the highly
conserved signaling pathways in cardiac development make the zebrafish a useful surrogate
model to evaluate the candidacy of NNT in LVNC (reviewed in ref 33). First, we conducted live
imaging of 2 dpf cmlc2:GFP control and nnt-a/nnt-b MO-injected larvae to visualize cardiac
45% vs. 12% affected embryos for MO vs. WT rescue; p<0.0001; Figure 1D). NNNexexxttt, wwwe e e
compparareded thee resscucue efficiency of NNT mRNA haarbrboring the 277Tyr-encncodo ing change found in T
hhhe LLLVNC patatatieeentntnt too thththatatat ooof f f WTWTWT.. BlBlBlininind d d scscscorrrinining gg of innnjecteteted d d ememembrryyoyosss shshshowowowedee thahahattt thththee e mumuutatatantntnt
eeescscscueuu was sigggnificccanttlylyy worrrsesese than WTWTWT (p=000.00002777), andndnd waaas ameeeliiiorateeed sigggnininificannntlylyly fffrooom ttthaaat
of MO (p<0.0001) (((FiFiFigugugure 1DDD),),), indicatting g that thehehe vvarianttt rrresee ulu ts in a a papapartrr ial loss of protein
fufuncnctitionon iin n ththisis aassssayayy aandnd isis a a lilikekelylyy hhypypypomomororphphp . . WWe e sasaw w nono ssigiggninifificacantnt ccarardidiacac ddefefecectsts iin n
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

DOI: 10.1161/CIRCGENETICS.115.001026
14
chamber morphology, contraction, and heart beat (n=10 larvae/injection batch, repeated twice).
First, we noted that nnt-a/nnt-b morphants display significant brachycardia in comparison to
their control counterparts (mean 91.5 vs. 66.8 beats/minute for control vs. morphants; p<0.0001).
Second, we observed marked contractile dysfunction in morphant hearts, as evidenced by a lack
of ventricular expansion during atrial contraction, with a concomitant generalized enlargement of
the atrium, a likely compensatory result of the ventricular defect (Figure 2; Supplementary
Movies).
Previous studies have shown that depletion of bona fide LVNC genes in zebrafish models
can result in cardiomyocyte proliferation defects9. Therefore, to investigate the cellular basis for
the contractile defects of nnt-a/nnt-b MO-injected larvae, we quantified the numbers of
proliferating versus non-proliferating cardiomyocytes. We utilized cmlc2:FUCCI, a dual
transgenic reporter line that indicates non-proliferating cardiomyocytes (mCherry-zCdt1, a G1
marker), and proliferating cardiomyocytes (Venus-hGeminin, an S/G2/M phase marker)18.
Confocal microscopy and semi-automated cell counting of larval heart sections revealed
significantly more non-proliferating cardiomyocytes in at 2 dpf in nnt-a/nnt-b morphants in
comparison to controls (Figure 3A, B; p<0.0001; n=12 larvae/injection batch, replicated).
However, the difference in cardiomyocyte counts did not persist until 3 dpf, a time point at
which morphants and controls did not display marked differences in non-proliferating
cardiomyocytes; this was consistent with a significant reduction in proliferating cardiomyocytes
in 3 dpf morphants vs. controls (Figure 3A, C; p<0.0001; n=12 larvae/injection batch, repeated
with similar results). These data suggest that the ventricular contractile defects may be the result
of aberrant cardiomyocyte proliferation in the developing heart.
he contractile defects of nnt-a/nnt-b MO-injected larvae, we quantified the numbebebersrr ooof f f
prolifereratating g vev rssusu non-proliferating cardiomyocycyytetes. We utilized cmlc2:2:FUCCI, a dual
rrrananansgs enic repepporororteteterrr liinenene thththatatat indnddicicicatatatesee nnnononon-ppprororolilil feraaatiiingg cccararardididiomomo yoyoyocycycytetet sss (m(m(mChherererryryry-z-z-zCdCdC t1t1t1, a a a G1G1G1
mmamarkrkrker), and ppprooolifferaatttinnng caararddidiomyoyoyocycyyteees (VVVeeenus-hhhGeeemimimininin,n,n, an S///G2/MMM phahahassese marrrkekeker)))188..
Confocal microscopppy y y anana d semimimi---auautomaated cell cououounting g ofoff lllaraa vav l hearart t t sesesections revealed
iigngng ifificicanantltly y y momorere nnonon-prprp ololififereratatining g g cacardrdioiomymyyococytyty eses iinn aat t 2 2 dpdppf f inin nnnnt-t a/a/nnnnt-t b b momorprpphahantnts s inin
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

DOI: 10.1161/CIRCGENETICS.115.001026
15
Discussion
Disorders with variable presentation can be challenging to diagnose. Although LVNC can be
diagnosed by echo-cardiogram, such tests are expensive and are not warranted without strong
prior indication that the patient has impaired cardiac function. Further, many cases may be
considered border-line for a noncompaction diagnosis by imaging. Single-gene tests can provide
an exact diagnosis but are likewise difficult and expensive to conduct on genetically
heterogeneous diseases where there are myriad potential genes to sequence. WES is delivered
less expensively than multiple gene tests and provides a relatively unbiased method to scan the
coding portions of the genome for causative mutations. Using this technique we were able to
identify mutations in TPM1 in one family, a rare cause of LVNC. Further, we found that 3/5 of
our families had mutations in MYH7 which likely caused their LVNC. The proportion of families
carrying MYH7 mutations was higher than expected based on previous reports. Investigation into
an expanded cohort of individuals found that 18.4% of them harbored mutations in MYH7
suggesting a potentially greater contribution of this locus than appreciated previously.
The combination of our human genetics data, the in vivo data from zebrafish embryos and
the prior observations in the mouse implicates mutations in NNT as contributory to LVNC.
Interestingly, homozygous mutations in NNT have been associated with glucocorticoid
deficiency, in which the adrenal cortex is unable to produce cortisol in response to stimulation by
ACTH40. Although no cardiac defects were reported in these subjects, it is not surprising that
defects in mitochondrial genes may lead to multiple phenotypes and that presentation may be
influenced by other factors. Multiple lines of evidence from in vivo functional testing in
zebrafish indicate that nnt suppression results in malformation of the ventricle, likely resulting in
contractility defects and brachycardia; our data suggest altered cell proliferation contributes to
dentify mutations in TPM1 in one family, a rare cause of LVNC. Further, we fouunund d d ththhatatat 333/5/5/5 fof
our famim lil es hhad mmutations in MYH7 which likely y cacaused their LVNC. TTheh proportion of famili7 es
cacaarrrryying MYH7H7H7 mumumutaaatititionononsss waww ss hihihighghgherere ttthahahan exexexpepp ctededed basssededed ooon nn prrreevevioioioususu rrrepepeportss.. InInInvevevests igggatatatioioionn n ininintot
anann eeexpxx anded cohhohorrrt offf innndiviiiduduuals foooununnd thattt 1118.4%%% of f f thththemmm hhharbbborrred mmmutttatitiiononns in MYMYM H7H7H7
uggesting a potentiialalallylyly ggreattererer ccono tributu ion of thihihiss locus thththanaa aappppreciciatatatedee ppreviously.
TThehe ccomombibinanatitionon oof f ouour r huhumaman n gegeg nenetiticscs ddatata,a,, tthehe inin vvivivoo ddatata a frfromom zzebebrarafifishsh eembmbryryyosos aandnd
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

DOI: 10.1161/CIRCGENETICS.115.001026
16
the cellular basis of this malformation. These data are consistent with other mouse and zebrafish
models of LVNC. First, Ki67-immunostaining of E13.5 Fkbp12-/- (also known as Fkbp1a) mouse
hearts demonstrated increased cell cycle activity in the myocardium in comparison to matched
controls41. Moreover, a recent study in which zebrafish were used to model LVNC showed that
suppression of prdm16 resulted in decreased cardiomyocyte counts and significantly reduced
proliferation from 2-4 dpf, an observation that was coincident with increased apoptosis9. Finally,
and most relevant to NNT, mouse models of Tafazzin, a protein critical to mitochondrial
function that when mutated gives rise to the syndromic LVNC phenotype of Barth syndrome42,
display altered cellular proliferation in embryonic hearts43. We do not know the precise
developmental mechanisms that link cardiomyocyte proliferative defects with ventricular
noncompaction defects. However, NNT is important for supporting the antioxidant capacity of
mitochondria and NNT deficient cells are likely under severe oxidative stress, a cellular context
with documented influence on cardiomyocyte cell cycle44. The implication of this pathway in the
pathogenesis of this disorder merits additional study, not least because drugs that reduce
oxidative stress may have a potential therapeutic role in select cases of LVNC. Furthermore, our
findings of brachycardia and abnormal contractility in the nnt in vivo model highlight potential
clinical implications. Numerous reports have associated LVNC in humans with depressed LV
systolic function; this has been associated independently with increased mortality in both
children and adults. Moreover, LVNC is reported increasingly in conjunction with congenital
heart disease 45–47. The implication of NNT in brachycardia may offer future understanding of
the mechanisms behind LVNC and concomitant CHD.
Identifying novel genetic loci in AD diseases is challenging because many genes contain
single, rare, protein changing mutations while far fewer have multiple mutations as would be the
developmental mechanisms that link cardiomyocyte proliferative defects with veeentntntririricuuulalalar r r
noncompmppactiiono ddefe ects. However, NNT is importaantnt for supporting the aantn ioxidant capacity of
mmmitooochondriaaa aandndnd NNNTNTNT dddefefeficici ieeentntnt cccelelellsss aaarerere llikikikelelely unnnddder sesesevevevererere oxixixidadadatititiveveve ssstrtrtress,s,, aaa cccelelellulululaaarrr cococontntntexexext t
wwiwithhh documenteddd iiinfluuuennnce ononon ccardiomomomyooocyttte cell cyyycllleee444444. ThThThe immmppplicatiiionnn ooof f tththis patatathwhwhwayyy in thhhe
pathogenesis of thiss dddisisi oro der mememeritst addditional studududy,y,y not lleaeaeastss bbecausesee dddrurr gsg that reduce
oxoxididatativive e ststreressss mmayayy hhavave e a a popop tetentntiaial l ththererapappeueutiticc rorolele iin n seselelectct ccasaseses oof f LVLVNCNC.. FFururththerermomorere,,, ouourr
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

DOI: 10.1161/CIRCGENETICS.115.001026
17
case in autosomal recessive diseases. For AD cases, such as those in the present study,
segregation in a single family likely will not provide enough statistical-power to identify a
single, novel causative gene. These approaches are limited in showing causality and biological
relevance to disease. In that regard, we and others have shown that functional studies are a
powerful and often necessary tool to both implicate novel causative genes in instances where
genetic arguments are underpowered and offer the added benefit of informing the direction of
effect of missense alleles32 33. Further development of in vivo or in vitro methods for which a
robust phenotypic readout is available will become increasingly important for the dissection of
disorders underscored by extreme genetic heterogeneity, and for which few members of the
pedigree are available for genetic analysis.
Acknowledgments: We would like to thank the families for their participation. We thank Joseph Yost for the cmlc2:GFP transgenic zebrafish line; Kenneth Poss for helpful discussions and critical reading of the manuscript; Amalia Kondyles, Kelly McKnight, and Kavita Praveen for assistance in determining morpholino efficiency; and Ben Carlson at the Duke University Light Microscopy Core Facility for assistance with image analysis. NK is a distinguished George W. Brumley Professor.
Funding Sources: Supported in part by grants from the National Human Genome Research
Institute (5 U54 HG003273, to Dr. Gibbs).
Conflict of Interest Disclosures: MNB is the founder of Codified Genomics LLC.
References:
1. Finsterer J. Cardiogenetics, neurogenetics, and pathogenetics of left ventricular hypertrabeculation/noncompaction. Pediatr Cardiol. 2009;30:659–681.
2. Sarma RJ, Chana A, Elkayam U. Left ventricular noncompaction. Prog Cardiovasc Dis.2010;52:264–273.
3. Chin TK, Perloff JK, Williams RG, Jue K, Mohrmann R. Isolated noncompaction of left ventricular myocardium. A study of eight cases. Circulation. 1990;82:507–513.
pedigree are available for genetic analysis.
AcAcAcknknknowledgdggmmments: We would like to thank the famamamilies for theiir r r papap rticippation. We thank JosephkYYYossst for the cmmmlclclc222:GFGFGFP P trtrtrananansgsgsgeneenicicc zzzebebbrararafishshh liiine; KKKennenenethtth PPPoooss foff rrr hehehelplplpfufuul dididiscscscusussisisiononons s anannd ddcrcrrittiici al reading ooof the mamm nussccrrript; Amaaaliia Kooonnndyleseses, KeKeelllllly y y McMM Knnniggght, aannddd KaKaKavvvita PPPraraaveveeennn forrr asssisis stststana ce in n n dedd teeerrmrminnininng g mooorprprphohoh liliinonono eeeffffici ieeencncncy;y aaannd BBBeenen CCCarrrlssononn aaat tht eee DDDukkeke UUUniveveverrrsittty Ligghghttt Micrososscococopypy Corore e FFaFa iciilililittyty for aasssssisisistance wiwiwiththth iiimaagege aananallylysisis. NNNK isisis aa dddisii tititingnguiishshshededed Geororgege WWW. Brumley Professor.
FuFundndiningg SoSoururceces:s: SSupuppoportrteded iinn papartrt bbyy grgranantsts ffroromm ththee NaNatitiononalal HHumumanan GGenenomomee ReReseseararchch
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

DOI: 10.1161/CIRCGENETICS.115.001026
18
4. Stöllberger C, Blazek G, Wegner C, Winkler-Dworak M, Finsterer J. Neuromuscular and cardiac comorbidity determines survival in 140 patients with left ventricular hypertrabeculation/noncompaction. Int J Cardiol. 2011;150:71–74.
5. Pignatelli RH, McMahon CJ, Dreyer WJ, Denfield SW, Price J, Belmont JW, et al. Clinical characterization of left ventricular noncompaction in children: a relatively common form of cardiomyopathy. Circulation. 2003;108:2672–2678.
6. Paterick TE, Umland MM, Jan MF, Ammar KA, Kramer C, Khandheria BK, et al. Left ventricular noncompaction: a 25-year odyssey. J Am Soc Echocardiogr. 2012;25:363-375.
7. Stöllberger C, Finsterer J. Trabeculation and left ventricular hypertrabeculation/noncompaction. J Am Soc Echocardiogr. 2004;17:1120-1121; author reply 1121.
8. Towbin JA, Lipshultz SE. Genetics of neonatal cardiomyopathy. Curr Opin Cardiol.1999;14:250–262.
9. Arndt A-K, Schafer S, Drenckhahn J-D, Sabeh MK, Plovie ER, Caliebe A, et al. Fine mapping of the 1p36 deletion syndrome identifies mutation of PRDM16 as a cause of cardiomyopathy. Am J Hum Genet. 2013;93:67–77.
10. Klaassen S, Probst S, Oechslin E, Gerull B, Krings G, Schuler P, et al. Mutations in sarcomere protein genes in left ventricular noncompaction. Circulation. 2008;117:2893–2901.
11. Luxán G, Casanova JC, Martínez-Poveda B, Prados B, D’Amato G, MacGrogan D, et al. Mutations in the NOTCH pathway regulator MIB1 cause left ventricular noncompaction cardiomyopathy. Nat Med. 2013;19:193–201.
12. Probst S, Oechslin E, Schuler P, Greutmann M, Boyé P, Knirsch W, et al. Sarcomere gene mutations in isolated left ventricular noncompaction cardiomyopathy do not predict clinical phenotype. Circ Cardiovasc Genet. 2011;4:367–374.
13. Xing Y, Ichida F, Matsuoka T, Isobe T, Ikemoto Y, Higaki T, et al. Genetic analysis in patients with left ventricular noncompaction and evidence for genetic heterogeneity. Mol Genet Metab. 2006;88:71–77.
14. Bainbridge MN, Wang M, Burgess DL, Kovar C, Rodesch MJ, D’Ascenzo M, et al. Whole exome capture in solution with 3 Gbp of data. Genome Biol. 2010;11:R62.
15. Bainbridge MN, Wang M, Wu Y, Newsham I, Muzny DM, Jefferies JL, et al. Targeted enrichment beyond the consensus coding DNA sequence exome reveals exons with higher variant densities. Genome Biol. 2011;12:R68.
9. Arndt A-K, Schafer S, Drenckhahn J-D, Sabeh MK, Plovie ER, Caliebe A, et aalal. FiFiFinenene mmmapapap ipingof the 1p36 deletion syndrome identifies mutation of PRDM16 as a cause of cardioioiommymyopopopatatathyhyhy. Am J HHumum GGene ett.. 2013;93:67–77.
10100. KKKlaassenn SS, , , PrPrProbbststst SSS,,, OeOeO chhhslslslininin EEE,, GeGeGeruuullllll BBB, Krrrinnngs GGG,, ScScSchuh lelelerr P,P,P, eet t t alalal... Muuutatatatititiononons s innn aarcccomere proteiiin gennness s ini lefefefttt ventntntrrricuuulaaar nooonnncommmpppactttioioion.n. iCiCircuuulaaation. 20000888;11117:222898993–––229290111.
111. LuLuLuxáxáxánn n G,G,G, CCCaaasanananovovova JCJCJC, MaMaMartrttíníníneeez-PPPovovovedededa aa BB,B, PPPraraadododos ss B,B,B, DDD’A’A’Amamamatototo GGG, , , MaMaMacGcGGrororogagagan n n D,D,, eet t alala ...Mutations in the NOTOTOTCHCHC ppatthwhwhwayay reggulu ator MIBBB111 cac use leleleftftft vventriculullararar noncompaction cardiomyopathy. NaNaat t t MeMeMed.d.d 2220101013;3;3;191919:1:11939393–2–2–2010101..
1212 PrProbobstst SS OeOechchslslinin EE ScSchuhulelerr PP GGrereututmamannnn MM BoBoyéyé PP KnKnirirscschh WW eett alal SaSarcrcomomereree gegenene
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

DOI: 10.1161/CIRCGENETICS.115.001026
19
16. Yang Y, Muzny DM, Reid JG, Bainbridge MN, Willis A, Ward PA, et al. Clinical whole-exome sequencing for the diagnosis of mendelian disorders. N Engl J Med. 2013;369:1502–1511.
17. Huang C-J, Tu C-T, Hsiao C-D, Hsieh F-J, Tsai H-J. Germ-line transmission of a myocardium-specific GFP transgene reveals critical regulatory elements in the cardiac myosin light chain 2 promoter of zebrafish. Dev Dyn Off Publ Am Assoc Anat. 2003;228:30–40.
18. Choi W-Y, Gemberling M, Wang J, Holdway JE, Shen M-C, Karlstrom RO, et al. In vivo monitoring of cardiomyocyte proliferation to identify chemical modifiers of heart regeneration. Dev Camb Engl. 2013;140:660–666.
19. Morrison AC, Bare LA, Chambless LE, Ellis SG, Malloy M, Kane JP, et al. Prediction of coronary heart disease risk using a genetic risk score: the Atherosclerosis Risk in Communities Study. Am J Epidemiol. 2007;166:28–35.
20. Liew CC, Sole MJ, Yamauchi-Takihara K, Kellam B, Anderson DH, Lin LP, et al. Complete sequence and organization of the human cardiac beta-myosin heavy chain gene. Nucleic Acids Res. 1990;18:3647–3651.
21. Kamisago M, Sharma SD, DePalma SR, Solomon S, Sharma P, McDonough B, et al. Mutations in sarcomere protein genes as a cause of dilated cardiomyopathy. N Engl J Med. 2000;343:1688–1696.
22. Chang B, Nishizawa T, Furutani M, Fujiki A, Tani M, Kawaguchi M, et al. Identification of a novel TPM1 mutation in a family with left ventricular noncompaction and sudden death. Mol Genet Metab. 2011;102:200–206.
23. Morita H, Rehm HL, Menesses A, McDonough B, Roberts AE, Kucherlapati R, et al. Shared genetic causes of cardiac hypertrophy in children and adults. N Engl J Med. 2008;358:1899–1908.
24. Cheng G, Porter JD. Transcriptional profile of rat extraocular muscle by serial analysis of gene expression. Invest Ophthalmol Vis Sci. 2002;43:1048–1058.
25. Arad M, Penas-Lado M, Monserrat L, Maron BJ, Sherrid M, Ho CY, et al. Gene mutations in apical hypertrophic cardiomyopathy. Circulation. 2005;112:2805–2811.
26. Frisso G, Limongelli G, Pacileo G, Del Giudice A, Forgione L, Calabrò P, et al. A child cohort study from southern Italy enlarges the genetic spectrum of hypertrophic cardiomyopathy. Clin Genet. 2009;76:91–101.
27. Kaneda T, Naruse C, Kawashima A, Fujino N, Oshima T, Namura M, et al. A novel beta-myosin heavy chain gene mutation, p.Met531Arg, identified in isolated left ventricular non-compaction in humans, results in left ventricular hypertrophy that progresses to dilation in a mouse model. Clin Sci Lond Engl. 1979. 2008;114:431–440.
equence and organization of the human cardiac beta-myosin heavy chain gene. Nuuuclclc eiic c c AcAciddssRes. 1990;18:3647–3651.
21. Kaamimisagogo MM,,, ShS arma SD, DePalma SR, Solommono S, Sharma P, McDoDonough B, et al. MuMuutatatatititions ininin ssarrcococomere protein genes as a cause ofofo dddilated cardiommmyoy paathththy. N Engl J Med. 20200000000;343:168888–8–1616169666...
22222. ChCC ang B,, Nisshhhizawawawa T, FuFuFururur tanii MMM, FFFujikikiki A, TaTaTanii MMM, KaKaKawaguguguchc i MMM, et alall... Idennntififificccatttioi n offf a nonoovevevell l TPTPTPM1M1M mmmututtatata ioioion ininin a fffamamamililily yy withthth lllefefeft tt veveventntntririricuuulalalar rr nonononcncn omomompapapactctctioioionn n annnd dd susuuddddddenenen dddeaeaaththth. . MoMoMol Genet Metab. 2011;1;1;102020 :200–2–2–206060 .
2323. . MoMoriritata HH,,, ReRehmhm HHL,L,, MMenenesesseses s A,A,, MMcDcDononououghghg BB,,, RoRobebertrtss AEAE,,, KuKuchchererlalapapap titi RR,,, etet aal.l. SShaharereddgegenenetiticc cacaususeses ooff cacardrdiaiacc hyhypepertrtrorophphyy inin cchihildldrerenn anandd adadulultstsyyy NN EnEnglgl JJ MMeded 20200808;3;35858:1:1898999–
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

DOI: 10.1161/CIRCGENETICS.115.001026
20
28. Waldmüller S, Erdmann J, Binner P, Gelbrich G, Pankuweit S, Geier C, et al. Novel correlations between the genotype and the phenotype of hypertrophic and dilated cardiomyopathy: results from the German Competence Network Heart Failure. Eur J Heart Fail.2011;13:1185–1192.
29. Huang T-T, Naeemuddin M, Elchuri S, Yamaguchi M, Kozy HM, Carlson EJ, et al. Genetic modifiers of the phenotype of mice deficient in mitochondrial superoxide dismutase. Hum Mol Genet. 2006;15:1187–1194.
30. Hoit BD, Kiatchoosakun S, Restivo J, Kirkpatrick D, Olszens K, Shao H, et al. Naturally occurring variation in cardiovascular traits among inbred mouse strains. Genomics. 2002;79:679–685.
31. Zieger B, Ware J. Cloning and deduced amino acid sequence of human nicotinamide nucleotide transhydrogenase. DNA Seq J DNA Seq Mapp. 1997;7:369–373.
32. Hoek JB, Rydström J. Physiological roles of nicotinamide nucleotide transhydrogenase. Biochem J. 1988;254:1–10.
33. Davis EE, Frangakis S, Katsanis N. Interpreting human genetic variation with in vivo zebrafish assays. Biochim Biophys Acta. 2014;1842:1960–1970.
34. Niederriter AR, Davis EE, Golzio C, Oh EC, Tsai I-C, Katsanis N. In vivo modeling of the morbid human genome using Danio rerio. J Vis Exp JoVE. 2013;e50338.
35. Borck G, Hög F, Dentici ML, Tan PL, Sowada N, Medeira A, et al. BRF1 mutations alter RNA polymerase III-dependent transcription and cause neurodevelopmental anomalies. Genome Res. 2015;25:155–166.
36. Khuchua Z, Yue Z, Batts L, Strauss AW. A zebrafish model of human Barth syndrome reveals the essential role of tafazzin in cardiac development and function. Circ Res.2006;99:201–208.
37. Van der Meer DLM, Marques IJ, Leito JTD, Besser J, Bakkers J, Schoonheere E, et al. Zebrafish cypher is important for somite formation and heart development. Dev Biol.2006;299:356–372.
38. Vogel B, Meder B, Just S, Laufer C, Berger I, Weber S, et al. In-vivo characterization of human dilated cardiomyopathy genes in zebrafish. Biochem Biophys Res Commun.2009;390:516–522.
39. Yeo S-Y, Kim M, Kim H-S, Huh T-L, Chitnis AB. Fluorescent protein expression driven by her4 regulatory elements reveals the spatiotemporal pattern of Notch signaling in the nervous system of zebrafish embryos. Dev Biol. 2007;301:555–567.
Biochem J. 1988;254:1–10. JJ
33. Davis EE, Frangakis S, Katsanis N. Interpreting human genetic variation withh iiinnn vivivivovovo zebrafisish h assaays. BiB ochim Biophys Acta. 2014;184242:1: 960–1970.
34344. NNNiederriteteer ARARAR, DaDaDavivivisss EEEE , , , GoGoGolzlzlzioioio CCC, OhOhh EEEC, TTTsaai I-I-C,C,C, KKKataa saaannnis s s N.N.N IIInnn vivv vooo mmmodododelele innng g g ofofof ttthehehe momm rrrbid human gggennnommee uuusini ggg DDDaniiiooo rerrriooo. J Viiis Exxxppp JooVEVEVE. 20202013;eee555033888.
35. BoBoBorcrcrckkk G,G,G, HHHögögög FFF,, DeDeDentntntici MLMLML,,, TaTaTan PLPLPL,,, SoSoSowawawadadada NNN, ,, MeMeMedededeiririra a a A,A, eeett t alalal... BRBRBRF1F1F1 mmmutututatatatiioionsnsns aaaltltltererer RNA polymerase IIIIII-d-d-depependeeentntnt traanscripption andd cccaua se neueueurororoded velopmpmmenenental anomalies. GenomeRes. 2015;25:155–111666666...
3636 KhKhucuchuhuaa ZZ YYueue ZZ BaBattttss LL SStrtrauaussss AAWW AA zzebebrarafifishsh mmododelel ooff huhumamann BaBartrthh sysyndndroromeme
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

DOI: 10.1161/CIRCGENETICS.115.001026
21
40. Meimaridou E, Kowalczyk J, Guasti L, Hughes CR, Wagner F, Frommolt P, et al. Mutations in NNT encoding nicotinamide nucleotide transhydrogenase cause familial glucocorticoid deficiency. Nat Genet. 2012;44:740–742.
41. Chen H, Zhang W, Li D, Cordes TM, Mark Payne R, Shou W. Analysis of ventricular hypertrabeculation and noncompaction using genetically engineered mouse models. Pediatr Cardiol. 2009;30:626–634.
42. Bione S, D’Adamo P, Maestrini E, Gedeon AK, Bolhuis PA, Toniolo D. A novel X-linked gene, G4.5. is responsible for Barth syndrome. Nat Genet. 1996;12:385–389.
43. Phoon CK, Acehan D, Schlame M, Stokes DL, Edelman-Novemsky I, Yu D, et al. Tafazzin knockdown in mice leads to a developmental cardiomyopathy with early diastolic dysfunction preceding myocardial noncompaction. J Am Heart Assoc. 2012;1. pii: jah3-e000455.
44. Puente BN, Kimura W, Muralidhar SA, Moon J, Amatruda JF, Phelps KL, et al. The oxygen-rich postnatal environment induces cardiomyocyte cell-cycle arrest through DNA damage response. Cell. 2014;157:565–579.
45. Brescia ST, Rossano JW, Pignatelli R, Jefferies JL, Price JF, Decker JA, et al. Mortality and sudden death in pediatric left ventricular noncompaction in a tertiary referral center. Circulation.2013;127:2202–2208.
46. Aras D, Tufekcioglu O, Ergun K, Ozeke O, Yildiz A, Topaloglu S, et al. Clinical features of isolated ventricular noncompaction in adults long-term clinical course, echocardiographic properties, and predictors of left ventricular failure. J Card Fail. 2006;12:726–733.
47. Vermeer AMC, van Engelen K, Postma AV, Baars MJH, Christiaans I, De Haij S, et al. Ebstein anomaly associated with left ventricular noncompaction: an autosomal dominant condition that can be caused by mutations in MYH7. Am J Med Genet C Semin Med Genet.2013;163C:178–184.
esponse. Cell. 2014;157:565–579.
45. Brescia ST, Rossano JW, Pignatelli R, Jefferies JL, Price JF, Decker JA, et al. MoMoMortrtrtalalalititityy y anananddd uddenn deae thh in pepep diatric left ventricular noncompmppacaction in a tertiary reffererral center. Circulation.
200131313;1;1;12727:2:222020202––22222208.
4464 . Aras D, Tufeeekkcciogggluu u O,O EEErrgrgun KKK, OzOzOzeke OOO, Yiiildddiz A,AA, TTTopopopalogggluu S, eeet al. CCCllinicaaal l feff aaatuuures ooff sssolololaatated ventricuuulaaar noooncccompppacacaction iiin nn adadadults looong---teeermmm ccclilliniiicaaal cooouururse, eeechhhoccacarrrdiogrrrapapaphhicc
propoppererertititieseses, , , ananand d d prprpredededicicctttorsrsrs of leleleftftft vvenenentriccculululararar fffaiaiailululurerere. J JJ CaCaCardrdrd FFFaiaiail.l 2220000006;6;6;121212:777262626–7–7–7333333...
47. Vermeer AMC,, vvvananan EEEngngngelee enenen KKK,,, PoPoPostststmamama AAAVVV, BBBaaaaaarsrsrs MMMJHJHJH,, , ChChChririristststiaiaiaanananss s I,I,I, DDDe ee Haij S, et al. EbEbststeiein n ananomomalaly y y asassosociciatateded wwitith h leleftft vvenentrtriciculularar nnononcocompmppacactitionon:: anan aaututososomomalal ddomomininanant t cocondndititioionn ththatat ccanan bbee cacaususeded bbyy mumutatatitiononss inin MMYHYH77 AmAm JJ MMeded GGenenetet CC SSememinin MMeded GGenenetet
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

DOI: 10.1161/CIRCGENETICS.115.001026
22
Table 1: LVNC Family sequencing results - for each of the five LVNC families in this study, the number of affected and unaffected individuals sequenced, the implicated candidate disease causing gene, mutation and in silico predictions are given. Pedigrees are available in Supplementary Figure 1.
Family ID
Affected Individuals Sequenced /Total in pedigree
Unaffected Individuals Sequenced /Total in pedigree
Gene Mutation(HG19 Coordinates)
PolyPhen-2 Prediction (score)
Mutation TasterPrediction
A 4/5 1/4 TPM1 p.D275H (chr15:g.23857430C>C)
Probably damaging (0.992)
Disease causing
B 5/7 0/2 MYH7 p.S648L (chr14:g.23896462G>A)
Possibly damaging (0.906)
Disease causing
C 2/2 2/6 NNT c.638_639insT (chr5:g.43619172insT) N/A Disease
causing
D 2/2 0/2 MYH7 p.Q163P (chr14:g.23901862T>G)
Probably damaging (0.979)
Disease causing
E 4/4 3/4 MYH7 p.E700G (chr14:g.23895236T>C)
Probably damaging (0.997)
Disease causing
4/5 1/4 TPM1 p.D275H(chr15:g.23857430C>C)
Probably damaginining g g (0.992)
DiDiDiseseseasasaseeecausing
5/7 0/0//222 MYMYMYH7H7H7 p.S6648488L (chr1114::g.22383838969 466622G2G>AAA)
PoPP sssibibiblyll dammmagagagininingg g (0.990006)))
DiDiDiseeeasasaseeecaauuusinggg
2/2/2/222 2/2/2/666 NNNNNNTTT c.c.c 6363638_8_8 6363639i9i9insnsTT T (chrhr5:55 g.436191727272ini sT) N/N/N/AAA DiDiiseseseasasaseee
causing
/2/2/22 /0/0/22 MYMYH7H7 ppp.Q1Q1Q1636363PPP (c(chrhr1414:g:g 223939010186862T2T>G>G))
PrPrProbobobababablylyly dddamamamagagaginininggg (0(0 997979))
DiDiDiseseseasasaseeecacaususiningg
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

DOI: 10.1161/CIRCGENETICS.115.001026
23
Figure Legends:
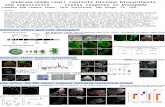
Figure 1: Genetic and Functional evaluation of NNT in LVNC. A. Schematic of the human
NNT locus on human chromosome 5 (top) showing non-coding exons (white boxes), and coding
exons (blue boxes). Double slash indicates large introns (> 5kb). A schematic of NNT protein is
shown below with functional domains Alanine dehydrogenase/PNT, N-terminal domain
(AlaDh_PNT_N, shown in green), Alanine dehydrogenase/PNT, C-terminal domain
(AlaDh_PNT_N, shown in purple) and twelve transmembrane domains (orange). The two
mutations identified in LVNC families are shown with asterisks. B. Schematic of NNT function
in mitochondria to regenerate NADPH in the cell; NADPH is then used to regenerate
Thioredoxin Reductase 2 (TrxR-2) and glutathione reductase (GR). C. Lateral view of
representative 3 dpf control (top) and nnt-a/b morphant (bottom) larvae showing cardiac edema
(arrow). D. In vivo complementation assay of NNT p.D277Y; quantification of cardiac edema in
3 dpf morphant zebrafish larvae (5 ng nnt-a/b MOs) with and without addition of 200 pg human
wild type NNT RNA (WT), and mutant RNA (p.D277Y). Statistically significant differences
were calculated with 2 tests and p<0.0001 are indicated (*); n=46-51 embryos/injection batch,
repeated three times with masked scoring.
Figure 2: nnt-a/nnt-b morphants display contractile dysfunction. Representative live ventral
views of 2 dpf cmlc2:GFP larvae at ventricular systole (top panels) and ventricular diastole
(bottom panels). Ventricles in control larvae expand normally as the atrium contracts (compare
white dashed circle to violet dashed circle in the bottom inset); ventricles in nnt-a/nnt-b
morphants fail to expand during atrial contraction (5 ng each MO injected/embryo). Dashed box
n mitochondria to regenerate NADPH in the cell; NADPH is then used to regeneeeraraattete
Thioreedodoxin ReR duductase 2 (TrxR-2) and glutathionee rreductase (GR). C. LaLateral view of
eeeprrresentativevee 333 dddpfpp cccononontrtrtrololol (tooop)p)p) aaandndn nnnnnnt-tt a/a/a/bbb mmmorphphphant t (b(b(bototottototom)m)) llarararvavav e e e shshshowininng g g cacacardrdr iaaaccc edededememema aa
aaarrrroowow). D. In vivvvooo commmppplemmmenenentat tionnn assssaaay oof NNNNTTT ppp.D2D2D277777YYY; quaaantttificatttiooon ofofof cardiiaiaccc eeedeeemaa innn
3 dpf morphant zebrrrafafafisisi h larvvvaaaeee (5(5 ng g nnn t-a/b MOOOs)s)) with aaandndnd wwithoutu aaaddddddition of 200 pg human
wiwildld tytyypepep NNNNTT RNRNAA (((WTWT),),), aandnd mmututanant t RNRNA A (p(p(p.D.D27277Y7Y).).) SStatatitiststicicalallylyy ssigiggninifificacantnt yyy didiffffererenencecess
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

DOI: 10.1161/CIRCGENETICS.115.001026
24
corresponds to the magnified image to the right of each large panel; L, left; R, right; A, atrium;
V, ventricle; scale bars, 100 m. nnt-a/nnt-b morphants also display brachycardia (mean 91.5 vs.
66.8 beats/minute, controls vs. morphants; p<0.0001; student’s t-test; n=10 larvae/injection,
repeated twice; see Supplemental movies).
Figure 3: Suppression of nnt-a/nnt-b in zebrafish results in altered cardiomyocyte proliferation.
A. Representative maximum intensity projections of cardiac sections from control and nnt-a/nnt-
b morphants (5 ng each MO injected/embryo) at 2 dpf and 3 dpf were used to monitor non-
proliferating (cmlc2:mCherry-zCdt1, red, G1 phase of the cell cycle) and proliferating
(cmlc2:Venus-hGeminin, green, S/G2/M phase of the cell cycle) cardiomyocyte counts.
Cardiomyocytes with only the green signal were counted as proliferating (exemplified by 2 dpf
nnt-a/nnt-b or 3 dpf control panels). Scale bars, 50 m; arrowheads point to the ventricle (V); or
atrium (A). B. Quantification of non-proliferating cardiomyocytes as indicated by red+green- or
red+green+ cells. C. Quantification of proliferating cardiomyocytes as indicated by red-green+
cells. p<0.0001; student’s t-test; n=12 larvae/injection batch, repeated once with similar results.
Error bars represent standard error of the mean (sem).
cmlc2:Venus-hGeminin, green, S/G2/M phase of the cell cycle) cardiomyocyte cococounununtststs...
Cardiooomymymyocoo ytyty ess wwwith only the green signal were ccououounted as proliferating g g (e(( xemplified by 2 dpf
nnnnnt---a/nnt-b or 333 dddppfp cccononontrtrtrololol ppanananelelels)s)s). ScScScalalale bababarsss, 5000 m;m;; aaarrrrrrowowowheeeaaadsss popopoinininttt ttto tthehehe vvvenenentrtrtriccclelele (((V)V)V);;; oroo
atttririr umumum (A). B.B.B. QQQuaaantiffficccationn n oofof nonnn---prprrollliferrrattting caaardddioioiomymyyocococyty ess aaas indddicccateeed d d byb rededed+++grrreeen- ooror
ed+green+ cells. C.. QuQuQuananantititififificacacatititiononon ooof f f prprprolololifififerereratatatinininggg cacacardrdrdioioi mymymyocococytytyteseses aaas s s ininindididicacacatttedee by red-green+
cececellllllsss. ppp<0<0<0 00.00000001;1;1; ssstututudededentntnt’sss ttt--teteteststst;;; nnn=1=1=1222 lalalarvrvrvaeaeae/i/i/injnjnjececectititiononon bbbatatatchchch, rererepepepeatatatededed oooncncnceee wiwiwiththth sssimimimilililararar rrresesesululultststs.
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

Gibbs and John L. JefferiesWang, Huyen Dinh, Donna Muzny, Ricardo Pignatelli, Nicholas Katsanis, Eric Boerwinkle, Richard
Matthew N. Bainbridge, Erica E. Davis, Wen-Yee Choi, Amy Dickson, Hugo R. Martinez, Min Are Associated with Left Ventricular NoncompactionNNTLoss of Function Mutations in
Print ISSN: 1942-325X. Online ISSN: 1942-3268 Copyright © 2015 American Heart Association, Inc. All rights reserved.
TX 75231is published by the American Heart Association, 7272 Greenville Avenue, Dallas,Circulation: Cardiovascular Genetics
published online May 29, 2015;Circ Cardiovasc Genet.
http://circgenetics.ahajournals.org/content/early/2015/05/29/CIRCGENETICS.115.001026World Wide Web at:
The online version of this article, along with updated information and services, is located on the
http://circgenetics.ahajournals.org/content/suppl/2015/05/29/CIRCGENETICS.115.001026.DC1Data Supplement (unedited) at:
http://circgenetics.ahajournals.org//subscriptions/
is online at: Circulation: Cardiovascular Genetics Information about subscribing to Subscriptions:
http://www.lww.com/reprints Information about reprints can be found online at: Reprints:
document. Permissions and Rights Question and Answer this process is available in the
located, click Request Permissions in the middle column of the Web page under Services. Further information aboutnot the Editorial Office. Once the online version of the published article for which permission is being requested is
can be obtained via RightsLink, a service of the Copyright Clearance Center,Circulation: Cardiovascular Genetics Requests for permissions to reproduce figures, tables, or portions of articles originally published inPermissions:
by guest on June 7, 2018http://circgenetics.ahajournals.org/
Dow
nloaded from

SUPPLEMENTAL MATERIAL
Supplementary Tables
Supplementary Table 1. All genes with shared rare mutations between WES affecteds and not shared
by WES unaffecteds. Bold, implicated LVNC gene. Underline, known cardiomyopathy gene.
Family Genes
A CANT1, CHRNE, ERCC8, EVPL, EXOC7, FOXA2, GPR98, KIAA1324, MRPS15, MTG1, MYH6, NBPF3, PDE4DIP, RSPH4A, SLC25A23, TMC1, TOR1B, TPM1, TRIP10, USP8, ZNF280D, ZNF644
B ANKRD36, CCDC150, CDC27, CNTNAP4, DCHS2, DNAH10, DRG1, MAL2, MYH7, OR11G2, PSAPL1, RP1L1, TRIL
C A4GNT, AGBL5, ALPK2, BHMT2, C2orf16, COL5A3, CR2, FBXO4, FCRL2, FLG, FLG, FRRS1, GSDMC, MAP1S, MMP12, NNT*, OR52W1, PNN, PSAP, RSPH3, TOR1AIP1, VPS41, ZNF180, ZNF181, ZNF229
D ACOT4, AHI1, ANAPC2, ASXL3, ATP4B, BBX, BMP1, BTBD10, CACNG2, CADM4, CASZ1, CCDC158, CCDC66, CHID1, COL6A1, CPE, DCX, DMXL2, DNM1, DUPD1, EPPK1, FBRSL1, FCHO2, GFM2, GPC1, GPR176, GREB1, HAL, HCCS, HOXB2, HSP90B1, INPP5D, KIF26A, LAMA1, LSG1, MAP2, MAST3, MED16, MEX3A, MRPL39, MTHFD1, MTHFD1L, MYH7, N4BP2, NAP1L1, NAV1, NEDD4L, NOTCH3, OR52I2, OSMR, PARP1, PEPD, PLXNA4, PPIP5K2, PPP1R12C, PTH2R, SBF1, SCN11A, SCN7A, SEMA3G, SI, SLC16A3, SLC22A5, SLC45A2, SLC45A4, SMPD4, SPEF2, STRAP, TCEA1, TIMM44, TMEM180, TOP2A, TRIM59, UMOD, UNKL, WDFY4, WDR4, ZNF354B, ZNF451, ZNF646
E ARSB, BRCA2, FAM38A, FRRS1, MAGI3, MYH7, NOVA1, RNF6, SLAIN1, TEP1

Supplementary Table 2. LVNC cohort data. LVNC cohort used for NNT sequencing. All cohort members
were diagnosed with LVNC. Gender and additional phenotype information is listed. Samples used for
MYH7 sequencing are indicated, with any discovered MYH7 mutations and any previous descriptions.
Patient ID# Gender Additional Phenotype
Used in MYH7 Sequencing Study
MYH7 Mutation(s)
Previous Description
L051 F
DISEASE ONSET-BIRTH. POST DELIVERY, AVA PRESENTED WITH PERSISTENT HYPOXIA AND RESPIRATORY DISTRESS, PULMONARY HYPERTENSION, HYPOTENSION AND WAS DIAGNOSED WITH LVNC AND VSD'S. DEVELOPED THROMBOSIS IN LEFT ATRIUM AND LEFT VENTRICLE. 1 N/A
L100 F
LVNC, VENTRICULAR SEPTAL DEFECT, SMALL NORMAL LV SIZE, LVEF 47% BY MRI 1 p.Q163P
L012 M Present at 15 days age-cyanosis 1 N/A
L030 F Genetic Analysis for alpha- dystrophin 1
p.R1045H, p.T547A
Frisso et al. CLIN GENET 2009
L034 M
AGE OF DISEASE ONSET: 6 YEARS, Dyspnea, sporadic palpitations. NYHA class II. Echocardiogram was compatible with non-compaction cardiomyopathy, mitral stenosis and mild mitral regurgitation. 1 N/A
L040 F
presented with cardiac arrest on day 7 of life. >25 of possible arrest with ischmicencepholopathy. Initial LV SF<15%-3 days later= he fxn but poss/prob noncompaction seen on 1 N/A

echo.
L044 F asymptomatic, identified by echocardiogram 1 N/A
L046 F age 13 so born in 1991;vheart failure 1 N/A
L054 M NON COMPACTION CM, HOCM 1 N/A
L058 F
1 p.M531R Kaneda et al. CS(L), 2008
L060 F
1 N/A L061 M
1 N/A
L131 F
CARDIAC US AT AGES 1 YEAR AND 6 YEARS - NORMAL; AT AGE OF 8, SYSTOLIC MURMUR WAS HEARD AND LEFT ATRIUM WAS MILDLY ENLARGED; 6/2005, CATHETHRISATION AND IN THE APEX OF LEFT VENTRICLE TRABECULATIONS WERE SEEN, HAS ANTICOAGULANT MEDICATION 1 N/A
L069 M
1 N/A L119 M patient is proband 1 N/A
L086 F
pronounced biventricular involvement. Died in 2002. 1 N/A
L025 F
21 MONTHS, severe left ventricular dysfunction; isolated left ventricular non-compaction. 1 N/A
L092 F
Symptoms include: heart failure, Dx of ventricular noncompaction 1 N/A
L094 F
LV non-compaction ascertained 2003 (age 34). Neonatal VSD, unrepaired. 1 N/A
L095 F VSD, LV DYSFUNCTION 1 N/A L098 M WPW, DCM, LVNC 1 N/A

L099 M
1 N/A L102 M
0 N/A
L104 F DX LVNC, NO SYMPTOMS 0 N/A L109 M Mitral Valve Prolapse 1 N/A L110 F LVNC, HCM 1 N/A L111 M LVNC, HCM 1 N/A L112 F
1 N/A
L117
1 p.R904H Waldmüller et al. EJHF , 2011
L122
1 N/A L123 M DX LVNC, NO SYMPTOMS 1 N/A L127 M
1 p.E1350K
L132 F
1 N/A
L134 M
Brother is affected with DCM, parents are first cousins 1 N/A
L135 M mother died at age 37 of cardiomyopathy 1 N/A
L137 F
Both siblings and mother affected with LVNC and HCM 1 p.L915P
L139 F Sister died at day 17 with LVNC 1 N/A
L141 M
0 N/A L143 M
0 N/A
L008 M Dilated Cardiomyopathy and hypotonia 1 N/A
L009 M
Present in infancy with cyanosis as a result of congenital heart disease. 1 N/A
L010 M
Present in 1997 (6 years old) with acute myocarditis associated with bradycardia due to varying heart block. 1 N/A
L011 M
Diagnosed in infancy with cyanotic congenital heart disease 1 N/A
L014 M Present at age 9 (July 01) with congestive failure 1 N/A
L020 M
0 N/A

L022 F
POST-PART. CM DX1990. NON-COMPACTION OF LV DX 1996. 1 p.L961P
L>R has been reported http://genepath.med.harvard.edu/seidman/cg3/muts/MYH7_mutations_TOC.html
L023 M
CHF. DCM WITH SPONG MYOCARDIUM AND VACTERL. 1 N/A
L026 M ISOLATED LVNC WITH HLHS 1 N/A
L031 F
Diagnosed with LVNC following a sudden cardiac arrest and her daughters have been diagnosed with an asymptomatic form on echocardiogram 1 p.R243H
http://genepath.med.harvard.edu/seidman/cg3/muts/MYH7_mutations_TOC.html
L050 F
0 N/A L079 F LVNC. 3 BEAT RUN OF VT 1 N/A L084 M
0 N/A
L085 F
mother and daughter both have LVNC, found to have same pathology 1 N/A
L087 F
CONGESTIVE HEART FAILURE AT 3 MONTHS OLD. PRESENTED WITH PULMONARY EDEMA. 1 N/A
L121 M
0 N/A
L129 M
INCREASED DYSPNEA AND LVH ON ECHOCARDIOGRAM. MRI SHOWS LVNC 1 N/A
L130 M
Early puberty, Pediatrician heard cardiac click sent for echo, LV non-compaction found; subsequently dad had echo showing DCM 1 N/A

Supplementary Table 3. Genes and rare variants identified in LVNC Family C that are shared between
WES affecteds, but not shared by WES unaffected.
Gene name
Nucleotide: Amino acid
Variant (heterozygous)
Nucleotide sequence
Mutation type Human disease
association (OMIM)
Mouse mutant phenotypes (MGI)
A4GNT c.G803T: p.W268L
NM_016161 nonsynonymous
cardiovascular (increased angiogenesis in the gastric mucosa), cellular, digestive/alimentary, immune, tumorigenesis
AGBL5 c.791_794del:
p.264_265del NM_0218314 frameshift
deletion
ALPK2 c.C772T: p.R258C NM_052947 nonsynonymous
BHMT2 c.C565T: p.P189S NM_017614 nonsynonymous
C2orf16 c.C730G: p.R244G NM_032266 nonsynonymous
COL5A3 c.C1648T: p.R550C
NM_015719 nonsynonymous
adipose, cellular, endocrine/exocrine, growth/size, homeostasis, integument, muscle
CR2 c.G641A: p.R214H
NM_001877 nonsynonymous Immunodeficiency, common variable, 7 [MIM614699]
hematopoietic, immune, mortality/aging
FBXO4 c.A676G: p.N226D
NM_012176 nonsynonymous
cellular, hematopoietic, immune, mortality/aging, tumorigenesis
FCRL2 c.G952T: p.E318X NM_030764 nonsense
hematopoietic, immune
FLG c.G4533C:
p.R1511S NM_002016 nonsynonymous Ichthyosis
vulgaris [MIM 146700]
cellular, craniofacial, growth/size, hearing/vestibular/ear, hematopoietic, homeostasis, immune, integument, mortality/aging
FLG c.G2280C: p.Q760H
NM_002016 nonsynonymous
FRRS1 c.T1292C: p.M431T
NM_001013660
nonsynonymous
mortality/aging
GSDMC c.G590A: p.S197N
NM_031415 nonsynonymous
behavior, craniofacial, digestive/alimentary, growth/size, hearing/vestibular/ear, homeostasis, integument, mortality/aging, respiratory

MAP1S c.C128G: p.S43C NM_018174 nonsynonymous
cardiovascular (cardiomyocytes in neonates and adult exhibit a 3-fold increase in dysfunctional mitochondria compared with wild-type mice), cellular, muscle
MMP12 c.G697T: p.A233S,
uc001phk.3 nonsynonymous
hematopoietic, homeostasis, immune, nervous system, reproductive
NNT* c.638_639insT:
p.R213fs NM_012343 frameshift
deletion Glucocorticoid deficiency 4 [MIM614736]
cellular, endocrine/exocrine, homeostasis, cardiovascular (lower LV shortening and increased LV weight, higher aortic ejection time)
OR52W1 c.G923A: p.R308Q
NM_00100517 nonsynonymous
PNN c.A1264G: p.S422G
NM_002687 nonsynonymous
embryogenesis, mortality/aging, vision/eye
PSAP c.C409G: p.L137V
NM_001042465V
nonsynonymous Combined SAP deficiency [MIM611721] Gaucher disease, atypical [610539]
Krabbe disease, atypical [MIM611722]
Metachromatic leukodystrophy due to SAP-b deficiency [MIM249900]
behavior, growth/size, hematopoietic, homeostasis, immune, mortality/aging, nervous system, reproductive, skeleton
RSPH3 c.C1126T: p.R376W
NM_031924 nonsynonymous
TOR1AIP1 c.T983G: p.I328S NM_015602 nonsynonymous
cellular, mortality/aging
VPS41 c.C881T: p.T294M
NM_014396 nonsynonymous
cellular, embryogenesis, mortality/aging
ZNF180 c.A1513G: p.S505G NM_013256 nonsynonymous
ZNF181 c.A352G: p.K118E
NM_001029997
nonsynonymous

ZNF229 c.G1537C: p.G513R
NM_014518 nonsynonymous

Supplementary Table 4. Morpholinos used for in vivo functional assays. Relationship of the variants
identified in LVNC family C to the zebrafish morpholinos (MO)s used for assays of physiological
relevance and variant pathogenicity. *Variants in HACL1, NAV2, RAB27A, and RBM28 are present in
<0.01% of controls (Exome Variant Server), shared by the affected individuals but do not segregate fully
with disease in the pedigree. **notch1a MO was published previously1.
Human gene
Variant in
LVNC Family
C*
Zebrafish
gene
MO sequence
NNT c.638_639insTp.R213fs
nnt-a ACCATTGTCAGAGGACTTACCCAGC
nnt-b GTAAGTGTGGATGTTTCACCTTCGC
HACL1 c.G1036T p.E346X
hacl1 GTTGAAATATTCATACCGTCCAGTC
NAV2 c.C3071G p.S1024W
nav2 TTTTGAAGTGTTTTTACCTTCAGCT
RAB27A c.C518G p.A173G
rab27a AAGGATTTTCTTACCCGTATTTCTC
RBM28 c.G1488C p.K496N
rbm28 ACTTGGTCAAACTTTTACCTGGTTT
NOTCH1 Not applicable notch1a** GAAACGGTTCATAACTCCGCCTCGG

Supplementary Figures Supplementary Figure 1A. Pedigree, sequencing platform and sequence coverage for each family A.
LVNC status for each individual is given: Black (affected), white (unaffected), gray (unknown status), *
indicates subject was capture sequenced.
1* 2*
3 4*
5* 6* 7 8 9

Supplementary Figure 1B. Pedigree, sequencing platform and sequence coverage for family B.
6
4
5*
9*
8*
7*
3*
2
1*

Supplementary Figure 1C. Pedigree, sequencing platform and sequence coverage for family C.
3
4
5*
6*
7*
8*
1
2

Supplementary Figure 1D. Pedigree, sequencing platform and sequence coverage for family D.
1
4*
3*
2

Supplementary Figure 1D. Pedigree, sequencing platform and sequence coverage for family D.
7*
8*
3
4*
5*
6*
1*
2*

Supplementary Figure 2. Knockdown efficiency of morpholinos
A. Schematic depicts the targeted D. rerio locus for each gene under investigation; white boxes,
untranslated regions; blue boxes, coding exons; red box, morpholino (MO) target site; arrows indicate
the position of RT-PCR primers (see B). B. Agarose gel images of nnt-a, nnt-b, hacl1, nav2, rab27a, and
rbm28 RT-PCR products show reduced (nnt-a, nnt-b) or alternatively spliced (hacl1, nav2, rab27a,
rbm28) transcript in MO-injected embryos. -actin is shown as a loading control. C. Quantification of the
effects of single or double nnt-a and nnt-b MO injection at increasing concentrations; cardiac edema
formation is used as a phenotypic readout for larval batches embryos scored at 3dpf. Comparisons of
equivalent doses for each of nnt-a or nnt-b MOs alone are not significant (3ng, p=0.13; 6ng, p=0.17, 9ng,
p=0.52). D. Quantification of control gene MO injection; cardiac edema formation is used as a
phenotypic readout for larval batches scored at 5dpf. Negative control genes (harboring variants shared
between the affected individuals in family C but not segregating with disease) show no significant
phenotype; notch1a, implicated previously in cardiac valve development shows marked cardiac edema.
n=50-100 embryos/injection batch, repeated twice with masked scoring.


Supplementary Movies
Supplementary Movie 1. Fluorescent video imaging of ventrally positioned 2 dpf control cmlc2:GFP
larva. Imaging was acquired at 8.77 frames/second at 12x magnification; anterior structures, left;
atrium, top chamber; ventricle, bottom chamber.
Supplementary Movie 2. Fluorescent video imaging of ventrally positioned 2 dpf nnt-a/nnt-b morphant
cmlc2:GFP larva. Imaging was acquired at 8.77 frames/second at 12x magnification; anterior structures,
left; atrium, top chamber; ventricle, bottom chamber.
References 1. Yeo S-Y, Kim M, Kim H-S, Huh T-L, Chitnis AB. Fluorescent protein expression driven by her4
regulatory elements reveals the spatiotemporal pattern of Notch signaling in the nervous system of zebrafish embryos. Dev Biol. 2007;301:555–567.