Lecture 8 Transcription Initiation Prokaryotic Eukaryotic Reading: Chapter 4 (115-116) Chapter 11...
-
Upload
ivan-willetts -
Category
Documents
-
view
218 -
download
3
Transcript of Lecture 8 Transcription Initiation Prokaryotic Eukaryotic Reading: Chapter 4 (115-116) Chapter 11...

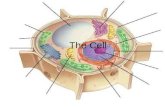
Lecture 8
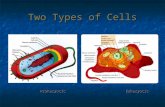
Transcription InitiationProkaryotic
Eukaryotic
Reading:
Chapter 4 (115-116)
Chapter 11
Molecular Biology syllabus web site

Bacterial transcription initiation
• RNA polymerase initiates transcription of most genes at a unique DNA position lying upstream of the coding sequence
• The base pair where transcription initiates is termed the transcription-initiation site or start site
• By convention, the transcription-initiation site in the DNA sequence is designated +1, and base pairs extending in the direction of transcription (downstream) are assigned positive numbers which those extending in the opposite direction (upstream) are assigned negative numbers
• Various proteins (RNA polymerase, activators, repressors) interact with DNA at or near the promoter to regulate transcription initiation

Differences in E. coli promoter sequences affect the frequency of transcription initiation

Lac Operon
• Beta Galactosidase (lacZ)
• Permease (lacY)
• Transacetylase (lacA)
lacZ lacY lacA
Lac promoter
mRNA
protein

LacZ (-galactosidase)




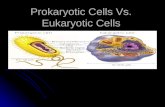
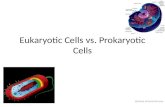
Eukaryotes

Promoter deletions to identify transcriptional control sequences




Promoter deletions to identify transcriptional control sequences

Run-off transcription: to map transcription initiation sites
METHOD
1. Isolate nuclei from specific tissues
2. Incubate: nuclear extract, restriction fragments containing start site, 32P-UTP
3. Synthesized RNA is separated by electrophoresis

Common Eukaryotic Promoter Consensus: TATA box
Alternate promoters:initiators- degenerate consensusGC-rich region (giving rise to multiple start sites)

Run-on transcription assay rate of transcription initiation
METHOD
1. Isolate nuclei from specific tissues
2. Label growing RNA chains with 32P-UTP
3. Hybridize to filters containing test cDNAs

S1 nuclease
mRNA
32P-labelled DNA probe
mRNA
32P-labelled DNA probe
denature
gel electrophoresis
S1 protection to assay transcription initiation rates

Enhancer elements
cis-acting sequences that serve to increase rate of transcription initiation regardless of their location near the gene (first discovered in SV40 viral genome).
S1 Protection Assay: to test rate of transcription initiation
METHOD
1. Transfect construct with/without enhancer into test cells
2. Isolate RNA and hybridize with 32P-labeled DNA probe
3. Treat with S1 nuclease, denature, subject to gel electrophoresis

Typical eukaryotic gene

Column purified fractions of DNA-binding proteins incubated with test promoter fragment
Gel-shift assays identify general region of protein-DNA interaction
METHOD
1. Isolate nuclei and prepare protein extract or purify DNA binding proteins
2. Incubate with restriction fragment or short DNA
3. Separate by agarose gel electrophoresis
Controls: non-specific DNA; no extract

METHOD
1. Isolate restriction fragment
2. Add DNA-binding protein
3. Partially digest with DNaseI
4. Separate by polyacrylamide gel electrophoresis
Controls: no protein
The footprint of RNA polymerase and lac repressor on the lac control region
DNase I footprinting assays identify specific regions of protein-DNA interactions

The lac control region contains three critical cis-acting sites
Figure 10-9

How to demonstrate the specificity of a putative transcription factor?
• In vitro
• In vivo

• In vitroMETHOD
1. Set up in vitro transcription reaction with DNA template, 32P-UTP, plus or minus transcription factor
2. Separate transcripts by gel electrophoresis
Controls: non-specific DNA; no protein

• In vivo
METHOD
1. Prepare two constructs:
a) encoding transcription factor X
b) reporter gene downstream of transcription factor binding site plus minimal promoter
2. Transfect cells and assay for reporter gene activity
Controls: transfect each plasmid alone

Transcription factors have separate domains
•DNA binding
•Transcription activation

Yeast Two-hybrid System- to fish out interacting transcription factors