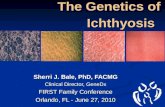
Lamellar Ichthyosis
description
Transcript of Lamellar Ichthyosis




• Autosomal recessive (ARCI-1)• Panethnic, rare (1 in 200,000)• Apparent at birth and is present throughout life• Genetically heterogenous (6 genes, 5 of them identified:
TGM1 (14q11.2), ABCA12 (2q34), 19p12-q12, 19p13; ALOXE3-ALOX12B (17p13), ichthyin (5q33)
• More than 90% mutant TGM1 gene (14q11.2)

• Infant is born encased in a hyperkeratotic translucent membrane, known as “collodion baby”.
• Crumpled ears, alopecia, eclabium, and ectropion.

The membrane sheds within 10-14 days and large, thick, brownish lamellar scales develop.

The child after 2 months showing lamellar ichthyosis(Aradhya et al. Clinical outcome of collodion baby: A retrospective review. Indian J Dermatol Venereol Leprol 2013;79:553)

• TGM1 consists of 15 exons • Encodes a 90 kDa, 817-amino-acid protein called transglutaminase-1
(TGase-1)

• Crosslinks proteins by catalyzing Ca2+ dependent formation of isopeptide b/w Glu from one protein and Lys in another.
• TGase-1 is responsible for cross-linking of ARP preserver proteins such as involucrin and loricrin during the formation of the cornified cell envelope (CCE)

• Essential in formation of the cornified cell envelope (CCE).
• The CCE, an insoluble structure composed of cross-linked proteins laid down on the inside of the plasma membrane in keratinocytes in the stratum corneum.
• The CCE acts as a mechanical barrier and protects against water loss and infectious agents.


Most Reported Mutations are located in exons: 2,3,4,5,6,7,8,13, 9,10,11,15
Working Flow Diagram

A>C transversion in exon 11(Reverse T>G)
Part of exon 11 sequence: Mutant vs. Normal


• NM_000359.2: c.1621A>C• Altered codon 541 from ACC into CCC thus changing the amino
acid threonine into proline • P22735: p.T541P

• The detected TGM1 exon 11 missense mutation. (a) the normal sequence (b) the homozygous c.A1621C mutantion (c) one of the heterozygous parents . The arrow indicates the mutation site.

• HGMD• Ensembl variation table• Literature• Novel

SIFT Results (Batch Protein)
Protein ID Substitution dbSNP ID Prediction Score Median Info Number of Seqs at position Comments
CCDS9622.1 T541P novel DAMAGING 0.00 2.77 45
* Low confidence means that the protein alignment does not have enough sequence diversity. Because the position artifically appears to be conserved, an amino acid may incorrectly predicted to be damaging.
(http://sift.jcvi.org/www/SIFT_BLink_submit.html)

• (http://genetics.bwh.harvard.edu/pph)
PolyPhen-2 report for P22735 T541PQueryProtein Acc Position AA1 AA2 Description
P22735 541 T P Canonical; RecName: Full=Protein-glutamine gamma-glutamyltransferase K; EC=2.3.2.13; AltName: Full=Epidermal TGase; AltName: Full=Transglutaminase K; Short=TG(K); Short=TGK; Short=TGase K; AltName: Full=Transglutaminase-1; Short=TGase-1; Length: 817
ResultsPrediction/Confidence PolyPhen-2 v2.2.2r398
HumDivThis mutation is predicted to be PROBABLY DAMAGING with a score of 0.998 (sensitivity: 0.27; specificity: 0.99)


• TGase-1 partial amino acid sequence (residues 524 to 574) alignments from a motif of the catalytic core domain. The amino acid sequence alignment by using the NCBI protein Blast analysis (http://blast.ncbi.nlm.nih.gov/Blast.cgi).

Several lines of evidence suggest the this mutation is pathogenic: (1)the mutation is inherited from the consanguineous parents and absent in
healthy control subjects (2)amino acid sequence alignment showed that the mutated threonine
residue is highly conserved across distant species i.e., it is important for the enzyme activity
(3)the altered amino acid is located in a critical region of the enzyme (the catalytic core domain) and in silico analysis using SIFT and PolyPhen-2 programs predicted that the amino acid change is damaging for the enzyme function
(4) threonine and proline have quite different characteristics: threonine fits into regular secondary structures proline behaves as a structural disruptor

• PCR-RFLP (HphI) results of the patient (lane 1), his mother and father (lanes 2 and 3, resepcetively), one homozygous normal control subject (lane 4) and a negative (no genomic DNA) control (lane 5). 238 bp, 124/114 bp.

• ARCI-1 (LI) in the present case is attributed to a novel “p.T541P” mutation in the catalytic core domain of the TGase-1 enzyme.
• The study further confirms the involvement of TGM1 mutations in LI. • This finding expands the mutations spectrum of TGM-1 gene and is
important for delivering proper counseling and diagnosis opportunities for the proband's family.
- NCBI/ClinVar Accession: SCV000148368