Journal of Proteome Research 2010 Breitkopf
Transcript of Journal of Proteome Research 2010 Breitkopf
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
1/39
Subscriber access provided by KU LEUVEN - BIOMEDICAL LIB
Journal of Proteome Research is published by the American Chemical Society. 1155Sixteenth Street N.W., Washington, DC 20036Published by American Chemical Society. Copyright American Chemical Society.
However, no copyright claim is made to original U.S. Government works, or worksproduced by employees of any Commonwealth realm Crown government in the courseof their duties.
Article
Proteomics Analysis of Cellular Imatinib Targets
and their Candidate Downstream EffectorsSusanne B Breitkopf, Felix S Oppermann, Gyrgy Kri, Markus Grammel, and Henrik Daub
J. Proteome Res., Just Accepted Manuscript DOI: 10.1021/pr1008527 Publication Date (Web): 27 September 2010
Downloaded from http://pubs.acs.org on October 8, 2010
Just Accepted
Just Accepted manuscripts have been peer-reviewed and accepted for publication. They are postedonline prior to technical editing, formatting for publication and author proofing. The American Chemical
Society provides Just Accepted as a free service to the research community to expedite thedissemination of scientific material as soon as possible after acceptance. Just Accepted manuscriptsappear in full in PDF format accompanied by an HTML abstract. Just Accepted manuscripts have been
fully peer reviewed, but should not be considered the official version of record. They are accessible to allreaders and citable by the Digital Object Identifier (DOI). Just Accepted is an optional service offered
to authors. Therefore, the Just Accepted Web site may not include all articles that will be publishedin the journal. After a manuscript is technically edited and formatted, it will be removed from the Just
Accepted Web site and published as an ASAP article. Note that technical editing may introduce minorchanges to the manuscript text and/or graphics which could affect content, and all legal disclaimers
and ethical guidelines that apply to the journal pertain. ACS cannot be held responsible for errorsor consequences arising from the use of information contained in these Just Accepted manuscripts.
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
2/39
1
Proteomics Analysis of Cellular Imatinib Targets and their Candidate
Downstream Effectors
Susanne B. Breitkopf1,,*, Felix S. Oppermann1,,*, Gyrgy Kri2,3,
Markus Grammel1, & Henrik Daub1,4,#
1Department of Molecular Biology, Max Planck Institute of Biochemistry, Am Klopferspitz 18,
82152 Martinsried, Germany. 2Vichem Chemie Ltd., Herman Ott u. 15., Budapest, 1022,
Hungary. 3Pathobiochemistry Research Group of the Hungarian Academy of Science,
Semmelweis University, Puskin u. 9., Budapest, 1088, Hungary. 4Kinaxo Biotechnologies
GmbH, Am Klopferspitz 19, 82152 Martinsried, Germany.
Present address: Beth Israel Deaconess Medical Center, Harvard Medical School, 3 Blackfan
Circle, Boston, MA 02115
Present address: Kinaxo Biotechnologies GmbH, Am Klopferspitz 19, 82152 Martinsried,
Germany.
Present address: Laboratory of Chemical Biology and Microbial Pathogenesis, The Rockefeller
University, 1230 York Avenue, New York, NY 10065
*These author contributed equally to this work.
#Correspondence: [email protected]
Running title: Quantitative kinase drug proteomics
Keywords: Kinase, inhibitor, imatinib, CML, SILAC, affinity purification, target
ge 1 of 38
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
3/39
2
Summary
Inhibition of de-regulated protein kinases by small molecule drugs has evolved into a major
therapeutic strategy for the treatment of human malignancies. Knowledge about direct cellular
targets of kinase-selective drugs and the identification of druggable downstream mediators of
oncogenic signaling are relevant for both initial therapy selection and the nomination of
alternative targets in case molecular resistance emerges. To address these issues, we
performed a proof-of-concept proteomics study designed to monitor drug effects on the
pharmacologically tractable subproteome isolated by affinity purification with immobilized, non-
selective kinase inhibitors. We applied this strategy to chronic myeloid leukemia cells that
express the transforming Bcr-Abl fusion kinase. We used SILAC to measure how cellular
treatment with the Bcr-Abl inhibitor imatinib affects protein binding to a generic kinase inhibitor
resin and further quantified site-specific phosphorylations on resin-retained proteins. Our
integrated approach indicated additional imatinib target candidates, such as flavine adenine
dinucleotide synthetase, as well as repressed phosphorylation events on downstream effectors
not yet implicated in imatinib-regulated signaling. These included activity-regulating
phosphorylations on the kinases Btk, Fer and focal adhesion kinase which may qualify them as
alternative target candidates in Bcr-Abl-driven oncogenesis. Our approach is rather generic and
may have various applications in kinase drug discovery.
Page 2
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
4/39
3
Introduction
Protein kinases are critical regulators in human cancer and play major roles in tumour cell
proliferation, migration and survival.1 Aberrant kinase activity has been identified as a major
factor contributing to disease progression in various human malignancies.2 The targeted
inhibition of protein kinases has therefore emerged as a major therapeutic approach and fuelled
the development of various kinase-selective drugs, such as cell-permeable small molecule
inhibitors, with the potential to address currently unmet medical needs in cancer therapy.3, 4
Imatinib (also known as imatinib mesylate, Gleevec, Glivec or STI571) was one of the first small
molecule inhibitors developed for the targeted inactivation of kinases in human cancer. Imatinib
efficiently blocks the activities of several tyrosine kinases including Abl, Kit and the platelet-
derived growth factor receptor and has demonstrated impressing clinical efficacy in human
malignancies such as chronic myeloid leukemia (CML).5 In most cases CML pathogenesis
results from the Philadelphia (Ph) chromosome translocation which generates the causative
BCR-ABL oncogene.6 By selective interference with the de-regulated Bcr-Abl kinase activity
imatinib treatment results in impressive and long-lasting responses in chronic-phase CML
patients.5 However, CML patients in advanced disease states such as the accelerated and blast
crisis phases typically relapse and acquire resistance to imatinib within several months.7 In the
majority of these cases, resistance formation is due to mutations in the kinase domain-encoding
region of the BCR-ABL oncogene, which selectively interfere with imatinib binding without
abrogating the catalytic activity of Abl tyrosine kinase.7-9 Molecular resistance of the targeted
Bcr-Abl oncoprotein in relapsed CML patients demonstrates its continued requirement for
disease progression. Structural data revealed that imatinib selectively interacts with an inactive
conformation of the Abl kinase, which is destabilized by many imatinib resistance-conferring
mutations.10 These mechanistic insights provided a rational basis for the development of
second-generation inhibitors, such as the small molecule drugs bosutinib and dasatinib, which
ge 3 of 38
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
5/39
4
target the active kinase conformation and thereby overcome imatinib resistance in many Abl
kinase variants.7-9 However, similar to imatinib, these second-generation drugs lack inhibitory
activity against the frequently occurring Thr-315 to Ile mutation in the Abl kinase domain, which
directly interferes with drug binding irrespective of the kinase conformation. Drug development
and clinical efforts are ongoing with the goal to address all possible drug-resistant Abl kinase
mutants.9 Alternative to targeted inhibition of the mutated, causative oncoprotein, therapeutic
intervention might also be directed against essential downstream mediators of Bcr-Abl. The Bcr-
Abl fusion protein possesses constitutive tyrosine kinase activity and assembles multi-protein
complexes that trigger proliferative and anti-apoptotic signaling as well as regulation of the actin
cytoskeleton.11, 12 Previous investigations have mostly been hypothesis-driven and placed
known signal transducing modules in Bcr-Abl signaling, such as the Ras/mitogen-activated
protein kinase (MAPK) cascades, phosphatidylinositol-3 kinase/Akt signaling and the signal
transducer and activator of transcription (STAT) pathway,7 and their concomitant activation is
implicated in the malignant transformation in Bcr-Abl-expressing leukemia cells. In addition to
these well documented pathways, Bcr-Abl might engage additional signal transducers that are
regulated by reversible phosphorylation events and have not been revealed by previous studies.
Recent developments in proteomics, including the availability of rapid, sensitive and highly
accurate hybrid ion trap-orbitrap mass spectrometers, improved phosphopeptide fractionation
procedures and breakthroughs in MS data processing and quantification, make MS-based
phosphoproteomics the method-of-choice for unbiased signal transduction analyses.13-20
Quantitative phosphorylation analyses enabled by stable isotope labeling by amino acids in cell
culture (SILAC) has been used to identify imatinib-induced tyrosine phosphorylation changes in
K562 cells21 and, more recently, for a global survey of phosphoproteome regulation upon
dasatinib treatment of the same cell line.22 The identification of downstream protein kinases with
essential roles in Bcr-Abl signal transmission would be of particular interest, as such knowledge
Page 4
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
6/39
5
might define alternative, small molecule-tractable targets in case of relapse due to drug-
insensitive Abl kinase mutants. In addition to phosphorylating their cellular substrates, protein
kinases regulate each other by reversible phosphorylation events and, moreover, many protein
kinases undergo autophosphorylation upon cellular activation.23 Thus, comprehensive
monitoring of imatinib-induced phosphorylation changes on protein kinases might allow for the
identification of druggable downstream targets in Bcr-Abl signaling, and thus contribute to the
nomination of new candidates for therapeutic intervention.
To analyze small molecule-tractable protein kinases with high analytical sensitivity we and
others have previously developed affinity chromatography procedures that employ combinations
of immobilized kinase inhibitors for selective kinase pre-fractionation from total cell extracts.24-26
This approach combined with SILAC enabled us to quantify the cell cycle regulation of more
than 200 protein kinases and to detect more than 1,000 distinct phosphorylation events on
these key signaling enzymes.24 A similar kinase enrichment strategy reported by Bantscheff et
al. was used to identify cellular targets of the clinical Bcr-Abl kinase inhibitors imatinib, dasatinib
and bosutinib as well as downstream signaling elements upon treatment of K562 cells with
these drugs.26 To further expand the knowledge obtained in this previous work, in particular to
identify additional kinase candidates downstream of direct imatinib targets, we revisited the
imatinib paradigm in our present study. Using imatinib-treated K562 CML cells as a model
system, we here present proof-of-concept for an integrated proteomics strategy that
quantitatively assesses both direct drug targets and their downstream signal transducers, and
report previously unknown target protein candidates falling in both categories.
ge 5 of 38
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
7/39
6
Experimental Section
Cell Culture. For all SILAC experiments, the human CML cell line K562 (ATCC, # CCL-243)
was cultured in suspension in RPMI1640 medium (Invitrogen) containing 10% dialyzed fetal
bovine serum (Invitrogen), 1% (10,000 units/ml) penicillin/(10 mg/ml) streptomycin (Invitrogen)
and either 45 mg/l unlabeled L-arginine and 76 mg/l unlabeled L-lysine (Arg 0, Lys0) or equimolar
amounts of L-[U-13C6]arginine and L-[2H4]lysine (Arg
6, Lys4) or L-[U-13C6,15N4] and L-[U-
13C6,15N2]lysine (Arg
10, Lys8) (Cambridge Isotope Laboratories or Sigma). K562 cells were grown
in SILAC medium for five cell doublings on 15 cm dishes and then transferred to spinner flasks
into 500 ml fresh SILAC medium at a cell density of 0.25 x 106 cells/ml. After two further rounds
of cell division, K562 cells were either treated with 1 M or 10 M imatinib mesylate (ACC
Corporation) or control-incubated with DMSO for 90 min before harvesting by centrifugation.
Our cell culture strategy yielded total cell numbers of 5 x 108 per labeling condition. Harvested
K562 cells were washed once with ice-cold PBS, snap-frozen in liquid nitrogen and stored at -
80C until cell lysis.
Cell Lysis and Kinase Enrichment. Kinase inhibitor resins containing the immobilizedcompounds VI16832, purvalanol B, bisindolylmaleimide X, AX14596 and SU6668 were
essentially prepared as described previously,24, 25, 27 with the only differences that 2 volumes of
1.5 mM (instead of previously 0.75 mM)24 VI16832 solution and 2 volumes of 5 mM (instead of
10 mM)25 bisindolylmaleimide X were coupled to 1 volume of aspirated epoxy-activated
Sepharose 6B for immobilization. For each of the two replicate experiments, we prepared a
mixed kinase inhibitor resin containing 0,5 ml of the VI16832 and purvalanol B resins and 0,33ml of the bisindolylmaleimide X, AX14596 and SU6668 resins. Frozen cell pellets from
differentially encoded and treated K562 cell populations were solubilized with 9 ml of lysis buffer
containing 50 mM Hepes-NaOH, pH 7.5, 150 mM NaCl, 0.5% Triton X-100, 1 mM EDTA, 1 mM
Page 6
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
8/39
7
EGTA, 1mM phenylmethylsulfonyl fluoride, 10 mM NaF, 2.5 mM Na3VO4, 50 ng/ml calyculin A
(Alexis Biochemicals, San Diego, CA), 10 g/ml aprotinin, 10 g/ml leupeptin, and 1%
phosphatase inhibitor mixtures 1 and 2 (Sigma) for 1 h at 4C. Cell debris was removed by
centrifugation (20 min at 13,000 rpm) and by further filtering through 0.22-m mixed esters of
cellulose membranes (Millipore). Protein concentration was measured using the BCA assay
(Pierce). 55 mg of each of the three differentially labeled K562 lysates were adjusted to a final
NaCl concentration of 1 M and a final volume of 10 ml. SILAC-encoded samples were then
subjected to parallel in vitroassociations with 0.7 ml mixed kinase inhibitor resin for 2 h at 4C in
the dark. Beads were then washed three times with 10 ml lysis buffer adjusted to 1 M NaCl and
twice with lysis buffer containing 150 mM NaCl. For elution of bound proteins, mixed kinase
inhibitor beads were repeatedly incubated for 10 min with 1.4 ml elution buffer (20 mM Tris-HCl
pH 7.5, 5 mM DTT, 0.5% SDS) at 50C. Aliquots of the resulting elution fractions were analyzed
by SDS-PAGE and silver staining. Protein-containing elution fractions were pooled and
lyophilized, and then resuspended with water in one tenths of the initial volume prior to protein
precipitation according to the protocol by Wessel & Flgge.28
Sample Preparation for Mass Spectrometry. In each of the two replicate analyses, 25% of
the kinase-enriched fraction was solubilized in 20 mM HEPES buffer (pH 7.5) containing 7 M
urea, 2 M thiourea, 1% n-octylglucoside and then reduced, alkylated and sequentially digested
with the endoprotease Lys-C (Wako) and modified trypsin (sequencing grade, Promega) as
described previously.13 The resulting peptide samples were then separated by strong cation
exchange chromatography on an KTA explorer system into a flow-through and 6 elution
fractions using a 1 ml Resource S column (GE Healthcare) according to a published
protocol.13
ge 7 of 38
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
9/39
8
The remaining protein pellets were dissolved in 1.5x LDS-Buffer and separated on a 10%
NuPage Bis-Tris gel (Invitrogen) according to the manufacturers instructions. Proteins were
stained using the Collodial Blue staining kit (Invitrogen). In both SILAC experiments, the gel was
cut into 16 slices followed by in-gel digestion with trypsin.29 20% of the resulting peptide
mixtures were mixed with an equal volume of 1% TFA, 5% ACN and then loaded on C 18
StageTips.30 After washing twice with buffer containing 0.5% acetic acid and 0.1% TFA bound
peptides were eluted with buffer containing 0.5% acetic acid and 80% ACN and then
concentrated in a Speed-Vac prior to further analysis.
The larger part of 80% of each fraction from the tryptic in-gel digests were subjected to
phosphopeptide enrichment using titanium dioxide (TiO2) microspheres.31, 32 The TiO2 beads
(GL Science, Tokyo, Japan) were first equilibrated by consecutive incubations with 20 mM
NH4OH in20% acetonitrile (ACN), pH 10.5, washing buffer (0.1%TFA, 50% ACN) and loading
buffer (5 g/liter 2,5-dihydrobenzoicacid in 55% ACN). Trypsin digests from adjacent gel slides
were combined to a total of 8 peptide samples for further phosphopeptide enrichment. Each of
them was adjusted to a final concentration of 30% ACN, 2 M urea and incubated with 5 mg
equilibrated TiO2 beads for 30 min at room temperature on a rotating wheel. Afterwards, beads
were washed once with 100 l of loading buffer, three times with 1.5 ml of washing buffer, and
phosphopeptides wereeluted by incubating twice with 30 l of 20 mM NH4OH in20% ACN, pH
10.5. Elution fractions were combined and passed through a C8 StageTip followed by a 30-l
rinse with 80% ACN, 0.5%acetic acid. After adjusting to a pH of 6, samples were concentrated
to 3 l and mixed with an equal volume of 4% ACN, 0.2% TFA. We further performed
phosphopeptide purifications with TiO2 microspheres from the seven SCX chromatography
fractions of in-solution digested, kinase-enriched samples in each of the two replicate
experiments. Additionally, total peptide extractions with C18 StageTips were done with 20%
aliquots of the SCX fractions in experiment 2.
Page 8
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
10/39
9
Mass Spectrometry Analysis. MS analyses were done as described previously13, 24. Briefly,
peptide separations were done on 15-cm analytical columns (75-m inner diameter) in-house
packed with 3-m C18 beads (Reprosil-AQ Pur, Dr. Maisch) using a nanoflow high pressure
liquid chromatography system (Agilent Technologies 1100). Peptides were eluted with 2 h
gradients from 5% to 40% ACN in 0.5% acetic acid and directly electrosprayed into a LTQ-
Orbitrap mass spectrometer (Thermo Fisher Scientific) by a nanoelectrospray ion source
(Proxeon Biosystems). The LTQ-Orbitrap was operated in the data-dependent mode to
automatically switch between full scan MS in the orbitrapanalyzer (with resolution r= 60,000 at
m/z400) and the fragmentationof the five most intense multiply-charged peptide ions by either
MS/MS or multi-stageactivation in the LTQ part of the instrument, the latter being triggered upon
neutral losses of 97.97, 48.99, or 32.66 m/z.33 For all full scan measurements in the orbitrap
detector a lock-mass strategy was used for internal calibration as described.34Typical mass
spectrometric conditions were: spray voltage, 2.4 kV; no sheath and auxiliary gas flow; heated
capillary temperature, 150C; normalized collision energy 35% for MSA in LTQ. The ion
selection threshold was 500 counts for MS2. An activation q = 0.25 and activation time of 30 ms
were used.
Peptide Identification, Quantification, and Data Analysis. All MS raw files from both
biological replicate analyses were collectively processed with the MaxQuant software suite
(version 1.0.13.12), which performs peak list generation, SILAC-based quantification, estimation
of false discovery rates, peptide to protein group assembly, and data filtration and presentation
as described.20 Data were searched against a concatenated forward and reversed version of the
human International Protein Index (IPI) database version 3.37 containing 69141 protein entries
and 175 frequently detected contaminants (such as porcine trypsin, human keratins and Lys-C)
using the Mascot search engine (Matrix Science; version 2.2.04). Cysteine
carbamidomethylation was set as a fixed modification and methionine oxidation, protein N-
ge 9 of 38
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
11/39
10
acetylation, loss of ammonia from N-terminal glutamine as well as phosphorylation of serine,
threonine and tyrosine residues were allowed as variable modifications. Spectra resulting from
isotopically labeled peptides, as revealed by presearch MaxQuant analysis of SILAC partners,
were searched with the fixed modifications Arg6 and Lys4 or Arg10 and Lys8, respectively,
whereas spectra for which a SILAC state could not be assigned before database searching
were searched with Arg6, Arg10, Lys4 and Lys8 as variable modifications. The accepted mass
tolerance was set to 7 p.p.m for precursor ions and to 0.5 Da for fragment ions. The minimum
required peptide length was 6 amino acids and up to three missed cleavage sites and three
isotopically labeled amino acids were permitted. The accepted FDR was 1% for both protein
and peptide identifications, and the cut-off for the posterior error probability (PEP) of peptides
was set to 10%. Phosphorylation site assignments were performed by a modified version of the
PTM scoring algorithm13 implemented in MaxQuant. Phosphorylation site assignments were
classified as class I sites in case of a localization probability of at least 0.75 and a score
difference of at least 5 to the second most likely assignment.
Network Analysis. All IPI identifiers of all quantified protein groups were matched to respective
Ensembl entries using BioMart and then uploaded to the Search Tool for the Retrieval of
Interacting Genes/Proteins (STRING) database (version 8.2).35 We retrieved interactions that
were of at least high confidence (score 0.7) based exclusively on experimental and database
knowledge while excluding all other prediction methods implemented in STRING (such as
textmining and coexpression). The resulting networks were visualized using Cytoscape36.
Additionally, we randomly selected subsets of the IPI database that contained the same number
of entries as present in our experimental data. This was repeated five times to determine the
average numbers of network nodes and edges in random protein selections by STRING
analysis.
Page 10
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
12/39
11
Data Availability.All raw data files from this study have been up-loaded to the Tranche file-
sharing system (ProteomeCommons.org, hash: 1jgzdG97a2b6CKjEqKrhQr2xSu6uyff7dft5qy
nd/cBqaOTO37Re2Ys7GHbDiOX5J+J8FQjtVQZ799wYWADqviLesoIAAAAAAABHsA==).
Furthermore, annotated phosphopeptide spectra for all identified class I sites have been
deposited (hash: G3KHiKwaE7x1r164vN4p1uMmbNrRDbVozsgmlmLi1qsAnXFPV2mcHTeyJ
JpQIqNtygeQbnZe3WtmmrsknUeyVmXkMh8AAAAAAAfdqQ==).
ge 11 of 38
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
13/39
12
Results and Discussion
Experimental Strategy. As judged by initial immunoblot analysis of total K562 cell lysates,
maximal repression of cellular tyrosine phosphorylation was evident after 45 min treatment with
10 M imatinib (data not shown). We further reasoned that serine/threonine phosphorylation
events located downstream in imatinib-regulated signaling might exhibit slower
dephosphorylation kinetics than direct tyrosine kinase substrates and therefore opted for 90 min
treatment in subsequent SILAC experiments. We also decided against longer stimulation times
of several hours used in earlier studies12, 26, as such treatment schemes might bear an
increased risk of accumulating secondary changes far away from the initial sites of imatinib
action. To enable sensitive and unbiased detection of imatinib effects on the kinase inhibitor-
tractable sub-proteome, we implemented SILAC for K562 leukemia cells grown in spinner
flasks. We differently encoded three populations of K562 cells by culturing them in medium
containing either normal arginine and lysine (Arg0 and Lys0) or combinations of heavier isotopic
variants of the two amino acids (Arg6 and Lys4, or Arg10 and Lys8) (Figure 1). Differently labeled
cells were treated for 90 min with either 1 M or 10 M imatinib, or control-incubated with
solvent prior to cell lysis. We subjected each of the resulting lysates to a separate in vitro
association with a mixture of five kinase inhibitor resins. This affinity purification strategy was
designed for comprehensive enrichment of drug-interacting protein kinases along with their
associating factors, and we used our previously established incubation conditions to ensure
preservation of cellular protein phosphorylation states.24, 25 Bound proteins were eluted from the
resin mixtures, and we pooled the kinase-enriched fractions from differentially encoded and
treated K562 cells prior to further sample processing. Three fourth of the combined material was
resolved by gel electrophoresis and in-gel digested with trypsin, followed by StageTip
extractions of total peptide samples and phosphopeptide purification with TiO2 beads. The
remaining inhibitor resin-enriched material was digested with trypsin in-solution prior to SCX
Page 12
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
14/39
13
chromatography and TiO2 enrichment of phosphorylated peptides (Figure 1). Thus, our
combined pre-fractionation strategy exploited the different fractionation principles of gel-based
and gel-free approaches for more comprehensive phosphopeptide coverage than possible with
either approach alone.24 All peptide and phosphopeptide fractions were analyzed by nanoscale
liquid chromatography-tandem MS (LC-MS/MS) analysis on a linear ion trap/orbitrap (LTQ-
Orbitrap) hybrid mass spectrometer. Moreover, to assess the biological reproducibility of
quantitative MS data, we repeated the whole analysis in a replicate experiment with a modified
SILAC scheme for the different treatment conditions. All resulting raw data files were then
collectively processed with the MaxQuant software suite for the integrated analysis of both site-
specific phosphorylation changes and protein binding to the kinase inhibitor resin upon imatinib
treatment. Our goal was to demonstrate proof-of-concept for a proteomics approach designed to
identify both regulated effector proteins, which are downstream of imatinib-inhibited kinases and
therefore exhibit repressed site-specific phosphorylations without changes in protein
abundance, and possible direct cellular imatinib targets, which are prevented from resin
interactions by bound imatinib.
Analysis of the Kinase Inhibitor-enriched Sub-proteome. In total, we identified more than
2,000 inhibitor resin-retained proteins with an accepted false-discovery rate of less than 1%.
Protein ratios for imatinib versus control-treated K562 cells could be determined for 1,275
distinct proteins, of which 683 were quantified in both biological replicate experiments (Figure
2A, Supplementary Table 1). Due to the kinase enrichment strategy we obtained such
quantitative data for more than 170 members of the protein kinase superfamily, which indicate
substantial enrichment considering that the kinome accounts for only 1.7% of the human
genome. Our affinity purifications with a mixture of broadly selective, ATP competitive inhibitors
fractionated for other likely direct binders that did not belong to the protein kinase superfamily,
ge 13 of 38
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
15/39
14
for example other types of nucleotide-utilizing enzymes. We detected many proteins falling into
such categories, including various dehydrogenases and lipid kinases.
In addition to protein quantification based on unphosphorylated peptides, phosphopeptide
enrichment allowed for quantification of more than 13,500 identified phosphopeptides, which
harbored 1,842 distinct phosphorylation sites that could be localized to specific serine, threonine
or tyrosine residues with high confidence (class I sites with a localization probability 0.75)
(Supplementary Tables 2 and 3). For 898 class I phosphorylation site ratios were determined in
both biological replicate experiments, and, notably, with 504 more than half of all repeatedly
quantified sites were detected on protein kinases (Figure 2A). The overlap of phosphorylation
sites quantified in both experiments was higher for protein kinases compared to all other
proteins, which might be due to a certain subset of non-specific binders in the latter group prone
to higher inter-experimental variability. Both experiments combined, we quantified as many as
868 distinct phosphorylation events on K562 cell-derived protein kinases which account for
three times as many phosphorylation sites on protein kinases compared to an earlier study.26
Thus, our current study considerably expands previous knowledge on phospho-modifications in
the expressed K562 cell kinome.
While protein and phosphorylation site ratios were determined for only 23% of all quantified
proteins, this was possible for the majority (64%) of the 213 quantified protein kinases (Figure
2B). About 81% of all identified site-specific phosphorylations were located on serine residues,
whereas phosphorylated threonines and tyrosines accounted for about 13% and 6% of all
phosphorylation sites, respectively (Figure 2C).
Furthermore, cellular interaction partners of direct inhibitor targets were also expected to be
captured by subsequent MS analysis in case such interactions were preserved during cell lysis
and affinity purification. In the present study, we used 1 M NaCl-containing buffer to promote
Page 14
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
16/39
15
protein kinase selectivity in the enrichment step.37, 38 Although such high salt concentrations can
disrupt hydrophilic protein-protein binding, the majority of protein kinase interactions with other
proteins are apparently not suppressed according to our recent comparison of kinase
enrichment in low and high salt conditions.38 Therefore, we reasoned that the current datasets
can be used to get an impression of the overall relationships within the affinity purified sub-
proteome. We used STRING to retrieve known interactions among all proteins for which
quantitative data was available from both biological replicate experiments. Out of these 900
proteins submitted to STRING 489 reappeared within a complex network, in which we specified
all nodes depending on whether they were identified as phosphoproteins and/or represented
protein kinases (Supplementary Figure 1, Supplementary Table 4). Notably, we detected as
many as 3435edges within the STRING-derived network, which were almost 20-fold more than
obtained for the same number of randomly selected IPI database identifiers (Supplementary
Figure 1). This indicated a high degree of network connectivity within the enriched sub-
proteome, which was in part due to the identification of many known kinase interactors including
several cyclins, SH2-domain containing proteins and regulatory kinase subunits (Supplementary
Figure 1, Supplementary Table 4). Moreover, we identified prominent modules of proteins
involved in translation, RNA processing and proteasomal protein degradation. Detection of
these rather abundant proteins might result from unspecific binding or sedimentation in the
inhibitor affinity purification step instead of specific interactions with coupled inhibitors or bound
inhibitor targets. However, we found the corresponding gene ontology (GO) biological process
categories highly overrepresented in the proteins detected upon kinase inhibitor enrichment
compared to those identified in a parallel analysis of K562 total cell extracts (data not shown),
thus pointing to preferred detection of these protein machineries as a by-product of our chemical
proteomics strategy.
ge 15 of 38
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
17/39
16
Identification of Imatinib-interacting Proteins. Due to the SILAC strategy combined with
parallel inhibitor affinity purifications we could identify proteins that exhibited decreased resin
binding upon prior imatinib treatment of K562 cells. To obtain reliable data, we filtered all
SILAC-based quantifications for proteins that were recorded in both biological replicate
experiments. Moreover, we considered only those proteins as target candidates which were
reproducibly quantified with SILAC ratios below 0.5 for affinity resin-retained proteins from 10
M imatinib versuscontrol incubated cells. (Supplementary Table 1). Evidently, proteins with
such properties comprise direct cellular imatinib targets as well as their interaction partners, as
monitored for Bcr-Abl and its associated signal transducers Grb2 and SHIP2. Grb2 is known to
bind to Bcr-Abl in a phosphotyrosine-dependent manner via its SH2 domain, whereas SHIP2
was reported to bind to the SH3 domain of Abl.7, 11 In accordance with the reported high imatinib
affinity of Bcr-Abl, its binding to the multi-inhibitor resin was already fully suppressed upon
exposure of K562 cells to 1 M imatinib (Figure 3, Supplementary Figure 2A).5 We detected
similar resin binding properties for discoidin domain receptor 1 (DDR1), a receptor tyrosine
kinase recently identified as a high affinity target by Bantscheff and colleagues.26 We further
monitored imatinib-dependent competition for quinone reductase 2 (NQO2) and the tyrosine
kinase Syk (Figure 4A). These two enzymes have been previously characterized as additional
imatinib targets.26, 39, 40 While the known high affinity for NQO2 was reflected by its nearly
complete competition at both imatinib concentrations of 1 M and 10 M, Syk binding was
prevented in a dose-dependent manner, with less than 40% still retained at the higher imatinib
concentration as indicated by an average binding ratio of 0.36 (Figure 3, Supplementary Figure
2A).Our results regarding Syk were consistent with earlier biochemical data, which identified
Syk as a low-affinity target with a reported Ki value of 5 M for imatinib.39 Furthermore, we
identified two phosphatidylinositol-4-kinase type-2 isoforms and (PIP4K2A and PIP4K2C)as
potential new imatinib targets (Figure 3, Supplementary Figure 2B). Although it cannot be
Page 16
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
18/39
17
formally excluded that these lipid kinases interacted indirectly with immobilized inhibitors and
imatinib, direct binding appears more likely due to their structural and functional similarities to
protein kinases. Imatinib-dependent displacement of these lipid kinases was similar as observed
for Syk, and the moderate effect of imatinib argues against their substantial cellular inhibition at
therapeutically relevant drug concentrations (Figure 3). However, as even minor structural
differences can significantly alter drug affinities, our data might warrant further testing of
phosphatidylinositol-4-kinases against related drugs such as nilotinib, NNO-406 or development
compounds based on the phenylaminopyridine scaffold of imatinib.7 Additionally, we identified
the flavine adenine dinucleotide (FAD) synthetase encoded by the FLAD1 gene as a new
potential target of imatinib. FAD synthetase is a key metabolic enzyme which catalyzes the
formation of FAD by adenylation of flavin mononucleotide (Figure 3, Supplementary Figure
2B).41 FAD represents a redox co-factor of many flavoproteins and is therefore essential for
many biological processes, suggesting that pharmacological inactivation of FAD synthetase
might cause toxicity. Notably, FAD is present as a prosthetic group in the described imatinib
target NQO2 and functions in the electron transfers catalyzed by this oxidoreductase. Thus, our
identification of FAD synthetase might point to common structural features that determine
imatinib binding to some FAD-utilizing or -containing enzymes. Alternatively, imatinib might
selectively interact with the ATP site of FAD synthetase. Binding ratios of FAD synthetase were
0.55 and 0.21 for 1 and 10 M imatinib-treated versus control incubated cells, respectively,
indicating that almost 80% of the enzyme was not retained by the affinity beads at the higher
imatinib dose. Thus, imatinib interfered with resin binding of FAD synthetase to a lesser extent
than observed for Bcr-Abl and NQO2, but had a more pronounced effect on this enzyme than
on Syk and PI4 kinases (Figure 3).
Identification of Imatinib-regulated Downstream Kinases. Our enrichment strategy enabled
the sensitive detection and quantification of protein kinases-derived phosphopeptides. To
ge 17 of 38
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
19/39
18
identify biologically reproducible effects induced by imatinib, we filtered our data for
phosphorylation sites that could be quantified and confidently assigned to specific residues in
both replicate experiments (Supplementary Table 1). As shown in Figure 4A, most quantified
phosphorylation sites were not affected by imatinib and were found in ratios close to one in both
experiments. Our goal was to obtain proof-of-concept that the identification of potentially drug-
regulated sites is feasible by our approach. We considered imatinib-induced changes as
relevant for further inspection in case phosphorylation sites were either consistently down- or
up-regulated by more than two-fold in both experiments, or exhibited an average regulation of at
least two-fold with both ratios differing by a factor of less than two. However, these data have to
be seen as preliminary in a sense that additional biological replicates would be needed to
enable the statistical evaluation of individual site ratios. Biologically reproducible down-
regulation according to the aforementioned criteria was observed for 70 distinct phosphorylation
sites upon cellular incubation with 10 M imatinib (Supplementary Table 3). Notably, regulation
on tyrosine residues was far more prominent than their prevalence among all phosphosites
quantified in kinase-enriched K562 cell fractions. We detected as many as 15 distinct tyrosine
phosphorylated residues and a similar number of Ser/Thr sites mapping to the Bcr-Abl
oncoprotein. These sites were found at very low ratios upon cellular imatinib treatment which
reflects the combined effect of cellular dephosphorylation and near complete prevention of Bcr-
Abl protein binding due to imatinib binding. However, these very low SILAC ratios obviate
reliable quantification of phosphorylation versus protein changes, and we therefore did not
further consider Bcr-Abl phosphorylation sites in our quantitative analysis. In case of all other
regulated phosphoproteins for which protein ratios were measured normalization of
phosphorylation changes was possible due to either less dramatic (as observed for the direct
target Syk) or no imatinib effect on the amount of resin-bound protein. Notably, most imatinib-
Page 18
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
20/39
19
regulated phosphorylations have not been reported in earlier studies (Supplementary Table 3),
including all down-regulated sites listed in Table 1 and discussed in the following part.
Syk phosphorylation site ratios were on average threefold more strongly reduced upon 10 M
imatinib compared to Syk protein binding, our results indicated cellular dephosphorylation at
residues such Tyr-323, Tyr-348 and Tyr-352. Previous reports revealed that Syk
phosphorylation on these sites creates binding sites for signaling proteins such as
phospholipase C, the guanine nucleotide exchange factor Vav, phosphatidylinositol 3-kinase
and c-Cbl42-45. Interestingly, while Tyr-348 and Tyr-352 (or the corresponding residues in mouse
Syk) were shown to positively contribute to cellular Syk function, Tyr-317 was characterized as
a negative regulatory site in Ag and Fc receptor signaling45-47. By extension, our results point to
possible functional modulation of Syk-mediated signaling at the higher imatinib dose in CML
cells.
The major goal of our phosphoproteomics analysis was the identification of protein kinases that
transduce signals emanating from direct imatinib interactors, as a strategy to identify potential
alternative targets in case of imatinib resistance formation due to Bcr-Abl mutations oroverexpression. In contrast to direct cellular imatinib targets, imatinib treatment would not affect
inhibitor resin binding of such downstream signaling kinases, but instead selectively repress
site-specific phosphorylations in our experimental approach. To identify such imatinib-induced
effects, we focused on phosphorylation sites that could be quantified and confidently assigned
to specific residues in both replicate experiments. Notably, we recorded effects on a number of
phosphorylation sites with reported regulatory functions. According to our analysis, imatinibtreatment exerted a dose-dependent effect on the phosphorylation of the cytoplasmic tyrosine
kinase BTK at Tyr-551, with a threefold down-regulation measured for the 10 M imatinib
concentration (Table 1). Quantification of non-phosphorylated peptides from BTK revealed that
ge 19 of 38
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
21/39
20
comparable protein amounts were retained by the inhibitor resin from imatinib-treated cells. We
observed similar regulation patterns for Tyr-714 of the cytoplasmic tyrosine kinase Fer and Tyr-
774 of the receptor tyrosine kinase EphB4 (Fig. 4B, Table 1). Our evidence of selective
dephosphorylation in the absence of protein changes points to these three tyrosine kinases as
potential downstream signaling elements of direct imatinib targets. Notably, these imatinib-
repressed phosphorylations occurred in the activation loop regions of the BTK, Fer and EphB4,
at a conserved position that stabilizes the active kinase conformation in a phosphorylation-
dependent manner.23, 48 Thus, our data further suggest cellular inhibition of BTK, Fer and EphB4
kinase activities upon imatinib, identifying them as candidate signal transducers of Bcr-Abl-
mediated and imatinib-sensitive leukemia cell transformation. It is noteworthy that in case of
BTK earlier data argue against an essential function in Bcr-Abl signaling, based on evidence
that BTK inactivation showed no inhibitory effect in Bcr-Abl-transformed murine cells.49
However, despite this data, cell-type specific requirements for BTK are conceivable, for example
in case of reduced signaling capacity of Bcr-Abl due to endogenous expression at considerably
lower levels compared to ectopic overexpression in murine model cell lines.
In our experiments, imatinib treatment of K562 cells markedly decreased the tyrosine
phosphorylation of focal adhesion kinase (FAK) at Tyr-883 according to the assigned IPI
database identifier (Table 1), which corresponds to Tyr-861 in the commonly used UniProt
knowledgebase entry FAK1_HUMAN. Phosphorylation of FAK at Tyr-861 is up-regulated in
Ras-transformed cells and required for Ras-mediated transformation.50 As constitutive Ras
activation represents a hallmark of Bcr-Abl transformation, our data point to FAK Tyr-861
phosphorylation as a previously unknown switch point in CML cell signaling. We further
detected dose-dependent dephosphorylation of protein kinase C (PKC) at Tyr-313 and Tyr-
334 upon imatinib treatment (Figure 4B, Table 1). These tyrosine residues reside in the hinge
region between the regulatory and catalytic domains of PKC and have been implicated in the
Page 20
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
22/39
21
regulation of apoptosis in glioma cells.51, 52 Phosphorylation at Tyr-313 plays a role in diverse
signaling responses including keratinocyte differentiation and thromboxane A2 generation in
platelets.53, 54 Thus, our identification of these regulation events in imatinib-treated CML cells
may reflect a role of PKCin the propagation of Bcr-Abl-induced signals and warrant further
examination of this kinase in leukemia cell transformation. We also detected imatinib-sensitive
tyrosine phosphorylation events in the N-terminal region of the Src family kinase Yes and the C-
terminal part of the cytoplasmic tyrosine kinase ACK. As these modifications have not been
functionally characterized yet, their possible roles in Bcr-Abl signaling cannot be inferred from
published data. In addition to imatinib effects on tyrosine phosphorylation levels we detected
inhibitor-repressed serine and threonine phosphorylations on protein kinases such as mitogen-
activated protein kinase kinase kinase 3 (MAP3K3), 90 kDa ribosomal protein S6 kinase
(RSK1), casein kinase 2 (CK22) and PCTAIRE1, with the latter exhibiting as much as 75%
reduction of Ser-71 phosphorylation even at the low inhibitor dose of 1 M imatinib (Table 1). To
explore potential relationships among all proteins that were either prevented from resin binding
or harbored down-regulated phosphorylation sites upon imatinib treatment, we mapped them on
known protein interactions by STRING analysis (Figure 5). Notably, imatinib-regulated
phosphoproteins such as BTK, Fer, FAK and others were extensively connected to direct
imatinib targets like Bcr-Abl and Syk, further emphasizing their putative roles as downstream
signal transducers.
Surprisingly, imatinib treatment of cells not only caused site-specific reduction of but also
triggered a subset of phosphorylations in the kinase inhibitor-enriched subproteome
(Supplementary Table 3). These were exclusively found on serine and threonine residues, thus
contrasting the high prevalence of tyrosine amongst the imatinib-repressed phosphorylations.
Almost half of all proteins with imatinib-induced phosphorylation sites have reported cell cycle
functions. For example, we detected increased phosphorylation on the protein kinases polo-like
ge 21 of 38
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
23/39
22
kinase 1 (Plk1), TTK, Wee1 and Myt1, which have well established functions in the entry and
progression through mitosis. Plk1, which is highly expressed in many leukemia cell lines, has
recently been described as a promising target for therapeutic intervention in hematological
malignancies.55 Notably, our analysis revealed Plk1 phosphorylation occurring at Thr-210 in the
activation loop, thus indicating enzymatic activation of the kinase upon short term imatinib
treatment. Although the statistical significance of this regulation needs to be verified and the
underlying molecular mechanisms remain to be elucidated, Plk1 activation may be part of a
cellular response to counteract the immediate cellular consequences of Bcr-Abl kinase
suppression. Thus, our results might warrant investigations how or whether therapeutic Plk1
inhibition synergizes with cellular Bcr-Abl inactivation regarding the anti-proliferative and
apoptotic effects on CML cells.
Page 22
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
24/39
23
Conclusions and Outlook
Targeted intervention strategies with kinase inhibitors have already made an enormous impact
on the treatment of several human cancers. The role of such therapies is likely to increase in the
years to come, considering the large number of kinase-selective drugs currently in pre-clinical
and clinical development.4 Proteomics approaches can contribute valuable information to such
efforts, including drug selectivity assessments in relevant biological systems and the
identification of alternative molecular targets for pharmacological intervention. Target flexibility in
cancer treatment is desirable, given that the selective pressure imposed on cancer cells
frequently leads to drug resistance due to desensitizing mutations in the targeted, disease-
causing oncogenes.7, 8 This has been extensively documented for the transforming Bcr-Abl
tyrosine kinase in imatinib-treated chronic myeloid leukemia (CML) patients, but represents a
pervading theme as evident from, for example, the occurrence of drug-resistant epidermal
growth factor receptor (EGFR) kinase mutants in non-small cell lung cancer therapy with the
EGFR inhibitors gefitinib and erlotinib.56-58 One potential strategy to overcome resistance could
involve inhibition of protein kinases, which are essential downstream mediators of oncogenic
signaling emanating from kinases such as Bcr-Abl. We have implemented a chemical
phosphoproteomics strategy to identify such transducers upon imatinib exposure, by targeted
analysis of a pharmacologically tractable sub-proteome isolated by affinity purification with non-
selective kinase inhibitors. These proof-of-concept experiments provide preliminary evidence for
so far unknown kinases in imatinib-regulated K562 cell signaling, which represent candidates for
further validation and functional studies. Our proteomics strategy is generic and can be applied
to other kinase inhibitors. For example, comprehensive analysis of essential and druggable
mediators of EGFR signaling in non-small cell lung cancer cells might define therapeutic back-
up strategies to overcome the frequent EGFR resistance upon prolonged gefitinib or erlotinib
treatment. Moreover, concomitant inhibition of both primary oncogenic kinases and their
ge 23 of 38
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
25/39
24
essential signal transducers might effectively counteract resistance formation, as individual
target mutations would not suffice to evade from such poly-pharmacological regimens.
In addition to the phosphoproteomic identification of signaling factors situated downstream of
direct imatinib targets, our integrated approach recapitulated known imatinib targets from K562
cells, as these were competed from the generic kinase inhibitor resin in lysates from imatinib
treated cells.26 Notably, this part of our analysis was not only confirmatory but also identified
previously unknown imatinib-interacting proteins such as the key metabolic enzyme FAD
synthetase. Our identification of such a key metabolic enzyme as off-target raises the issue of
potential dose-limiting toxicity resulting from its likely pharmacological inhibition. We think further
in vitroand cellular studies are warranted to verify enzymatic inhibition of FAD synthetase by
imatinib and by related compounds in clinical development. Taken together, our sensitive
proteomics approach that integrates kinase inhibitor selectivity analysis with phosphoproteome
quantification in the kinase-enriched sub-proteome should have considerable utility for discovery
and development efforts aiming for improved targeted intervention strategies in human
malignancies.
Page 24
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
26/39
25
Acknowledgment.We thank Axel Ullrich for his generous support of our work. We thank
Matthias Mann and Jrgen Cox for early access to the MaxQuant software. We further thank
Jesper Olsen and Kirti Sharma for help and advice. This work was supported by a grant from
the Novartis-Stiftung fr therapeutische Forschung.
Supporting Information Available: Supplementary Table 1, list of all protein groups
identified in this study. Supplementary Table 2, peptide evidence data for all identified peptides.
Supplementary Table 3, all identified class I phosphorylation sites. Supplementary Table 4,
STRING-derived interactions among all kinase inhibitor resin-bound proteins that were
quantified with protein ratios and/or class I phosphorylation site ratios in both biological replicate
experiments. Supplementary Table 5, STRING-derived interactions among all proteins with
either imatinib-reduced kinase inhibitor resin binding or were imatinib-inhibited phosphorylation
sites. Supplementary Figure 1, interaction network constituted by inhibitor resin-bound proteins.
Supplementary Figure 2, representative MS spectra of K562 cell proteins exhibiting reduced
inhibitor resin binding upon cellular imatinib treatment.
ge 25 of 38
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
27/39
26
References
1. Blume-Jensen, P.; Hunter, T., Oncogenic kinase signalling. Nature2001, 411, (6835), 355-65.
2. Krause, D. S.; Van Etten, R. A., Tyrosine kinases as targets for cancer therapy. N Engl J Med
2005, 353, (2), 172-87.
3. Gschwind, A.; Fischer, O. M.; Ullrich, A., The discovery of receptor tyrosine kinases: targets for
cancer therapy. Nat Rev Cancer2004, 4, (5), 361-70.
4. Zhang, J.; Yang, P. L.; Gray, N. S., Targeting cancer with small molecule kinase inhibitors. Nat
Rev Cancer2009, 9, (1), 28-39.
5. Capdeville, R.; Buchdunger, E.; Zimmermann, J.; Matter, A., Glivec (STI571, imatinib), a
rationally developed, targeted anticancer drug. Nat Rev Drug Discov2002, 1, (7), 493-502.
6. Faderl, S.; Talpaz, M.; Estrov, Z.; O'Brien, S.; Kurzrock, R.; Kantarjian, H. M., The biology ofchronic myeloid leukemia. N Engl J Med1999, 341, (3), 164-72.
7. Weisberg, E.; Manley, P. W.; Cowan-Jacob, S. W.; Hochhaus, A.; Griffin, J. D., Second
generation inhibitors of BCR-ABL for the treatment of imatinib-resistant chronic myeloid leukaemia. Nat
Rev Cancer2007, 7, (5), 345-56.
8. Daub, H.; Specht, K.; Ullrich, A., Strategies to overcome resistance to targeted protein kinase
inhibitors. Nat Rev Drug Discov2004, 3, (12), 1001-10.
9. Quintas-Cardama, A.; Kantarjian, H.; Cortes, J., Flying under the radar: the new wave of BCR-
ABL inhibitors. Nat Rev Drug Discov2007, 6, (10), 834-48.
10. Schindler, T.; Bornmann, W.; Pellicena, P.; Miller, W. T.; Clarkson, B.; Kuriyan, J., Structural
mechanism for STI-571 inhibition of abelson tyrosine kinase. Science2000, 289, (5486), 1938-42.
11. Brehme, M.; Hantschel, O.; Colinge, J.; Kaupe, I.; Planyavsky, M.; Kocher, T.; Mechtler, K.;
Bennett, K. L.; Superti-Furga, G., Charting the molecular network of the drug target Bcr-Abl. Proc Natl
Acad Sci U S A 2009, 106, (18), 7414-9.
12. Preisinger, C.; Kolch, W., The Bcr-Abl kinase regulates the actin cytoskeleton via a GADS/Slp-
76/Nck1 adaptor protein pathway. Cell Signal2010, 22, (5), 848-56.
13. Olsen, J. V.; Blagoev, B.; Gnad, F.; Macek, B.; Kumar, C.; Mortensen, P.; Mann, M., Global, in
vivo, and site-specific phosphorylation dynamics in signaling networks. Cell2006, 127, (3), 635-48.
Page 26
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
28/39
27
14. Schreiber, T. B.; Mausbacher, N.; Breitkopf, S. B.; Grundner-Culemann, K.; Daub, H.,
Quantitative phosphoproteomics--an emerging key technology in signal-transduction research.
Proteomics2008, 8, (21), 4416-32.
15. Lemeer, S.; Heck, A. J., The phosphoproteomics data explosion. Curr Opin Chem Biol2009, 13,
(4), 414-20.
16. Dephoure, N.; Zhou, C.; Villen, J.; Beausoleil, S. A.; Bakalarski, C. E.; Elledge, S. J.; Gygi, S. P.,
A quantitative atlas of mitotic phosphorylation. Proc Natl Acad Sci U S A 2008, 105, (31), 10762-7.
17. Choudhary, C.; Olsen, J. V.; Brandts, C.; Cox, J.; Reddy, P. N.; Bohmer, F. D.; Gerke, V.;
Schmidt-Arras, D. E.; Berdel, W. E.; Muller-Tidow, C.; Mann, M.; Serve, H., Mislocalized activation of
oncogenic RTKs switches downstream signaling outcomes. Mol Cell2009, 36, (2), 326-39.
18. Grimsrud, P. A.; Swaney, D. L.; Wenger, C. D.; Beauchene, N. A.; Coon, J. J.,
Phosphoproteomics for the masses. ACS Chemical Biology2010, 5, (1), 105-19.
19. Zhao, Y.; Jensen, O. N., Modification-specific proteomics: strategies for characterization of post-
translational modifications using enrichment techniques. Proteomics2009, 9, (20), 4632-41.
20. Cox, J.; Mann, M., MaxQuant enables high peptide identification rates, individualized p.p.b.-range
mass accuracies and proteome-wide protein quantification. Nat Biotechnol2008, 26, (12), 1367-72.
21. Goss, V. L.; Lee, K. A.; Moritz, A.; Nardone, J.; Spek, E. J.; MacNeill, J.; Rush, J.; Comb, M. J.;
Polakiewicz, R. D., A common phosphotyrosine signature for the Bcr-Abl kinase. Blood2006, 107, (12),
4888-97.
22. Pan, C.; Olsen, J. V.; Daub, H.; Mann, M., Global effects of kinase inhibitors on signaling
networks revealed by quantitative phosphoproteomics. Mol Cell Proteomics2009, 8, (12), 2796-808.
23. Nolen, B.; Taylor, S.; Ghosh, G., Regulation of protein kinases; controlling activity through
activation segment conformation. Mol Cell2004, 15, (5), 661-75.
24. Daub, H.; Olsen, J. V.; Bairlein, M.; Gnad, F.; Oppermann, F. S.; Korner, R.; Greff, Z.; Keri, G.;
Stemmann, O.; Mann, M., Kinase-selective enrichment enables quantitative phosphoproteomics of the
kinome across the cell cycle. Mol Cell2008, 31, (3), 438-48.
25. Wissing, J.; Jansch, L.; Nimtz, M.; Dieterich, G.; Hornberger, R.; Keri, G.; Wehland, J.; Daub, H.,
Proteomics analysis of protein kinases by target class-selective prefractionation and tandem mass
spectrometry. Mol Cell Proteomics2007, 6, (3), 537-47.
ge 27 of 38
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
29/39
28
26. Bantscheff, M.; Eberhard, D.; Abraham, Y.; Bastuck, S.; Boesche, M.; Hobson, S.; Mathieson, T.;
Perrin, J.; Raida, M.; Rau, C.; Reader, V.; Sweetman, G.; Bauer, A.; Bouwmeester, T.; Hopf, C.; Kruse,
U.; Neubauer, G.; Ramsden, N.; Rick, J.; Kuster, B.; Drewes, G., Quantitative chemical proteomics
reveals mechanisms of action of clinical ABL kinase inhibitors. Nat Biotechnol2007, 25, (9), 1035-44.
27. Godl, K.; Gruss, O. J.; Eickhoff, J.; Wissing, J.; Blencke, S.; Weber, M.; Degen, H.; Brehmer, D.;
Orfi, L.; Horvath, Z.; Keri, G.; Muller, S.; Cotten, M.; Ullrich, A.; Daub, H., Proteomic characterization of
the angiogenesis inhibitor SU6668 reveals multiple impacts on cellular kinase signaling. Cancer Res
2005, 65, (15), 6919-26.
28. Wessel, D.; Flgge, U. I., A method for the quantitative recovery of protein in dilute solution in the
presence of detergents and lipids. Analytical Biochemistry1984, 138, (1), 141-143.
29. Shevchenko, A.; Tomas, H.; Havlis, J.; Olsen, J. V.; Mann, M., In-gel digestion for mass
spectrometric characterization of proteins and proteomes. Nat Protoc2006, 1, (6), 2856-60.
30. Rappsilber, J.; Mann, M.; Ishihama, Y., Protocol for micro-purification, enrichment, pre-
fractionation and storage of peptides for proteomics using StageTips. Nat Protoc2007, 2, (8), 1896-906.
31. Larsen, M. R.; Thingholm, T. E.; Jensen, O. N.; Roepstorff, P.; Jorgensen, T. J., Highly selective
enrichment of phosphorylated peptides from peptide mixtures using titanium dioxide microcolumns. Mol
Cell Proteomics2005, 4, (7), 873-86.
32. Pinkse, M. W.; Uitto, P. M.; Hilhorst, M. J.; Ooms, B.; Heck, A. J., Selective isolation at the
femtomole level of phosphopeptides from proteolytic digests using 2D-NanoLC-ESI-MS/MS and titanium
oxide precolumns. Anal Chem2004, 76, (14), 3935-43.
33. Schroeder, M. J.; Shabanowitz, J.; Schwartz, J. C.; Hunt, D. F.; Coon, J. J., A neutral loss
activation method for improved phosphopeptide sequence analysis by quadrupole ion trap mass
spectrometry. Anal Chem2004, 76, (13), 3590-8.
34. Olsen, J. V.; de Godoy, L. M. F.; Li, G.; Macek, B.; Mortensen, P.; Pesch, R.; Makarov, A.; Lange,
O.; Horning, S.; Mann, M., Parts per Million Mass Accuracy on an Orbitrap Mass Spectrometer via Lock
Mass Injection into a C-trap. Mol Cell Proteomics2005, 4, (12), 2010-2021.
35. Jensen, L. J.; Kuhn, M.; Stark, M.; Chaffron, S.; Creevey, C.; Muller, J.; Doerks, T.; Julien, P.;Roth, A.; Simonovic, M.; Bork, P.; von Mering, C., STRING 8--a global view on proteins and their
functional interactions in 630 organisms. Nucl. Acids Res. 2009, 37, (suppl_1), D412-416.
36. Cline, M. S.; Smoot, M.; Cerami, E.; Kuchinsky, A.; Landys, N.; Workman, C.; Christmas, R.;
Avila-Campilo, I.; Creech, M.; Gross, B.; Hanspers, K.; Isserlin, R.; Kelley, R.; Killcoyne, S.; Lotia, S.;
Page 28
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
30/39
29
Maere, S.; Morris, J.; Ono, K.; Pavlovic, V.; Pico, A. R.; Vailaya, A.; Wang, P. L.; Adler, A.; Conklin, B. R.;
Hood, L.; Kuiper, M.; Sander, C.; Schmulevich, I.; Schwikowski, B.; Warner, G. J.; Ideker, T.; Bader, G.
D., Integration of biological networks and gene expression data using Cytoscape. Nat Protoc 2007, 2,
(10), 2366-82.
37. Godl, K.; Wissing, J.; Kurtenbach, A.; Habenberger, P.; Blencke, S.; Gutbrod, H.; Salassidis, K.;
Stein-Gerlach, M.; Missio, A.; Cotten, M.; Daub, H., An efficient proteomics method to identify the cellular
targets of protein kinase inhibitors. Proc Natl Acad Sci U S A 2003, 100, (26), 15434-9.
38. Sharma, K.; Kumar, C.; Keri, G.; Breitkopf, S. B.; Oppermann, F. S.; Daub, H., Quantitative
analysis of kinase-proximal signaling in lipopolysaccharide-induced innate immune response. J Proteome
Res2010, 9, (5), 2539-49.
39. Atwell, S.; Adams, J. M.; Badger, J.; Buchanan, M. D.; Feil, I. K.; Froning, K. J.; Gao, X.; Hendle,
J.; Keegan, K.; Leon, B. C.; Muller-Dieckmann, H. J.; Nienaber, V. L.; Noland, B. W.; Post, K.;
Rajashankar, K. R.; Ramos, A.; Russell, M.; Burley, S. K.; Buchanan, S. G., A novel mode of Gleevec
binding is revealed by the structure of spleen tyrosine kinase. J Biol Chem2004, 279, (53), 55827-32.
40. Rix, U.; Hantschel, O.; Durnberger, G.; Remsing Rix, L. L.; Planyavsky, M.; Fernbach, N. V.;
Kaupe, I.; Bennett, K. L.; Valent, P.; Colinge, J.; Kocher, T.; Superti-Furga, G., Chemical proteomic
profiles of the BCR-ABL inhibitors imatinib, nilotinib, and dasatinib reveal novel kinase and nonkinase
targets. Blood2007, 110, (12), 4055-63.
41. Huerta, C.; Borek, D.; Machius, M.; Grishin, N. V.; Zhang, H., Structure and mechanism of a
eukaryotic FMN adenylyltransferase. J Mol Biol2009, 389, (2), 388-400.
42. Law, C. L.; Chandran, K. A.; Sidorenko, S. P.; Clark, E. A., Phospholipase C-gamma1 interacts
with conserved phosphotyrosyl residues in the linker region of Syk and is a substrate for Syk. Mol Cell
Biol1996, 16, (4), 1305-15.
43. Schymeinsky, J.; Sindrilaru, A.; Frommhold, D.; Sperandio, M.; Gerstl, R.; Then, C.; Mocsai, A.;
Scharffetter-Kochanek, K.; Walzog, B., The Vav binding site of the non-receptor tyrosine kinase Syk at
Tyr 348 is critical for beta2 integrin (CD11/CD18)-mediated neutrophil migration. Blood2006, 108, (12),
3919-27.
44. Schymeinsky, J.; Then, C.; Sindrilaru, A.; Gerstl, R.; Jakus, Z.; Tybulewicz, V. L.; Scharffetter-
Kochanek, K.; Walzog, B., Syk-mediated translocation of PI3Kdelta to the leading edge controls
lamellipodium formation and migration of leukocytes. PLoS One2007, 2, (11), e1132.
ge 29 of 38
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
31/39
30
45. Yankee, T. M.; Keshvara, L. M.; Sawasdikosol, S.; Harrison, M. L.; Geahlen, R. L., Inhibition of
signaling through the B cell antigen receptor by the protooncogene product, c-Cbl, requires Syk tyrosine
317 and the c-Cbl phosphotyrosine-binding domain. J Immunol1999, 163, (11), 5827-35.
46. Simon, M.; Vanes, L.; Geahlen, R. L.; Tybulewicz, V. L., Distinct roles for the linker region
tyrosines of Syk in FcepsilonRI signaling in primary mast cells. J Biol Chem2005, 280, (6), 4510-7.
47. Keshvara, L. M.; Isaacson, C. C.; Yankee, T. M.; Sarac, R.; Harrison, M. L.; Geahlen, R. L., Syk-
and Lyn-dependent phosphorylation of Syk on multiple tyrosines following B cell activation includes a site
that negatively regulates signaling. J Immunol1998, 161, (10), 5276-83.
48. Oppermann, F. S.; Gnad, F.; Olsen, J. V.; Hornberger, R.; Greff, Z.; Keri, G.; Mann, M.; Daub, H.,
Large-scale proteomics analysis of the human kinome. Mol Cell Proteomics2009, 8, (7), 1751-64.
49. MacPartlin, M.; Smith, A. M.; Druker, B. J.; Honigberg, L. A.; Deininger, M. W., Bruton's tyrosine
kinase is not essential for Bcr-Abl-mediated transformation of lymphoid or myeloid cells. Leukemia2008,22, (7), 1354-60.
50. Lim, Y.; Han, I.; Jeon, J.; Park, H.; Bahk, Y. Y.; Oh, E. S., Phosphorylation of focal adhesion
kinase at tyrosine 861 is crucial for Ras transformation of fibroblasts. J Biol Chem2004, 279, (28), 29060-
5.
51. Lu, W.; Finnis, S.; Xiang, C.; Lee, H. K.; Markowitz, Y.; Okhrimenko, H.; Brodie, C., Tyrosine 311
is phosphorylated by c-Abl and promotes the apoptotic effect of PKCdelta in glioma cells. Biochem
Biophys Res Commun2007, 352, (2), 431-6.
52. Lu, W.; Lee, H. K.; Xiang, C.; Finniss, S.; Brodie, C., The phosphorylation of tyrosine 332 is
necessary for the caspase 3-dependent cleavage of PKCdelta and the regulation of cell apoptosis. Cell
Signal2007, 19, (10), 2165-73.
53. Murugappan, S.; Shankar, H.; Bhamidipati, S.; Dorsam, R. T.; Jin, J.; Kunapuli, S. P., Molecular
mechanism and functional implications of thrombin-mediated tyrosine phosphorylation of PKCdelta in
platelets. Blood2005, 106, (2), 550-7.
54. Balasubramanian, S.; Zhu, L.; Eckert, R. L., Apigenin inhibition of involucrin gene expression is
associated with a specific reduction in phosphorylation of protein kinase Cdelta Tyr311. J Biol Chem2006, 281, (47), 36162-72.
55. Ikezoe, T.; Yang, J.; Nishioka, C.; Takezaki, Y.; Tasaka, T.; Togitani, K.; Koeffler, H. P.;
Yokoyama, A., A novel treatment strategy targeting polo-like kinase 1 in hematological malignancies.
Leukemia2009, 23, (9), 1564-76.
Page 30
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
32/39
31
56. Kobayashi, S.; Boggon, T. J.; Dayaram, T.; Janne, P. A.; Kocher, O.; Meyerson, M.; Johnson, B.
E.; Eck, M. J.; Tenen, D. G.; Halmos, B., EGFR mutation and resistance of non-small-cell lung cancer to
gefitinib. N Engl J Med2005, 352, (8), 786-92.
57. Pao, W.; Miller, V. A.; Politi, K. A.; Riely, G. J.; Somwar, R.; Zakowski, M. F.; Kris, M. G.; Varmus,
H., Acquired resistance of lung adenocarcinomas to gefitinib or erlotinib is associated with a second
mutation in the EGFR kinase domain. PLoS Medicine2005, 2, (3), e73.
58. Blencke, S.; Ullrich, A.; Daub, H., Mutation of threonine 766 in the epidermal growth factor
receptor reveals a hotspot for resistance formation against selective tyrosine kinase inhibitors. J Biol
Chem2003, 278, (17), 15435-40.
ge 31 of 38
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
33/39
32
Figure Legends
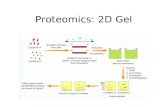
Figure 1. Schematic illustration of the experimental strategy. Three populations of K562 CML
cells were SILAC-encoded with normal or isotopically labeled arginine and lysine and either 1
M imatinib, 10 M imatinib or DMSO before lysis. Two biological replicate experiments were
performed with different SILAC schemes. In both experiments, each of the three lysates was
subjected to separate affinity purification with a mixture of five kinase inhibitor resins. Resin-
bound proteins were eluted and pooled. Subsequently, fractions of the pooled kinase-enriched
material were separated by gel electrophoresis prior to or by SCX chromatography after
digestion with trypsin. Phosphopeptides were enriched with titanium dioxide beads and total
peptide samples were prepared by desalting with C18-StageTips from all peptide fractions. All
resulting samples were analyzed by LC-MS/MS on an LTQ-Orbitrap.
Figure 2. Overview of results from SILAC experiments. (A) Comparison of the two independent
SILAC experiments regarding the quantified proteins and quantified phosphorylation sites with
confident site localization (class I sites with p 0.75). Numbers are separately shown for all
proteins and for protein kinases. (B) Numbers of all proteins and protein kinases for which
protein ratios and class I phosphorylation site ratios were quantified. The overlapping regions
indicate the protein and protein kinase numbers for which both protein and phosphorylation site
ratios were obtained within this study. (C) Numbers and distribution of serine, threonine and
tyrosine phosphorylation for all quantified class I phosphorylation sites are shown for each of the
two biological replicate experiments and the overlap between the two experiments.
Figure 3. Quantified K562 proteins exhibiting dose-dependent suppression of kinase inhibitor
resin binding upon cellular imatinib treatment. Average values are shown for K562 cell proteins
Page 32
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
34/39
33
quantified in both replicate experiments whose interaction with the immobilized kinase inhibitors
was reduced by at least 50% upon 10 M imatinib treatment of K562 cells.
Figure 4. Quantitative phosphoproteomics of imatinib effects on kinase inhibitor beads-captured
proteins. (A) Scatter plot comparison of log2 transformed class I phosphorylation site ratios upon
10 M cellular imatinib treatment. Sites that were consistently down-regulated (up-regulated) in
both biological replicate experiments are highlighted in red (green). Moreover, phosphorylation
ratios for Tyr-714 of FER and for Tyr-313 of PKC are indicated. (B) Examples of imatinib-
regulated downstream kinases identified by quantitative MS. SILAC spectra are shown forimatinib-regulated, pTyr-containing phosphopeptides (left panels) and unchanged, non-
phosphorylated peptides (right panels) derived from the protein kinases Fer and PKC in
experiment 1. Values for measured and pooling error-corrected (normalized) SILAC ratios are
shown.
Figure 5. Interactome of imatinib-regulated proteins. All proteins that either exhibited reduced
kinase inhibitor resin binding or which harbored imatinib-repressed phosphorylation sites were
used for STRING analysis. The resulting network was visualized with Cytoscape.
ge 33 of 38
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
35/39
DMSO 1 M imatinib 10 M imatinibExp. 1
1 M imatinib 10 M imatinib DMSOExp. 2
0 0Arg /Lys 6 4Arg /Lys 10 8Arg /Lys
K562 K562 K562
0 0Arg /Lys 6 4Arg /Lys 10 8Arg /LysK562 K562 K562
Cell lysis
NH
OH
O
NH
O
SU6668
N
N N
N
HO
Cl
COOH
Purvalanol B
N
NH2
HN
N
O O
BisX
N ONH
OH2N
VI16832
O
N
N
HN Cl
F
O
H2N
AX14596
Elution
Pooling
LC-MS/MS analysis on LTQ-Orbitrap
Trypticdigestion
Trypticdigestion
TiO2TiO2
Titanspherephosphopeptideenrichment
C18desalting
StageTip
TiO2TiO2
C18 C18
Figure 1
Page 34
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
36/39
137
33 13
898574
370
487 683
105
504
201 163
47
30
136
Proteins quantifiedwith pSTY ratios
Proteins quantifiedwith protein ratios901 356374
Experiment 2
Experiment 1
All proteins Protein kinases
Quantifiedprotein ratios
QuantifiedpSTY ratios
A
B
C
All proteins Protein kinases
Figure 2
81.25% 13.93% 4.82%
79.10% 15.14% 5.76%
81.07% 13.25% 5.68%
1472Exp. 1 1196 205 71
1268Exp. 2 1003 192 73
898Both 728 119 51
Class I total pSer pThr pTyr
ge 35 of 38
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
37/39
Figure 3
Page 36
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
38/39
7 04.0 705 .0 706.0 707.0 708.0 709.0
m/z0
10
20
30
40
50
60
70
80
90
100
704.30
705.81
704.80706.31
707.80708.30
706.81705.30
708.81707.30 709.31
703.79
0LysDMSO control
4Lys1 M imatinib
8Lys10 M imatinib
FER: QEDGGV SSSGLKpY
704 705 706 707 708 709 710 711
m/z0
10
20
30
40
50
60
70
80
90
100707.02 709.35
709.68706.69
704.01
707.36
710.01
704.34707.69
710.35709.01
708.02704.67 710.68708.69
705.01
703.34703.67
0ArgDMSO control
6Arg1 M imatinib
10Arg10 M imatinib
FER: ESHGKPGEYVLSVYSDGQR
1019 1020 1021 1022 1023 1024 1025 1026 1027 1028 1029 1030
m/z0
10
20
30
40
50
60
70
80
90
100
1019.95
1023.971024.47
1027.96
1020.45 1024.981028.46
1028.961025.48
1026.441020.96 1029.47
1018.95 1019.45
0LysDMSO control
4Lys1 M imatinib
8Lys10 M imatinib
6 79 .0 6 80 .0 6 81 .0 6 82 .0 6 83 .0 6 84 .0 6 85 .0m/z0
10
20
30
40
50
60
70
80
90
100
683.39
678.89
681.90 683.89
679.39684.39
682.40
679.89682.89
684.89680.39
678.39 681.40
0ArgDMSO control
6Arg1 M imatinib
10Arg10 M imatinib
Relativeabundance
Relativeabundance
Relativeabundance
Relative
abundance
Measured Normalized1 M imatinib / control 0.38 0.52
10 M imatinib / control 0.18 0.25
Measured Normalized1 M imatinib / control 0.77 1.21
10 M imatinib / control 0.78 1.17
Measured Normalized1 M imatinib / control 0.43 0.6210 M imatinib / control 0.21 0.36
Measured Normalized1 M imatinib / control 0.85 1.1310 M imatinib / control 0.83 1.06
A
B
Down-regulated
Up-regulatedOther
Log (10 M imatinib / control) - exp. 12
Lo
g
(10M
imati
nib
/co
ntro
l)-
exp
.2
2
pSTY sites
Figure 4
FER-Tyr-714
ge 37 of 38
ACS Paragon Plus Environment
Journal of Proteome Research
-
8/8/2019 Journal of Proteome Research 2010 Breitkopf
39/39
Regulated pSTY ratio
Regulated protein ratio
Protein kinase
Non protein kinase
BCR-ABL SHIP2
Figure 5
Page 38Journal of Proteome Research