Introduction to RNA-Seq in GeneSpring NGS Software · Introduction to RNA-Seq • In a few years,...
Transcript of Introduction to RNA-Seq in GeneSpring NGS Software · Introduction to RNA-Seq • In a few years,...

Learn more at www.genespring.com
Introduction to RNA-Seq in GeneSpring NGS Software
Dipa Roy Choudhury, Ph.D. Strand Scientific Intelligence and Agilent Technologies

Introduction to RNA-Seq
• In a few years, massively parallel cDNA sequencing, or RNA-seq,
has allowed many advances in the characterization and
quantification of transcriptomes.
• Rapidly decreasing sequencing cost and massively-parallel
sequencing technologies has resulted in a dramatic increase in
the quantity of data that needs to be analyzed.
• Therefore, the need of the day is to build a tool that will enable
the analysis and integration of data produced on multiple
platforms and using multiple methods.
• Agilent has designed NGS analysis in GeneSpring keeping in
mind the biologist, who is interested in answering REAL biological
questions and does not want to become a bioinformatics expert
just to do their work.

GeneSpring
NGS
Agilent SureSelect Target Enrichment
5 Map Reads to the
Ref. Genome 6 Quality Control on
Mapping 7 Detect
SNPs/InDels or
Diff. Spliced
Genes
8 Find Biological
Relevance for
your Results

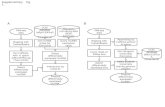
GeneSpring NGS Provides Downstream
Analysis of Next-Gen Sequencing data
Data File (Reads + Quality)
Control Software
Data File (Reads + Quality)
ELAND/BIOSCOPE/BWA…
Reads aligned to genome
Reads aligned to genome
GeneSpring NGS
Primary Analysis Tertiary Analysis Secondary Analysis
FASTQ, … BAM, …
FASTQ

Questions we can seek to answer using
GeneSpring RNA-Seq Workflow are
What are the differentially expressed genes?
What are the differentially spliced genes?
Are there any SNPs in the transcriptome?
Can we identify gene fusion events?
Can we identify novel genes, novel exons, and novel splice junctions?

6
Import Data and Annotations

Download Human build HG18 from the
Agilent Server
8
Open Human
“tree” Select hg18 and
Homologene
Groups
Click [UPDATE]
to start the
download
Annotations files
can be quite
large

Baits and Targets for SureSelect Catalog
Kits are Pre-loaded
9
List of Agilent
SureSelect Catalog
Kits

Creating a new Experiment
10
Click “New
Experiment” icon to
start new experiment
Or select “New
Experiment” from
Project menu

Create New GeneSpring NGS Experiment
Provide some useful and descriptive name
Be sure to select “NGS” as the Analysis type

Importing Data: Choosing Metadata New organisms supported on demand
Indicate organism,
build and transcript
model
Prepackaged
annotations
available for a
variety of
organisms
Support for Illumina,
Life Technology, 454
Roche
Support for single end,
paired end, and mate
pair protocols

Provide Information on Sequencing platform and
library layout for SureSelect specific kits
13
Be sure to select the
previously loaded
Target region

GeneSpring NGS
Organization Of The Windows
14
Project and Experiment Navigator
Workflow Browser
Region View

15
Quality Control

Perform QC on reads and filter anomalous reads View reads by tile/lane and remove reads in anomalous tiles
Base
qualities in
the same
tiles show
bad
Perfect match
reads and too
many
mismatch
reads in this
tile.

Quality Inspection
Open Quality Control Manager
Press Compute to
calculate the Library
QC metrics
Press Compute to
calculate the Library
QC metrics

July 11 Page 18
QC Manager feature in SureSelect experiments
Off-target reads
can be removed
to focus analysis
Targeted Regions
(SureSelect Baits)
Off-target reads
can be removed
to focus analysis

Determine the expression values for
each gene

Run Quantification to determine raw gene counts Which reads contribute to a gene’s count?
These reads
do NOT
contribute
Only Reads
overlapping exonic
regions contribute to
the read count
Multiply mapping
reads contribute
fractionally to the
count

Quantification
Quantify Genes, Transcripts and Exons
Reverted back to the All
Aligned Reads list to
establish a base line. Feel
free to try a different
(filtered) read list
Unchecked will only count
reads falling completely
inside an exon.
New genes and exons are
discovered using
conservation data
(Conservation track in
Annotations)

Filter genes by RPKM: Results of Filtering
Profile plot of
genes that pass
the filter criteria
Profile plot of
genes that pass
the filter criteria

Handling overlapping genes Which reads contribute to a gene’s count?
These reads contribute to both
genes except for ABI data
which is strand specific
Overlapping gene on
negative strand
Gene on positive
strand

Determine expression values normalized across samples

Scatter Plot between Two Replicates

Detection of differentially expressed genes

Output of Differential Expression Analysis
P-values,
Corrected p-
values and fold
changes
Volcano Plot

Determine differentially spliced genes
detected

34
Identifying Differentially Spliced Genes
Compute the proportion of a
gene’s count that can be ascribed to
a particular transcript.
If the proportion for a particular
transcript changes substantially across conditions, the gene
is said to be differentially spliced.

The Challenge: Deconvoluting Transcript Read Counts
Which of the 4
transcripts do
these reads
come from?

Differential Splicing Analysis
View Results in Gene View
Ensure Splicing
Analysis Results
Entiy list is selected
Click “Gene
View” icon to
show Gene
View

Differential Splicing Analysis
Gene View
This transcript
is expressed
less in Tumor
Gene’s
RPKM
4 Transcript
RPKMs
4 Transcript
RPKMs
4 Transcript
RPKMs
Transcript
RPKM
This transcript
is expressed
more in Tumor
Possible
New Exon
Possible
New Exon

Determine SNPs be determined in the
transcriptome

GeneSpring NGS has a built-in SNP calling algorithm
Set Filters for SNP
statistical significance
Set filters for min number of
overlapping reads and min
number of overlapping
variant reads

GeneSpring NGS calls transcript effects for
each SNP and allows filtering of SNPs based on
these effects
Types of effects
predicted Change in
Amino Acid for
Non-
synonymous
SNPs

Viewing SNPs in the Genome Browser
Color-coded
indicator for a
Homozygous
SNP
Known in
dbSNP
GeneSpring
NGS SNP Call
In a Repeat
Region

Determine chimeric transcripts or
fusion genes

Identify Fusion Genes
In a K562 Leukemia cell line, GeneSpring NGS confirms the well-known
BCR-ABL1 gene fusion.
Filters set on the
Genome
Browser to show
only trans-
located reads
Several reads pairs for
the BCR gene on
chr22 with mates
translocated to the
ABL1 gene on chr9
The
corresponding
paired reads for
the ABL1 gene on
chr9

Detection of novel genes, exons, and
splice junctions

Identify Novel Exons and Genes
In a mouse myoblast study, GeneSpring NGS determines
a new exon for the FHL3 gene
Read clumps not aligned
with a known exon
Novel exon
determined by
GeneSpring NGS,
probably a new
transcription
start site
Add exon to gene if close to or within the
gene, otherwise call it a new
gene

Identify Novel Splice Junctions In a brain tissue expression study, GeneSpring NGS determines a new splice
junction in the DTX3 gene when considering only Refseq transcripts; this novel
splice junction is corroborated by a UCSC transcript.
Solid lines show
spliced reads
connecting the
1st and 3rd exons
of the RefSeq
transcript
The
corresponding
novel splice
junction found by
GeneSpring
NGS
Indeed, a known
UCSC transcript
that is not
present in
RefSeq validates
this discovery

49
Biological Contextualization
Pathway Analysis

Agilent GeneSpring NGS for SureSelect
Display the Results on a Pathway

Questions we can seek to answer using
GeneSpring RNA-Seq Workflow
What are the differentially expressed genes?
What are the differentially spliced genes?
Are there any SNPs in the transcriptome?
Can we identify gene fusion events?
Can we identify novel genes, novel exons, and novel splice junctions?

Summary
•Differential expression and splicing analysis
•Novel gene, exon and alternative splicing
discovery
•Gene Fusion Analysis
•SNP & InDel discovery and annotation with
dbSNP

Summary in General
•GeneSpring NGS supports both SureSelect RNASeq
experiments as well as RNA Seq experiments that don’t
use Sure Select
•The workflow steps in GeneSpring NGS application are
application specific and changes based on whether you
are analyzing a DNA-SEQ or RNA-SEQ experiment.
•It is possible to integrate data produced on multiple
platforms and using multiple methods in the same project
in GeneSpring.
•Multiple different visualization tools available to query the
data.

55
http://www.AVADIS-NGS.com