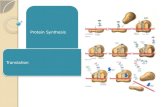
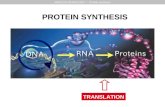
I. Structure and Mechanism: Protein Synthesis
description
Transcript of I. Structure and Mechanism: Protein Synthesis

I. Structure and Mechanism: Protein Synthesis
“Mechanism of the peptidyl-transfer reaction of the prokaryotic ribosome”
By: Trang Bui

Overview
• Introduction-Ribosome
• Structure of Peptidyl Transferase Center
• Mechanism of Peptide Bond Formation• Acid-base catalysis (enthalpic)
• Proper substrate positioning (entropic)
• Future Prospects

Ribosome• Prokaryote
– 50S subunit:• 23S rRNA and 5S rRNA• 31 proteins
– 30S subunit:• 26S rRNA• 21 proteins
• Eukaryote– 60S subunit:
• 28S rRNA, 5.8S rRNA, & 5S rRNA
• 49 proteins
– 40S subunit:• 18S rRNA• 33 proteins

The acceptor ends of A-site and P-site tRNAs are located in the 50S subunit facing the 30S subunit

Review: peptide bond formation

Peptidyl-transferase • Peptidyl transferase (PT) center is
the site for peptide bond formation• Located on 50S• Composed of RNA only and no
protein is found within 15 Ǻ
What can you conclude from this?Ribosome Ribozyme
L27 protein might be involved - deletion of 3 a.a. leads to impaired activity
- only exits in some organisms - suggesting that L27 facilitates the
proper placement of the tRNA at the PTC

Structure of the PT center
•Conserved & are located at the core of the PT center•tRNA analog (A site)•tRNA analog (P site)•23S rRNA bases

Knowing that ribosome is a ribozyme, what mechanism(s) do you think
might be used to form peptide bonds between 20 different a.a.?

Chemistry of peptide-bond formation
1: Deprotonation of the amino group2: Nucleophilic attack and formation of the zwitterionic tetrahedral intermediate3: Deprotonation and formation of the negatively charged intermediate4: Product formation and protonation of the leaving oxygen

Mechanism of Peptide Bond Formation
1. Contribution of general acid-base catalysis
2. Contribution of substrate positioning

Acid-base Catalysis
• In general, this means either proton transfer or abstraction
• Transition state: chemicals bonds are in the process of being made and broken
• Unstable + and – charges are being developed
• Stabilizing charges catalyzes the reaction by lowering the energy of the transition state

Acid-base Catalysis
• Facilitates the activation of weak nucleophiles
• Stabilization of poor leaving groups

Role of active site residue A2451 as general acid-base catalyst
• Crystallized ribosomes with a transition analog CCdA-pPuro
• N3 A2451 is in close contact (3 Ǻ) with a transition state analog of peptide bond formation, CCdA-pPuro
• A2451U mutation led to significantly reduced activity
• Thought to be involved in acid-base catalysis

N3 of A2451
• X-ray structures of the ribosome with a P-site tRNA A76 2’OH instead of 2’-deoxyadenosine
• Within hydrogen bond distance of the -amino group only in the pre-reaction state
• Intermediate’s oxyanion points away from the A2451-N3
A siteP site23S rRNA bases
WRONG

Experimental Procedure
Acid-base Catalysis

A site• Aminoacyl-tRNA binds to the A site in the range of 10 s-1
• The intrinsic rate of peptide-bond formation was estimated to be > 300 s-1
• Thus, the mechanism of peptide-bond formation cannot be studied with the native aa-tRNA under current experimental capabilities.
• Adio et al. (2006) examined the contribution of acid-base catalysis to peptide bond formation using ribosomes from E.coli with native aa-tRNA

Experimental Procedure
• The -amino group of aa-tRNA has a pKa of 8.
• Assuming that an ionization group (pKa = 7) on the ribosome is involved in the reaction, the rate of the peptide bond formation should be pH dependent.
• Since the accommodation step is pH independent, the reaction rate may become lower than the accommodation rate at a certain pH.
• Measured rates of aa-tRNA accommodation and peptide bond formation in the pH range between 6 and 9.
• pH<6 = EF-Tu precipitation
• pH>9 = tRNA cleavage

A-site Accommodation
• Measuring the rate of A-site accommodation using FRET– fMet-tRNA was labeled with a fluorescence donor, fluorescein– Phe-tRNA was labeled with a fluorescence quencher, QSY35– Time course of accommodation was measured using the
stopped-flow method• Conclusion: the rates of accommodation were identical at pH
values of 6,7, and 8
Curve 1: Phe-tRNA (QSY)Curve 2: Phe-tRNA

Peptide Bond Formation
• Time course of accommodation was measured using the quench flow method– f[3H]Met-tRNA
– [14C]Phe-tRNA
• Conclusion: The rate of peptide bond formation was independent of pH and indistinguishable from the rate of accommodation

Conclusion from pH reactions
• This can be explained in 2 ways:
1) The ionization group has no affect on the rate of the PT reaction
2) The chemical step was so fast that the contribution from the protonation step was insignificant
If you can’t study the chemistry step using native aa-tRNA, what can you do?

Uncoupling the Chemistry Step from the Accommodation Step
Phelac-tRNA• -OH is the nucleophile instead of -NH2
• Does not change the catalytic mechanism
• Formation of ester bond rate limiting step
• Measuring the reaction rate at pH 6-9 reveals that the reaction rate is independent of pH
• This indicates that acid-base catalysis is not used to a great extent

Peptide Bond Formation
1. Contribution of general acid-base catalysis
2. Contribution of substrate positioning

2’OH of A76• CCdA-p-Puro inhibits peptidyl
transferase activity
• Substituting 2’-OH of A76 by either 2’-deoxy or 2’-fluoro reduce the activity ~10^6 fold
• 2’-OH receives a proton from the a-amino group while simultaneously protonates the leaving 3’OH—proton shuttle
• 2’-OH of A76 orients the nucleophile
A site tRNA substrateP site tRNA substrateRibosome residuesHOH* might be used for a proton shuttle

Other Groups Involvement??
2’-OH of A2451
- Substitution of 2’OH of A2451 by hydrogen impairs peptidyl-transferase activity
- Interacts directly with the 2’OH group of the P-site tRNA
A site tRNA susbtratesP site tRNA substratesRibosome residues

• The ribosome brings ~10^7 fold enhancement in the rate of PT reaction compared with the second-order reaction in solution
• Entropy of activation is lowered
• Enthalpy of activation is the same for both reactions
• In acid-base and covalent catalysis, enzymes act by lowering the activation enthalpy

Mechanism of Peptide-bond Formation
• Conclusion: entropic catalysis is the major catalytic mechanism of peptide bond formation
Intra-reactant proton shuttling via the 2’OH of A76 of the P-site tRNA

Future Prospects
• Obtain structural and mechanistic information for eukaryotic ribosomes
• Examine the second important function of the PT center: termination of protein synthesis