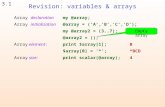
Global array
-
Upload
alfreda-tonelli -
Category
Science
-
view
95 -
download
0
Transcript of Global array

Global Array: DNA Oligonucleotide Microbial Diagnostic Microarray (MDM)
Tonelli Alfreda
13/09/2009

Introduction Pathogen isolation and identification considerations
enrichment and selective culture media appropriate incubation conditions
temperature atmosphere
various identification kits for confirmation interpretation of results is often subjective
qualified personnel essential Time to diagnosis a limiting factor
13/09/2009

Introduction A diagnostic test that will identify pathogens and zoonotic organisms, because of the great number of organisms, to this effect microarrays were chosen because they are; high throughput method easy to perform harmonizeable easy to interpret ? ? ?
13/09/2009

What is an MDM? An MDM is a Microbial Diagnostic
MicroarrayHigh throughput method for the detection of multiple genes belonging to one or more organismOligonucleotide microarray specific oligonucleotides, 35-70 nucleotides,
in length are chemically synthesized the oligonucleotides are spotted, at defined
positions, onto a glass slide to construct the array
binding of fluorescently labelled probes to the oligonucleotides is detected spectrophotometrically13/09/2009

What does an MDM look like ?
13/09/2009

Genes present on an MDM?
Probes to all genes of organisms of interest in veterinary and medical field Probes should be common within the genera inter and intra species specific
Probes are designed in such a fashion as to have few x-reactors Bacteria have different virulence mechanisms virulence genes are they unique to species?
13/09/2009

MDM Genes used in the design of microarray are; Housekeeping genes
16S rRNA, 16S-23S rRNA intergenic transcribed spacer
region (ITS), rpoB; RNA polymerase Beta Hsp60/groEL; Heat shock protein recA; Recombinase A gyrB; Gyrase Beta
13/09/2009

MDM Virulence genes
bacterial toxins adhesins important virulence factors
facilitate colonisation cause tissue damage
13/09/2009

Sample MDM: Pig zoonoses
The bacteria that will be detected will belong to the following genera
Campylobacter sp Staphylococcus sp Enterococcus sp Listeria sp Clostridium sp Salmonella sp Streptococcus sp Yersinia sp and others.
13/09/2009

Plan of the investigation1. Literature Search for pathogenic
organisms2. Identification in pathogens of
housekeeping genes virulence genes
3. Collection of all genes of interest in FASTA format
4. “Unique Sequence” search with OligoPicker2.3.2
5. “Unique Sequence” is checked for homology in BLAST (NCBI) and other BLAST engines 13/09/2009

Plan of the investigationaccording to species
ERIC PATRIC NMPDR Other BLAST engines particular to speciesbeing investigated
5. Choice of control sequences to be included in microarray
Negative and positive 6. Design of the MDM
13/09/2009

Plan of the investigation7. Choice reference strains to be used for
validation8. Purchase of reference strains9. Extraction of DNA from reference
strains10. Elaboration of Hybridization protocol:
Standard Operating Procedure for the MDM
11. Choice of software12. Hybridization of Reference strains13/09/2009

Plan of the investigation13. Analysis of Results ???14. Hybridization of field strains15. Analysis of Results ???16. Extraction of DNA from Test material17. Hybridization of Test material18. Analysis of Results ??? 19. Comparison of fluorescent profiles of
strains, HOW?
13/09/2009

Preliminary data Preliminary hybridization results are available at the BrIzsP-Db database
http: candida.bri.nrc,ca/Italy2/Microarray version V2.0 for strains of Listeria sp Campylobacter sp
Cluster analysis may be performed by using Four different methods Eight different algorithms
13/09/2009

Analysis of results Analysis of results is rather complex when 8096 fluorescent signals must be interpreted A interactive database of results was constructed which automatically transforms the fluorescent signals and performs cluster analysis with various mathematical algorithms
Organism Probes
Campylobacter 503
Yersinia 325
Salmonella 621
Listeria 142
Streptococcus 295
Clostridium 116
Others 22
Total 2024
13/09/2009

BrIzP-Db (database)
13/09/2009

Interpretation of data
13/09/2009

Plan of work for the next year
MIAME compliance Finish work with Reference Strains Application of Learning Algorithms to Data Hybridize to slides biological samples
13/09/2009

Timetable2008 2009 2010Sep-Dec Jan-Apr
May-Aug
Sep-Dec Jan-Apr
May-Nov
2nd set of farm samples1
Storm samples2
Species screening
Multilocus sequence typing
Survival studies
Thesis preparation 1One week in October; 2Three storms to be sampled13/09/2009

Hybridization Protocol
13/09/2009