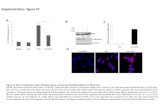
Figure S1 Figure S1. Modular structure of rice proteins encompassing the SPX domain. Different...
-
Upload
philip-cannon -
Category
Documents
-
view
218 -
download
2
Transcript of Figure S1 Figure S1. Modular structure of rice proteins encompassing the SPX domain. Different...

Figure S1
Figure S1. Modular structure of rice proteins encompassing the SPX domain. Different colors indicate the amino acid ranges of the domains. Purple, SPX domain; blue, EXS domain; yellow, MFS domain; red, RING-type Zinc finger domain; pink, no conserved domain.

Figure S2
Figure S2. Expression analysis of OsSPX-MFS1, OsSPX-MFS2 and OsSPX-MFS3 in leaves and roots.

Figure S3
Figure S3. Expression analysis of OsSPX-MFS1, OsSPX-MFS2, OsSPX-MFS3 and osa-miR827 under different Pi starvation treatments. Seeds were geminated and growth on nutrient solution with different Pi concentration for three weeks. Roots were sampled separately for analysis. All data are means of three biological replicates with error bars indicating SD. Expression of OsACTIN was used as the internal control. nd indicates non-detectable.

Figure S4
acctctctagttcattttgtgagagagctctacccagctccgctcccatattagatagaactgaaactgtttggttga
gcttcagctccaggagagatggagctagagctggagccgtgccaaacaggaccatctaggtactgacacgtgta G-boxgaaaatacttcccggagctttggttcttctgttccactccacccatcggcacaggcgggcgtctgcactctgggcctc
tggcgtgcggcgcccgcgcgtggtctggttgcgcagcgcgctgcgccgggtggaatccgttgcggcggcgtacac
cgtcacggcgtcacacactcacaccagcgccgcgcgccgcgggggacgtcagggactcgcgcgggccatgcag
gttgggctggctctcggcctctcgcctgtatctcaagggggtagcagcccattgcgtcggtagacctaggactcccc
ccgtgggccggtcacgtggagtcgtacgtggccctcgtcgatagcgccgcaggcctctctcaccacaagtccacg G-boxatgctgccccgaggatggttgtacacatggtagaacagtccgagaggggaacaattttgcctacaaattatccaca E-boxactttgcgagcgtttcatccgttccgaacagtccccttcctctgctgccccatcatcgctcatatctggattttttttttg
aaaggagtatatctggatatatataataccccttcgtactcgtaaagaaaattgtttaggacaatgtttaagtcaga
ccttgaaaatataaatcataaataactctcaaattgttgagtttgaaaatataaaaattatataaatagatttgtctt TATA box TATA boxgaaaaatattttcataaaagtatacatataatactttcaataaatatttttttatagaaacaataagtaaaagttgtat
tttagagaccgtgtcactgtccaaaatgactttctctacgagtaagagagagtattcctttttttttcatgactggaca W-box W-box accgcaaaatccacgcgttcgtccgctggtacaactcctccctgtcacggctaatttaccaagaccgttgatccaaa
tctacgctcatcatctccacctagcagaagaggcagtacagtcagccagtggcacagcgtcagcgtcactacaga
gtgtgcacgtacatacatgttgctgactgtctcctgcttaattacttgatgcttccatgtcgtcgtgcttctgcggggg W-boxacggtttattcctcgcgaatatgcctctaattcgctggcatcaacgtaccatcatcccttccgtgcacgcacctaccc P1BSgatccaaaatatatccttcgcttcaatacagcactagcagcttcatctcattcgccccatcttccagtcttccaggcgt
ccagctccagtttcttctcattgacaatatatccagattatgctctgggcttgctaattgctactagtagaatactagg
ggtggtcgtggtgtgcaggtgagattattgctacCCATGAACCTGTTTTGTTGCTGGTCAT
CTAGCTACCCGTGCATGCCTGGAGATTGGAGAATAATTGACGATGCA
GCAGTCGGCTTATTGGCTCTTGGGCACGCGTGGTTAGATGACCATCA
GCAAACAAGTTCGTGAG
Figure S4. Nucleotide sequence of the promoter and gene region of osa-miR827. The letters in capital indicates the precursor sequence of osa-miR827. The cis-elements of the promoter region were colored in red.

Figure S5
Figure S5. Expression analysis ofOsSPX-MFS1 (A), OsSPX-MFS2 (B), OsSPX-MFS3 (C) and Pri-miR827 (D) in osa-miR827 overexpression lines and msf1in the roots. Seeds were geminated and growth on nutrient solution with high Pi concentration (HP, 10 mg/L) and low Pi concentration (LP, 0.1 mg/L) for three weeks. Total RNA was extracted from the roots and used for qRT-PCR. All data are means of three biological replicates with error bars indicating SD. Expression of OsACTIN was used as the internal control. WT, wild type; Oe1/2/3, osa-miR827 overexpression lines; mfs1, OsSPX-MFS1 mutant line NC2634.

Primer Name Forward ( 5` → 3` ) Reverse ( 5` → 3` )OsSPX-MFS1 GCGATTCTTGGGTGTACTGT CGTGAAAGCAACGACAGGTTOsSPX-MFS2 CCTCCTGAATGTAACCCTG AGGAACTGTGTCAACTGCTTOsSPX-MFS3 CGATTGGTTCAGCATACAGA GTGCTGAGACGACATACTGGPre-miR827 TCTAGCTACCCGTGCATGCC GCGTGCCCAAGAGCCAATAAOsSPX1 GACCAGCTTCTACCATCAAACG AGTTCCTGCTGCTCCTCTGGOsIPS1 AAGGGCAGGGCACACTCCACATTATC ATTAGAGCAAGGACCGAAACACAAACOsPAP10 ATACTGGCAGCCGACGGATGA GAGGGAGCTGGAGCGGAGAAOsACP1 AGATGGAGGAGACGCTGCTG TCGCAATCCTCACACACACATOsPT2 CACAAACTTCCTCGGTATGCT GAAACCCCACAAATCCACAACOsPT3 AGACGGTGGTTCAACAGAG GCAAACCAAACTAACAAAATACCOsPT7 CTTGCTCGGCTTCCTCCT GCCATTATTTCATCATCCAGCOsPT8 CCTACTTGTGTTTGTCTATGTG GTGCCAAATTGCTGGTCTGTos17-3' ATTGTTAGGTTGCAAGTTAGTTAAGANC2634 GTTTTGCGGTTCTGCTTCTC TCCACCAAATCCTGACCAACOsSPX-MFS1:Pro ACTCCACAAGGATTGGCTCCG CTTGTCGTTCAATACCTGTATCmiR827-Oe GATCGGATCCGGGTGGTCGTGGTGTGCAGGT GATCGGATCCCAAATGAACCAGAGAATCGAC
Table S1. Sequences of the primer sets used in the study

Gene name Gene locus Protein length cDNA accession No. Total EST
OsSPX-MFS1 Loc_Os04g48390 696 AK066068 9
OsSPX-MFS2 Loc_Os02g45520 697 AK241359 18
OsSPX-MFS3 Loc_Os06g03860 698 AK243620 182
OsSPX-MFS4 Loc_Os09g34990 570 NF NF
Table S2. Information of OsSPX-MFS genes in rice