Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation
-
Upload
nishoanth-ramanathan -
Category
Education
-
view
660 -
download
0
description
Transcript of Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation

Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation
Mario Halic and Danesh Moazed
Nishoanth S Ramanathan
MS Molecular and Cell Biology
Presented by,

I. IntroductionII. GoalIII. Results and
ImplicationsIV. KeyV. Procedures involvedVI. Reference

Introduction
• Fission yeast A.K.A Schizosaccharomyces pombe• siRNAs and miRNAs Translation inhibition and RNAi
dsRNADICER
siRNA Argonuate
RITS or RDRC
Specific sequencesInactivation of target RNAsRNAi
trig
gers
gu
ides

Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
Contd…
• RITS complx = Argonuate + Tas3 + Chp1• Tas3 = GW motif protein linking Chp1 and Ago• Chp1 = chromodomain containing ptn that interacts
with H3K9 methylated nucleosomes• RITS associates with chromatin via
i. siRNA in Ago1 with nascent transcriptsii. Chp1 with H3K9 methylated nucleosomes
• This association leads to CLRC recruitment to swi6 and Chp2
• CLRC = Clr4 Rik1 Cul4 methyltransferase/Ub ligase Complx
• Swi6 and Chp2 = chromodomain ptns

Heterochromatin vs Euchromatin

Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
Contd…
• In addition to RITS, Fission yeast contains ARC.• ARC = Argonuate siRNA Chaperone Complex• ARC = Ago1 + Arb1 + Arb2 + duplex siRNA
• Silencing activity is amplified by RdRP• RdRP = RNA dependent RNA Polymerase• Fission yeast RdRP is Rdp1• RDRC = RNA dependent RNA Polymerase complex• RDRC = Rdp1 + Hrr1 + Cid12• Hrr1 = Dead box RNA helicase• Cid12 = Nucleotidyl transferase domain containing
ptn

Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
How siRNA generation and
Heterochromatin assembly
initiated???Models can explain HOW!
It is suggested that trigger siRNAs are produced from centromeric transcripts

Models for Biogenesis of intial siRNA from centromeric repeats
Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed

GOAL
EVIDENCE:
• Ago1 associated small RNAs target RDRC to centromeric transcripts.
• priRNAs act through argonaute in H3K9 methylation triggering siRNA amplification.
• siRNAs undergo processing at their 3’ ends which involves Cid12 and Cid14 nucleotransferases and trimming mediated by exosome.
Examine the mechanisms that mediate small RNA generation from the fission yeast centromeric repeat sequences.
Results suggests : A transcriptome surveillance mechanism based on the random association of small RNAs with Argonautes triggers RNA- mediated heterochromatin formation with DNA repeats.
Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed

RESULTS
Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
• Distinguish models• Determine siRNA levels
• Radioactive labelling of Ligation Oligont
• Small RNA, Lig Oligont and Bridge mixed, denatured and anealed with T4 DNA Ligase
• Excess Oligont is dephosphorylated
• Detected by PAGE

Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
Contd…
• Detect Centromeric small RNAs in WT and mutant cells by splinted ligation
• Diff amt of Ago asso. Small RNAs = Relative input• 50 fold more sensitive that N.Blot

Contd…
Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
• >100 fold reduction• dcr1r and rdp1r levels
are similar
No siRNAs are produced from dsRNA through sense-antisense pairing
• Presence of small RNAs in dcr1r
Existence of DICER independent small RNA generation

Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
Contd…
• Small RNAs in WT & Mutant cells plotted on cnt region of chromosome 1
• Cid12r 10 fold more siRNA present
Low levels of siRNA generated in a Cid12 Indepenfent fashion

Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
Contd…
• Rdp1 over expn restores the dg and dh siRNAs to WT levels

Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
Contd…
Essential role of cid12 in RNAi
• Silencing Assay by growth on complete selective (EMMC-Leu) and 5-Fluoroorotic acid (FOA) medium, respectively
• Rdp1 over expni. Rescues cid12 loss
of silencing phenotype of a imr1R::ura4+ reporter gene
ii. Fails to restore IRC3

Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
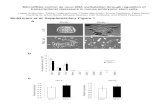
Concise RESULTS section -1
Cl4r cells dg siRNA
if10 folds
H3K9 Independent
H.C. IndependentDicer Dependent
RDRC Dependent
swi6r cells dh siRNA
dg siRNA
Reduced by more than 10 foldModest reduction ~2 fold
arb1r cellsarb2r cells Similar to swi6r cells
dg repeats
dh repeats
H.C. Independent siRNA generation
H.C. Dependent siRNA generation

Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
Contd…
• Chp1 subunit is not required for dg siRNA accumulation
• Tas3 is required
rules out a requirement for the full RITS complex in heterochromatin independent siRNA generation

Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
Contd…
• Ago1D580A has a loss of slicer activity
• dg RNAs produced in the lanes containing Ago1D580A are similar to dcr1r cells and less than that of the Cl4r cells
Heterochromatin independent dg RNAs amplification involves Ago mediated slicing or recognition of dg templates as in Model 2 and also implies the requirement of triggering dg small RNAs

Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
Contd…
• total number of small RNAs was similar in dcr1r and rdp1r cells
Sense and Antisense strands do not form high levels of dsRNA that can be processed by DICER.

Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
Concise RESULTS section -2
• High level of antisense txn is unique in S. pombe• Antisense small RNA per mRNA is negligible compare
to Antisense small RNA per dg/dh repeats• In addition the majority of priRNAs mapping to coding
regions showed a striking enrichment toward the 30 ends of annotated transcripts, often overlapping 30 untranslated sequences or regions just after transcription termination
Existence of a relationship between priRNA biogenesis at these regions and RNA processing events associated with transcription termination and also provide further support for the proposal that priRNAs originate from degradation products of single stranded transcripts

Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
Contd…
• PAZ/MID mutants are termed Ago1 A3
• Staining for all RNAs with Sybr Green showed that 21–24 nt small RNAs and 30–40 nt small RNAs were enriched in Ago1 from dcr1r cells
• Ago1A3 cells abolished H.C. dependent gene silencing of reporter gene.
Association of dcr1 independent small RNA with Ago1

Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
Concise RESULTS section -3
• The DICER independent small RNAs are termed as priRNAs (Primal RNAs)
• ago1r cells shown greater H3K9 methylation defect than dcr1r cells and rdp1r cells
• H3K9 methylation in ago1r cells were generally close to clr4r cells
RNAi independent H3K9 methylation is Ago1 dependent
• In dcr1r cells it is found that only priRNAs are present
priRNA promote low levels of H3K9 methylation

Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
Contd…
• To determine whether Ago1-dependent, but Dcr1-independent, H3K9 methylation required the PAZ and MID domains of Ago1, which were required for priRNA binding
• Ago1A3 present, H3K9 methylation was similar to Ago1r cells present, H3K9 methylation
Ago1 promote DICER indepencent methylation which requires small RNA binding domain.

Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
Concise RESULTS section -4• Ago1D580A has
i. Inactive slicerii. Dominant –ve effect on H.C. formationiii. Inhibits H3K9 methylationiv. Do not promote dcr1 independent methylationv. ds siRNA do not release passenger strand
• Centromeric siRNA associated with Ago1D580A had broad size from 21 – 27 nt
• Ago1 can bind larger precursors and then be trimmed to 22nt siRNAs after the release of the passenger strand which is a similar case as to priRNAs
• 3’ end mismatch of WT was similar to that of 3’end mismatch of the mutant suggesting that nucleotidyl activity is not required for large size

Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
Contd…
• priRNA mediated H3K9 meth requires RITS or not???• In ago1r cells, Chp1r cells and Tas3r cells are studied

Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
Contd…
• Chp1r, Tas3r-dcr1r & Tas3r cells shows higher methylation than Ago1r cells & equal to dcr1r cells
Ago1 promote H3K9 methylation indepencent of RITS

Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
Concise RESULTS section -5
Cell Type 1 Cell Type 2 priRNA level Inference
Ago1D580A dcr1r SIMILARSlicer based mechanism not involved
dcr1r cells Dcr1r rdp1r SIMILARRdp1 is not
involved
- eri+ cells Unaffected
priRNA are ss & generated
from ss precursors.eri is a ds specific ribonuclease.

Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
Concise RESULTS section -6• dh (–) strand transcript ∞ priRNA (–) strand levels• cid14 helps in the RNA degradation or processing of
RNA to exosome• degradation of small RNAs is at least partly mediated
by the exosome• Sequencing of Ago1 bound small RNAs from
Dis3Mutdcr1+ shows the similarity to WT but 3% of them had longer transcripts and from Dis3Mutdcr1r shows the presence of priRNAs but their levels are lower than dcr1r cells
An exosome may be involved

Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
MODEL

RESULTS – KEY POINTS
Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
• Detection of Ago1 bound small RNAs in RNAi and Heterochromatin Mutants
• High throughput sequencing reveals 3’ end processing of siRNAs by Nucleotidyl tranferases
• Heterochromatin independent siRNA generation from dg transcripts
• piRNA primarily target centromeric repeats and require PAZ/MID domain for Ago binding
• Role of piRNA in promoting H3K9 methylation• Ago1 bound siRNAs are trimmed after release of
passenger strand• Final Model for initiation of siRNA generation and
Hetrochromatin assembly

Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
PROCEDURES INVOLVED
• Protein Affinity Purification, RNA Synthesis, and Cid12 Activity Assays
• siRNA Purification and Detection by immunoaffinity purification of FLAG-Ago1 with an M2 affinity resin
• Strain Construction• Chromatin Immunoprecipitation• Statistical significance was calculated with the
Student’s t test• The ligated reaction was gel purified and used for
first-strand complementary DNA (cDNA) synthesis. cDNA was amplified with PCR
• Analysis by Maq (http://maq.sourceforge.net/) and/or Novoalign (http://www.novocraft.com/)

Dicer-Independent Primal RNAs Trigger RNAi and Heterochromatin Formation by Mario Halic and Danesh Moazed
References
• Aoki, K., Moriguchi, H., Yoshioka, T., Okawa, K., and Tabara, H. (2007). In vitro analyses of the production and activity of secondary small interfering RNAs in C. elegans. EMBO J. 26, 5007–5019.
• Aravin, A.A., Hannon, G.J., and Brennecke, J. (2007). The Piwi-piRNA pathway provides an adaptive defense in the transposon arms race. Science 318, 761–764.
• Baulcombe, D. (2004). RNA silencing in plants. Nature 431, 356–363.
• Bu¨ hler, M., Verdel, A., and Moazed, D. (2006). Tethering RITS to a nascent transcript initiates RNAi- and heterochromatin-dependent gene silencing Cell 125, 873–886.