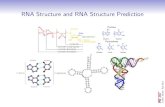
Class 5: RNA Structure Prediction
description
Transcript of Class 5: RNA Structure Prediction

.
Class 5:RNA Structure Prediction

RNA types
Messenger RNA (mRNA) Encodes protein sequences
Transfer RNA (tRNA) Adaptor between mRNA molecules and amino-acids
(protein building blocks) Ribosomal RNA (rRNA)
Part of the ribosome, a machine for translating mRNA to proteins
mi-RNA (micro-) Sn-RNA (small nuclear) RNA-I (interfering) Srp-RNA (Signal Recognition Particle)

Functions of RNAs
Information Transfer: mRNA
Codon -> Amino Acid adapter: tRNA
Enzymatic Reactions:
Other base pairing functions: ???
Structural:
Metabolic: ???
Regulatory: RNAi

RNA World Hypothesis
Before the “invention” of DNA and protein, early organisms relied on RNA for both genetic and enzymatic processes
DNA was a selective advantage because it greatly enhanced the fidelity of genetic replication
Proteins were a selective advantage because they make much more efficient enzymes
Remnants of the RNA world remain today in catalytic RNAs in ribosomes, polymereases and slicing molecules

Why is RNA structure important?
Messenger RNA is a linear, unstructured sequence, encoding an amino-acid sequence
Most non-coding RNA’s adopt 3D structures and catalyse bio-chemical reactions.
Predicting structure of a new RNA => information about its function

Terminology of RNA structure
RNA: a polymer of four different nucleotide subunits: adenine (A) , cytosine (C), guanine (G)and uracil (U)
Unlike DNA, RNA is a single stranded molecule folding intra-molecularly to form secondary structures.
RNA secondary structure = set of base pairings in the three dimensional structure of the molecule
G-C has 3 hydrogen bonds A-U has 2 hydrogen bonds Base pairs are almost always stacked onto other pairs,
creating stems.

Base Pairing in RNA
guanine cytosine
adenine uracil

Non-canonical pairs and pseudoknots
In addition to A-U and G-C pairs, non-canonical pairs also occur. Most common one is G-U pair.
G-U is thermodynamically favourable as Watson-Crick pairs (A-U, G-C) .
Base pairs almost always occur in nested fashion. Exception: pseudoknots.

Elements of RNA
secondary
structure

RNA Secondary Structure(more…)

AGCTACGGAGCGATCTCCGAGCTTTCGAGAAAGCCTCTATTAGC

RNA Tertiary Structure
•Do not obey “parantheses rule”

tRNA structure

Structure vs Sequence
Homologous RNA’s that have common secondary structure without sharing significant sequence similarity are important.
It is advantageous to search conserved secondary structure in addition to conserved sequence in databases.

Two Problems
1. RNA secondary structure for a single sequence. The dynamic programming algorithms –
Nussinov and Zuker, SCFG algorithms.
2. Analysis of multiple alignments of families of RNA’s.
Covariance Models – used for both multiple alignment and database searches.

Problem I: Structure Prediction
Input: An RNA sequence X
Output: Most likely secondary structure of X
Algorithms: Nussinov, CYK, MFOLD, …

Problem II: RNA family modeling
Input: A family for RNA sequence X1, …, XN sharing a common secondary structure
Aligned / Not aligned
Output: A probabilistic generative model representing the RNA family
Model: Covariance model