Chapter 6: Metabolism and Enzymes The sum total of an organism’s chemical reactions is called...
-
Upload
walter-lambert -
Category
Documents
-
view
227 -
download
2
Transcript of Chapter 6: Metabolism and Enzymes The sum total of an organism’s chemical reactions is called...



Chapter 6: Metabolism and Enzymes

The sum total of an organism’s chemical reactions is called metabolism.
The chemistry of life is organized into Metabolic Pathways

Catabolic pathways release energy by breaking down complex molecules to simpler compounds
Metabolic Pathways are made of:
ATP

Anabolic pathways consume energy to build complicated molecules from simpler compounds.
Metabolic Pathways are made of:
ATP

Catabolic Pathways and Anabolic Pathways act in tandem
Metabolic Pathways :

Fig. 6.1 The inset shows the first two steps in the catabolic pathway that breaks down glucose.

Chemical reactions can be classified as either exergonic or endergonic based on energy.
An exergonic reaction proceeds with a net release of energy.
Fig. 6.6a
C6H12O6 + 6O2 -> 6CO2 + 6H2O
CATABOLIC PATHWAY
Free energy of this reaction is negative (G)

An endergonic reaction is one that absorbs energy from its surroundings.– Endergonic reactions store energy
ANABOLIC PATHWAY
6CO2 + 6H2O -> C6H12O6 + 6O2
Free energy of this reaction is positive (G)

Exergonic Reactions and Endergonic Reactions are Coupled using ATP
ATP (adenosine triphosphate) is a type of nucleotide ATP has the nitrogenous base adenine, the sugar ribose, and a chain
of 3 phosphate groups
ATP: Adenine Triphosphate

Bonds between PO4 groups can be broken to release energy
This is a hydrolysis reaction
ATP: Adenosine Triphosphate

PO 4 released is tagged to a reactant
Reactant is phosphorylated and now able to undergo the chemical reaction
ATP can be regenerated
ATP: High NRG PO4 Bond Transfer

Energy FactsEnergy is used by the cell for: Mechanical Work
(movement), Transport (of macromolecules into and out of cells), and Chemical Work (drive endergonic reactions in anabolic pathways)
Plants transform light to chemical energy; they do not produce energy.

Activation energy: Energy needed by reactants to make the products (NRG Barrier)
Activation Energy is used to make transition state complexes whose bonds are strained. Then, products form from these transition state complxes by breaking and making new bonds.


Activation energy: Enzymes lower the Activation Energy needed for a reaction
Enzymes drive most chemical reactions in the body

Activation energy: Enzymes lower the Activation Energy needed for a reaction
Enzymes drive most chemical reactions in the body
Lactose ----------------------------- > Glucose + GalactoseLactase (lactaid pills)

ENZYMESEnzyme discovery to benefit homeland security
Enzymes are Proteins (names end in ase)

Enzymes:Are Substrate Specific
Enzyme names have 2 parts: First part – which substrate it acts
on/what product is formed Second part – what it does
Malate Dehydrogenase
Citrate synthase

Enzymes:Are Substrate Specific
1. Oxidoreductases: Oxidation – reduction; (dehydrogenase, reductase, oxidase)
2. Transferases: Transfer functional groups; (transferase; phosphorylase)
3. Hydrolases: hydrolytic cleavage of C-O, C-N, C-C bonds (phosphatase; protease)
4. Lyases – cleave bonds (decarboxylase)5. Isomerases - geometric or structural changes within a
molecule (epimerase or isomerase)6. Ligases - joining together of two molecules using ATP
(synthetase; ligase)
Skip details


Enzymes: Active SiteEnzymes act on Substrates (reactants) Active Site: is a pocket or groove on the surface
of the enzyme into which the substrate fits (often active siteis a nonpolar environment)
Substrate binds to active site by hydrogen bonding/weak forces to form an enzyme-substrate complex

Enzymes: Active Sites Active Site binding helps the reaction by:
•Substrates are placed in the correct orientation for the reaction.
•Puts stress on bonds that must be broken, making it easier to reach the transition state.
•R groups at the active site may create a conducive microenvironment for a specific reaction.
•Enzymes may even bind covalently to substrates in an intermediate step before returning to normal.

Induced Fit Model For Enzyme Action: Substrate binding to active site causes a
change in enzyme shape around active site The active site is molded into a tighter fit
around the substrates Substrates are held in close contact; EA is
lowered; products are formed and released Enzyme regains original shape; it is reused
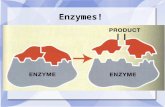
Enzymes: How do they work?

A single molecule of enzyme can catalyze 1000’s of reactions per second

Enzymes can catalyze both forward and backward reactions
Direction of a reaction depends on accumulation/removal of product
Enzymes: How do they work?
Malate Dehydrogenase

Substrate Concentration Temperature pH Cofactors Coenzymes Competitive Inhibitors Non-competitiveInhibitor Allosteric Regulation and Co-operativity Feedback Inhibition
Enzymes: Factors affecting their function (nine of them! - know these)

Enzymes: Factors affecting their function
The environment of the cell affects the structure of enzymes (proteins) at the secondary/tertiary/quarternary levelsThe effect is perceived as a change in reaction rate (rate of formation of product)

1) Substrate ConcentrationEnzymes: Factors affecting their function
Low [S]: an increase in substrate speeds binding to available active sites ([E] >>>> [S])
High [S]: Enzyme is SATURATED (all active sites are occupied) (([S] >>>> [E])
At High [S]: To increase reaction rate, increase [E]

2) TemperatureEnzymes: Factors affecting their function
Low Temp: insufficient collisions between active site and substratesHigh Temp: Enzyme can denature and the folding can unravel, damaging functionality
Optimal temp: Human enzymes – 37.5 oc

3) pH – determines state of acidic/basic functional ‘R’ groups on amino acids, and hydrogen bonding
Enzymes: Factors affecting their function
Low/High pH: changes the above interactions and alters folding of enzyme protein/active site
Optimal pH: Human enzymes – pH 6-8

4) Cofactors –inorganic helpers. Bind permanently or reversibly to enzyme. What’s the biochemical reason?
• Examples: zinc, iron, and copper.
Enzymes: Factors affecting their function
Photosystem I multienzyme complexes

5) Coenzymes – organic helpers (same idea as cofactors). Bind reversibly to enzymes - can be re-used.
Examples:vitamins.
Enzymes: Factors affecting their function

Inhibitors:6) Competitive Inhibitors: Active Site Directed
Inhibitors - bind to active site and inhibit binding of substrate
Enzymes: Factors affecting their function

Inhibitors:7) NonCompetitive Inhibitors: Bind to a
different site on enzyme called Allosteric Site - diminishes bindng of substrate to active site
Enzymes: Factors affecting their function
Minimata Bay – Japan (Mercury Poisoning)
Nerve Gas Used on Kurds by Iraq - 1993

8) Allosteric Regulation: Enzyme has several subunits - hangs around in “active” or “inactive” state
Effect of binding of regulator (inhibitor/activator) to one subunit is translated to other subunits (cooperativity)
Enzymes: Factors affecting their function

9) Feedback Inhibition - a metabolic pathway is turned off by its end product. End product becomes a inhibitor for a very early step enzyme. Very common!
Enzymes: Factors affecting their function

Rate of a Reaction Reaction Rate- the change in the concentration of a reactant or product with time. (M/s)
General equation for a reaction:– A → B– Reactant → Product
In order to monitor a reaction’s speed or rate, we can look at one of two things:– Decrease in [ reactant ]– Increase in [ product ]– Can be represented as:
rate = - Δ [A] / Δ t or
rate = Δ [B] / Δ t

Rate of a Reaction

Rate of a Reaction

Rate Calculations
How do we calculate the rate of a reaction?– We first need this information:
• Time (s) • [reactant]• Or [product]• [ ] means concentration• Decrease in [reactant]/time OR• Increase in [product]/time• Calculate the slope of your graph - X axis - time; Y axis - you
choose• (Y2-Y1)/(x2-x1)

Reaction rate calculationAppearance of Glucose in an Enzyme Catalysed Reaction with Lactaid Pills
Containing Lactase
0
5
10
15
20
25
30
35
40
45
0 10 30 60 120 180
Time in Seconds
Product Concentration (mM)
[Product] mM
Initial Stage of Reaction: [Substrate] is:[Enzyme] is:Number of Active Sites available to catalyze this reaction is:Calculation of Initial Reaction rate: Ri =( y2-y1)/(x2-x1) => Rise/Run

Reaction rate calculationAppearance of Glucose in an Enzyme Catalysed Reaction with Lactaid Pills
Containing Lactase
0
5
10
15
20
25
30
35
40
45
0 10 30 60 120 180
Time in Seconds
Product Concentration (mM)
[Product] mM
Final Stage of Reaction:[Substrate] is:[Enzyme] is:Number of Active Sites available to catalyze this reaction is:Calculation of Final Reaction rate: Rf =( y2-y1)/(x2-x1) = Maximum Velocity: Vmax