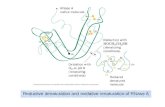
RNase mitochondrial RNA processing correctly cleaves a novel R ...
Biophysical and Population Genetic Models Predict the ... · Samples were separately ground in...
Transcript of Biophysical and Population Genetic Models Predict the ... · Samples were separately ground in...
Current Biology, Volume 26
Supplemental Information
Biophysical and Population Genetic Models Predict
the Presence of ``Phantom'' Stepping Stones
Connecting Mid-Atlantic Ridge Vent Ecosystems
Corinna Breusing, Arne Biastoch, Annika Drews, Anna Metaxas, Didier Jollivet, Robert C.Vrijenhoek, Till Bayer, Frank Melzner, Lizbeth Sayavedra, Jillian M. Petersen, NicoleDubilier, Markus B. Schilhabel, Philip Rosenstiel, and Thorsten B.H. Reusch
Supplemental Figure Legends
Figure S1. Related to Experimental Procedures (Larval dispersal modeling) and Supplemental Experimental
Procedures (Biophysical model). (A) Vertical profile of speed (mean and standard deviation, based on 5-daily data)
above Lucky Strike. (B) Stick-plot of bottom currents at the Lucky Strike vent field. The vector plot shows 5-day
average horizontal velocities (i.e., including directions) directly above the seafloor. Mean current maps of
VIKING20 at 1850 m depth (model level 29) for (C) northwestward and (D) southeastward bottom flow (identified
from B). Shaded is model bathymetry at horizontal model resolution and with intervals of vertical model resolution
Figure S2. Related to Experimental Procedures (Larval dispersal modeling) and Supplemental Experimental
Procedures (Biophysical model). Comparison of sea surface height (SSH) mean (A and C, in cm) and variance (B
and D, in cm²) between AVISO satellite (A, B) and VIKING20 model (C, D) data. SSH is a dynamic quantity for
upper-ocean current velocity. White stars indicate locations of the three modeled vent sites Menez Gwen, Lucky
Strike and Rainbow. The altimeter products were produced by Ssalto/Duacs and distributed by AVISO, with support
from Cnes (http://www.aviso.altimetry.fr/duacs/)
Supplemental Tables
Table S1. Related to Figure 1. Bathymodiolus sampling localities along the Mid-Atlantic Ridge
Locality Abbr. Latitude Longitude Depth [m] Cruise* Dive Samples
Menez Gwen MG‡,1
37°50.7'N 31°31.2'W 813–860 M:82/3
PP:BIOBAZ
690/27–761/6
N/A 50
Lucky Strike LS 37°17.0'N 32°15.0'W 1710 AT:03/3 3120 30
Rainbow RB 36°14.0'N 33°54.0'W 2251 AT:03/3 3121–3122 30
Broken Spur BS 29°10.0'N 43°10.0'W 3350 AT:03/3
AT:05/3
3125
3676 30
Snake Pit SP 23°22.0'N 44°56.0'W 3480 AT:03/3
AT:05/3
3129
3672–3674 30
Irina (Logatchev) IR‡,2
14°45.2'N 44°58.8'W 3020–3034 MSM:04/3 244/9–267/1
282/3 48
Quest (Logatchev) QS 14°45.2'N 44°58.8'W 3024–3047 MSM:04/3 267/7–271/5
313/2 30
Semenov SM 13°30.8'N 44°57.8'W 2432 PP:ODEMAR 541 40
Clueless CL‡,3
04°48.2'S 12°22.3'W 2992–2995 M:78/2
ATA
302/15
52/11 30
Lilliput LP‡,4
09°32.8'S 13°12.6'W 1489–1491 M:78/2 319/8–335/5 30
*Research vessels: M = Meteor, PP = Pourquoi Pas?, AT = Atlantis, MSM = Maria S. Merian, ATA = L'Atalante
‡5–6 samples from these sites were used for RNA sequencing to obtain reference transcriptomes for the putative
species: 1) B. azoricus, 2) B. puteoserpentis, 3) B. sp. 5°S, 4) B. sp. 9°S
Table S2. Related to Experimental Procedures (SNP marker design and DNA extraction and genotyping). Fluidigm
primer information (5' to 3'), putative gene functions and population allele frequencies of the 94 SNP markers
designed in this study. The comment column indicates which locus was used in which analyses (if only a subset of
SNPs was investigated). ASP1 = SNP allele detected with allele-specific primer 1, ASP2 = SNP allele detected with
allele-specific primer 2, SNP_SEQ = sequence of the amplified fragment containing the SNP, ASP1_SEQ =
sequence of allele-specific primer 1, ASP2_SEQ = sequence of allele-specific primer 2, LSP_SEQ = sequence of
locus-specific reverse primer, STA_SEQ = sequence of forward primer for specific target amplification, AMP_GC =
proportional GC content, REFSEQ = blast annotation in the RefSeq protein database, SWISSPROT = blast
annotation in the Swiss-Prot database, TREMBL = blast annotation in the TrEMBL database, FREQ_ASP1 =
frequency of SNP amplified with ASP1 in the respective population. Please refer to separate Excel sheet
Table S3. Related to Experimental Procedures (MSAT marker design and DNA extraction and genotyping). Primer
pools and sequences for the 9 microsatellites that were used in this study. c64270_g3_i1 was excluded from the
analysis of contemporary migration rates due to low polymorphism
Pool Locus Primer 5'-label Sequence Fragment [bp] Period
1
c27696_g2_i1 BMAR1F
BMAR1R
HEX
GCGTTATAACACCAAAACTCT
AACCACAGTCATTCACAAGG 127–159 4
c63755_g5_i1 BMAR4F
BMAR4R
HEX
TGGAGGCCAGGTTGTTTT
TGTTGCAAAGGGACATAAACAG 180–252 4
c60427_g1_i2 BMAR14F
BMAR14R
6-FAM
CCACATCAACACAAGTAGAAAGC
AACCTGTTGTGTTCACCGTC 183–210 3
2
c61565_g2_i3 BMAR5F
BMAR5R
6-FAM
GAAAAGTCAGCACCATGGCT
TGCTTTGTCCTGTAAACGCT 192–213 3
c55227_g3_i3 BMAR10F
BMAR10R
HEX
GACGTTTGACCAGAATAGGGG
GGGTTCTGGACAAATTCTCTGT 131–200 3
c43280_g1_i1 BMAR12F
BMAR12R
NED
GCTTCGCTTCTTCCTTTCTTCT
GGCAAGACAATAATTCCAGACGA 135–204 3
3
c42547_g1_i1 BMAR8F
BMAR8R
6-FAM
AAAAGCTGGGTTATATACTGCA
AGGGGTTGTATGACTAGGAAC 241–273 4
c50564_g1_i1 BMAR16F
BMAR16R
HEX
CACTCTTACAGGCTAGGATCCA
CGTCTTTCTTCCCCACAACA 246–267 3
c64270_g3_i1 BMAR21F
BMAR21R
NED
TGGTCGAATGAAGAGGAGCT
ACTTGTATCCATGGCCTCCT 159–189 3
Table S4. Related to Experimental Procedures (FST-outlier test). FST-outlier test using BayeScan. Only naturally
selected loci (consistent q < 0.1) are shown. Positive α values indicate positive selection, while negative ones imply
balancing or purifying selection. Prob = posterior probability of the model, log10(PO) = logarithm of Posterior Odds
to base 10 for the model, type of selection = positive (+) or negative/balancing (‒) selection between pairs of species.
No significant outliers within species were found
Locus Prob log10(PO) α FST Type of selection
c54079_g1_i1 0.8302 0.6891 0.9354 0.6406 + B. azoricus B. puteoserpentis
c44266_g1_i1 0.9754 1.5982 -1.7525 0.1975 ‒ B. azoricus B. puteoserpentis
c54250_g1_i1 0.8524 0.7615 -1.6606 0.2203 ‒ B. azoricus B. puteoserpentis
c14746_g1_i1 0.8396 0.7188 -1.6874 0.2187 ‒ B. azoricus B. puteoserpentis
c61080_g8_i1 0.8816 0.8718 1.0554 0.6608 + B. azoricus B. puteoserpentis
c62359_g8_i4 0.9078 0.9932 1.0744 0.6643 + B. azoricus B. puteoserpentis
c63335_g5_i2 0.9508 1.2860 -1.2576 0.2618 ‒ B. azoricus B. puteoserpentis
c34434_g1_i1 0.9848 1.8114 1.2899 0.7004 + B. azoricus B. puteoserpentis
c54783_g1_i2 0.9970 2.5215 2.0027 0.7980 + B. sp. 5°S B. puteoserpentis
c35009_g1_i1 1.0000 1000.0000 -2.7882 0.0936 ‒ B. azoricus all other species
c63790_g3_i1 0.9998 3.6988 -1.9354 0.1708 ‒ B. azoricus B. puteoserpentis
c58708_g7_i1 0.9996 3.3977 1.7073 0.7621 + B. azoricus B. puteoserpentis
c42320_g1_i1 0.8720 0.8332 -1.1034 0.2867 ‒ B. azoricus B. puteoserpentis
c60277_g2_i2 0.9336 1.1479 -1.1985 0.2708 ‒ B. azoricus B. puteoserpentis
c42562_g1_i1 0.9732 1.5600 1.3626 0.7110 + B. azoricus B. puteoserpentis
Table S5. Related to Figure 3. Results of the BayesAss analysis for the pooled data set using 44 neutral molecular markers. The table shows the mean
immigration rates (fraction of individuals that are derived from source per generation) ± 95% confidence interval. Numbers in bold denote rates that were outside
the 95% confidence interval for uninformative data. NMAR = Menez Gwen, Lucky Strike, Rainbow; BS = Broken Spur; MMAR = Snake Pit, Irina, Quest,
Semenov; CL = Clueless; LP = Lilliput
SOURCE
DE
ST
INA
TIO
N
NMAR BS MMAR CL LP
NMAR 0.9884 (0.9772; 0.9996) 0.0029 (-0.0028; 0.0086) 0.0029 (-0.0028; 0.0086) 0.0029 (-0.0028; 0.0086) 0.0029 (-0.0028; 0.0086)
BS 0.0259 (-0.0029; 0.0547) 0.7084 (0.6672; 0.7496) 0.2488 (0.1957; 0.3019) 0.0085 (-0.0080; 0.0250) 0.0085 (-0.0080; 0.0250)
MMAR 0.0022 (-0.0021; 0.0065) 0.0024 (-0.0021; 0.0069) 0.9894 (0.9800; 0.9988) 0.0039 (-0.0020; 0.0098) 0.0022 (-0.0021; 0.0065)
CL 0.0104 (-0.0092; 0.0300) 0.0104 (-0.0092; 0.0300) 0.0104 (-0.0092; 0.0300) 0.9532 (0.9115; 0.9949) 0.0156 (-0.0120; 0.0432)
LP 0.0083 (-0.0076; 0.0242) 0.0083 (-0.0076; 0.0242) 0.0084 (-0.0075; 0.0243) 0.0090 (-0.0081; 0.0261) 0.9660 (0.9350; 0.9970)
Table S6. Related to Experimental Procedures (De novo transcriptome assembly). Basic assembly statistics for the
raw and filtered de novo transcriptomes of the four Bathymodiolus (sub-)species. Contigs = number of all contigs
including splice variants. Loci = number of different "genes". CEGs = 248 low copy core genes that are ultra-
conserved in eukaryotes and are used for assessing the completeness and quality of assemblies, ORFs = unique
contigs with open reading frame
B. azoricus B. puteoserpentis B. sp. 5°S B. sp. 9°S
Raw
Contigs 173300 177587 225279 345535
Loci 122630 124196 158654 293714
Median contig length [bp] 439 442 412 279
Mean contig length [bp] 882.22 887.30 826.36 464.49
Assembled bases 152889522 157572715 186162113 160498529
GC content [%] 34.42 34.26 34.18 33.87
Filtered
Contigs 108448 134384 132632 319766
Loci 71371 91462 84326 274115
Median contig length [bp] 616 515 589 286
Mean contig length [bp] 1081.89 970.14 1037.03 457.37
Assembled bases 117328861 130370673 137543597 146252403
GC content [%] 34.77 34.44 34.54 33.74
CEGs [%]
Partial 100 99.60 100 95.56
Complete 99.19 97.58 97.18 78.23
Proteins
BLAST 36946 35870 39906 36863
Swiss-Prot 23707 24052 26581 22531
TrEMBL 36562 35528 39496 36316
RefSeq 32360 31415 34830 31434
ORFs 49268 49441 55137 48776
Supplemental Experimental Procedures
RNA extraction and high-throughput transcriptome sequencing
Samples were separately ground in liquid nitrogen and incubated at 4°C in RNAlater overnight to prepare samples
for RNA purification. This step has improved RNA isolation efficiency from mussel tissues in our lab. RNA was
subsequently isolated using a modified protocol of the RNeasy Mini Kit (Qiagen, Germany) for spin technology. To
ensure complete tissue homogenization, we prolonged the incubation time in RLT-β-mercaptoethanol buffer to 10
min and disrupted the samples with VWR VDI 12 and QIAshredder homogenizers (Qiagen, Germany) before adding
the lysates to the extraction columns. Prior to RNA elution, samples were put under a fumehood for 3 min to let the
remaining ethanol evaporate and then incubated for 10 min in RNAse free water (elution solution) on ice to improve
diffusion into the column membrane. Potential DNA contaminations were removed with the DNA free DNAse
Treatment & Removal Kit (Invitrogen/Ambion, Germany). RNA integrity and concentration were measured on the
Experion Automated Electrophoresis Station using the Experion RNA StdSens Analysis Kit (Bio-Rad, Germany)
before submitting samples to the Institute of Clinical Molecular Biology (Kiel, Germany) for 2×101 bp paired-end
RNA sequencing on an Illumina HiSeq2000 machine (Illumina, USA). 45 indexed sequencing libraries were
constructed from poly(A)-selected RNA with the TruSeq RNA Sample Preparation Kit v2 (350‒550 bp insert size),
while cluster generation was performed with the TruSeq PE Cluster Kit v3-cBot-HS (Illumina, USA). As the six LP
samples were needed for further RNA expression analyses of the bacterial endosymbionts [S1], RNA-Seq libraries
for these samples were not poly(A)-enriched and three of them were sequenced in a strand-specific way. For
ensuring sufficient read coverage [S2], six unstranded libraries were multiplexed on each lane of an Illumina
flowcell, whereas the three stranded LP libraries were sequenced on a separate lane.
De novo transcriptome assembly
Raw reads from multiplexed paired-end sequencing were first quality checked with FASTQC v0.10.1
(http://www.bioinformatics.babraham.ac.uk/projects/fastqc/) and then processed with FASTQ_ILLUMINA_FILTER
v0.1 (http://cancan.cshl.edu/labmembers/gordon/fastq_illumina_ filter/) to remove Y-tagged reads that failed to pass
the internal quality filter of the Casava 1.8+ pipeline. As transcriptome accuracy was more important than
completeness for our purpose, we opted for a stringent trimming strategy [S3, S4]. Adapter and read clipping was
performed with Flexbar v2.4 [S5], allowing for adapter removal in any part of the read, a minimum match of 10 bp
between adapter and sequence and not more than one mismatch or gap per 10 bases overlap. The custom adapter file
contained all Illumina oligonucleotide sequences used by the TruSeq Kits and their reverse complements. To account
for an increased error rate near the 5' end [S6] the first 15 bp of each read were removed (as suggested by the
FASTQC output), while the 3' end was trimmed until a minimum base quality of 20 (Illumina 1.8+ phred score
encoding) was reached. Reads shorter than 50 bp after processing or those with more than 5 uncalled bases were
discarded. Ribosomal sequences were removed from non-poly(A)-selected libraries with riboPicker v0.4.3 [S7],
using the following settings: -i 95 -c 90 -l 45 -dbs rrnadb. Filtered sequences were subsequently aligned against
unpublished symbiont genomes from B. azoricus (MG) and B. sp. 9°S (LP) via Bowtie2 v2.1.0 [S8] to separate the
bacterial reads (parameters: -I 0 -X 600 --very-sensitive). All paired-end sequences that did not match concordantly
were kept for assembly. To obtain comprehensive reference transcriptomes, we merged reads from all individuals
and tissues per (sub-)species. De novo assemblies were done with Trinity r20140413 [S9] using the PasaFly
algorithm with jaccard-clip option to reduce creation of chimeric transcripts. An in silico normalization to 30–50X
coverage was performed with an allowed kmer coverage deviation of 100% to decrease computational resources
(exception: LP reads were not normalized due to lower coverage). The group pairs distance was set to 600 bp, while
the minimum contig length was left at its default of 200 bp. Potential assembly artifacts with FPKM values < 1 were
removed with Trinity's support scripts "align_and_estimate_ abundance.pl" and "filter_fasta_by_rsem_values.pl",
choosing Bowtie2 v2.1.0 and RSEM v1.2.12 [S10] as read mapping and expression analysis software, respectively
(parameters: --no-mixed --no-discordant --gbar 99999999 --dpad 0 --very-sensitive). To avoid exclusion of real, but
lowly expressed transcripts, we re-integrated contigs from the discarded data set into the cleaned assembly, if they
had a significant BLASTx hit in the TrEMBL, Swiss-Prot or RefSeq protein databases (e-value threshold: 1e-20;
releases from April 2014) or contained an ORF of at least 100 aa based on maximum likelihood scores and Pfam AB
homologies as determined by TransDecoder rel16JAN2014 (https://transdecoder.github.io/). The final assembly was
clustered with an identity threshold of 99% with CD-HIT-EST v4.6.1 [S11], using the additional settings: -n 9 -r 1 -T
0 -M 50000 -g 1 -aS 0.99 -uS 0.01. This approach seemed to produce optimal results for our data sets in terms of
assembly quality and completeness as evaluated by the CEGMA pipeline ([S12]; Table S6).
SNP marker design
The B. azoricus transcriptome was chosen as a reference for inter- and intra-specific SNP detection. Cleaned tissue-
specific paired-end reads of each individual were first aligned with Bowtie2 v2.1.0 in the "very-sensitive" mode.
Subsequently, Picard tools v1.93 (http://picard.sourceforge.net) was used to mark PCR duplicates and merge the
tissue-specific BAM files for each specimen. To prevent base mismatch due to misalignment in indel regions, reads
were locally realigned with the IndelRealigner suite in GATK v3.0.0 [S13]. SNP calling was performed using the
mpileup2snp command in VarScan v2.3.7 [S14, S15]. MPILEUP files were created from each realigned BAM file
with SAMtools v1.1 [S16], applying the following settings: -C 50 -E -d 700 -Q 20 -q 10. SNPs were called, if the
respective position had a minimum read depth and base quality of 30, if each variant was supported with at least 15
reads and if the variant allele had a minimum frequency of 5%. The p-value threshold for variant calling was set to
0.05, while the minimum frequency for homozygote calls was fixed at 0.80. Variants with more than 90% support on
one strand were ignored. Raw SNP outputs for each individual were checked for false positive calls with VarScan's
"fpfilter.pl" script and all unique variants that passed the filter were saved in a list. The SNP calling procedure was
then repeated on the whole data set containing all individuals and the consensus between this approach and the pass
list was taken as the true SNP set. This set was further filtered with Reads2SNP v2.0 [S17, S18] to remove potential
paralogous variants (parameters: -min 15 -th1 0.95 -par 1 -th2 0.05 -nbth 16 -aeb -tol -spa -fis 0.0 -opt newton -rlg 50
-bqt 30 -rgt 10 -rleading 10 -rtrailing 20). Putative species-diagnostic SNPs were identified with the vcf-contrast
command in VCFtools v0.1.12a [S19] and based on the NOVELAL description in the resulting VCF files. The final
marker design only focused on bi-allelic SNPs that had a BLAST annotation, which yielded 80160 SNP variants. A
pre-selection of likely diagnostic and undiagnostic contigs containing these variants was blasted against a draft
genome of B. azoricus (T. Takeshi, unpublished dataset, Bathymodiolus genome JST project, coord. N. Satoh and A.
Tanguy) that had been made accessible to validate the uniqueness of the SNP regions and define exon-intron
boundaries for primer design. Cross-species amplifiability was evaluated based on the presence of conserved regions
using IGV v2.3 [S20]. Primers were designed in the Fluidigm D3 system (San Francisco, USA; Table S2).
MSAT marker design
MSATs were identified in the B. azoricus de novo transcriptome using TRF v4.07b [S21]. Weights for match,
mismatch and indels were set to 2, 7 and 7, respectively, while detection parameters were left at their defaults. All
detected MSATs with period sizes > 4 were discarded. To ensure amplifiability across species, we compared the
MSAT contigs to all other Bathymodiolus transcriptomes using NCBI BLAST v2.2.29+ [S22] with an e-value
threshold of 1e-20 and kept only those that had a significant hit in at least 2 out of 4 transcriptomes. This approach
yielded 83 potential MSAT candidates. We used the Geneious alignment algorithm in Geneious v8.1.3
(http://www.geneious.com; [S23]) to multiple align MSAT contigs and search for conserved regions within 150 bp
upstream and downstream of the repeat motif. Primer design was done in Primer3Web v4.0.0 [S24] with default
settings except that the minimum annealing temperature was lowered to 54°C and the maximum temperature
difference between forward and reverse primer was set to 1.5°C. Positions that contained SNPs or indels or were
outside the 150 bp flanking regions were excluded. Under these conditions primers for 21 gene regions could be
designed. Primer pairs were preliminarily triplexed into seven pools based on expected fragment lengths and
annealing temperatures. According to these theoretical groupings, unlabelled reverse primers and 5'-fluorescent
labeled forward primers (6-FAM, HEX or NED) were ordered at Metabion (Planegg, Germany) and Applied
Biosystems (Warrington, Cheshire, UK). After testing primers in single reactions and checking for polymorphism in
about 24 individuals per species, we kept 9 primer pairs, which were arranged in three pools (Table S3).
DNA extraction
DNA was extracted with the DNeasy Blood & Tissue Plate Kit (Qiagen, Germany) according to manufacturer's
suggestions with one modification. To increase DNA yield we conducted a second elution step with the previously
isolated DNA. DNA concentration and purity was measured on a NanoDrop ND-1000 spectrophotometer (Peqlab,
Germany).
SNP genotyping
SNPs were genotyped following the Fluidigm 96×96 SNP Type Genotyping protocol with optimized volumes of the
used reagents. Prior to end-point fluorescence detection on the Biomark HD system (Fluidigm, USA) we performed
a pre-amplification step with 16 cycles to make sure that enough DNA was available for each reaction. In each chip
run we included two negative controls to enable data normalization. SNP genotypes were called in the Fluidigm SNP
Genotyping Analysis software using the recommended SNP Type Normalization method. All genotype calls were
inspected by eye and manually adjusted, if necessary.
MSAT genotyping
MSATs were amplified in 10 µl triplex reactions containing 2–4 pmol of each primer, 5 µl Multiplex PCR Master
Mix (Qiagen, Germany), 1 µl template DNA and a variable amount of HPLC-H2O. Hot start at 95°C for 15 min was
followed by 26 cycles of 94°C for 30 s, 56°C for 30 s and 72°C for 1 min on Veriti Thermal Cyclers (Applied
Biosystems, Germany). The final extension was done at 72°C for 20 min. 1 µl of each PCR product was mixed with
8.75 µl Hi-Di formamide and 0.25 µl GeneScan 350 ROX Size Standard (Applied Biosystems, Germany) and then
denatured for 2 min at 95°C. Capillary electrophoresis was run on an ABI 3130xl Genetic Analyzer (Applied
Biosystems, Germany). MSAT genotypes were read with GeneMarker v1.91 [S25] using default settings in the run
wizard. Size standards for each sample were checked for correctness and adjusted by hand, if necessary. Bins were
constructed for each MSAT considering period size, peak frequency and height.
Analysis of mitochondrial ND4
Mitochondrial ND4 was amplified in a 15 µl PCR reaction using universal primers [S26, S27] and the DreamTaq
DNA Polymerase (Fermentas, Germany) as recommended by the manufacturer. PCRs were run following the
thermal cycling protocol with 28 cycles and an annealing temperature of 55°C. PCR products were submitted to the
Institute of Clinical Molecular Biology (Kiel, Germany) for bi-directional Sanger sequencing on an ABI 3730xl
Genetic Analyzer (Applied Biosystems, Germany). Sequence analysis and haplotype encoding was done as described
previously [S28].
Genetic structure and differentiation
STRUCTURE v2.3.4 [S29, S30] was used to determine the degree of genetic subdivision along the MAR. The
admixture model with correlated allele frequencies was iterated 107 times after a burnin period of 10
6. To identify the
number of genetic groups K (112), we corrected the posterior probabilities according to Evanno et al. [S31] and
investigated all bar plots by eye as suggested by Meirmans [S32]. STRUCTURE plots were converted to vector
graphics using "bar_plotter.rb" (http://evolution.unibas.ch/salzburger/software.htm). Pairwise FST values between all
sampled localities were computed in Arlequin v3.5.1.2 [S33] using a non-parametric permutation procedure (10000
replications). Post-hoc corrections were done with the Benjamini-Yekutieli False Discovery Rate method [S34].
FST-outlier test
We used BayeScan v2.1 [S35] to identify selectively adaptive FST-outlier loci that would bias the estimation of
contemporary migration rates. To investigate inter- and intra-specific patterns of selection and assess robustness of
the estimates the analysis was run with (1) the combined data set of all sampled sites, (2) data subsets including
populations from only two of the previously described species (for all pairwise comparisons), and (3) data subsets
including populations from only one species (if multiple sites had been sampled). Outliers were considered as
significant, if they had a q value < 0.1 and were identified in both the combined data set (1) and at least one of the
between-species (2) or within-species (3) data subsets. Parameter settings for each run followed suggestions in the
manual.
Biophysical model
We used the particle tracking tool Ariane [S36] with velocity data from the North Atlantic OGCM NEMO [S37] to
model dispersal between the three known vent sites MG, LS and RB. The configuration contains an eddy-resolving,
1/20° grid of the (sub-)polar North Atlantic region, VIKING20 (~30°N to 85°N; [S38, S39]), which is hosted within
the global 1/4° ORCA025 grid [S40] by using a two-way nesting approach [S41]. In the region of interest the
horizontal resolution of the grid is about 4.4 km. In the vertical, 46 depth layers are specified, the spacing of which
increases from 6 m at the surface to 250 m in the deep ocean. Characteristic layer thicknesses at the depths of the
simulated vents were between 134 m and 237 m. The seafloor of the model is defined by the ETOPO2 bathymetric
database [S42], where a partial-cell representation is used for the bottom topography [S40]. To simulate vertical
mixing a 1.5-level turbulent kinetic energy scheme is applied [S43], while discrete bi-Laplacian and isoneutral
Laplacian operators are used to model viscosity and diffusion, respectively. The model is initialized with
climatological salinity and temperature fields from Levitus et al. [S44]. Following a 30-year spin-up, simulations are
forced with atmospheric fluxes based on the CORE2 hindcast [S45, S46] with 6-hourly (wind speed, humidity, and
atmospheric temperature), daily (short- and long-wave radiation), and monthly (rain and snow) resolutions, and an
interannual variability for the time period 1948–2007. Simulated velocities are stored as 5-daily three-dimensional
averages. The VIKING20 model itself has been rigorously validated in previous studies [S39, S47] and has been
used for dispersal studies in the upper ocean [S48]. To demonstrate the reliability of our model simulations in the
area of interest, we compared sea surface height for the investigated years between AVISO satellite data and
VIKING20. Both mean and variance fields show that the model simulates the detailed current structure in the
subtropical gyre with sufficient detail, including the Azores Current seen as a variance maximum at 34°N (Figure
S2). In addition, we checked for the presence of small scale current variation around the model vent locations. It is
clear that the depths and specific bathymetric settings of the individual vents are crucial. For example, LS is deeply
located in the rift valley, with connections to the northern and southern flank of the MAR (Figure S1). As a result,
currents and its associated variability are bottom-intensified. The bathymetric setting causes a bi-directional
behavior, with current reversal on short timescales which matches the characteristics of observations [S49]. Coupling
between deep and surface flow is most likely indirect and appeared to be a complex result of oceanographic
conditions northwest and southeast of the MAR. A more thorough investigation of the dynamics of deep flow was
beyond the scope of this study. The most southern vent, RB, might occasionally be enveloped by the Azores Current.
In some years its mesoscale variability seems to be connected to the deep flow and might influence retention rates of
larval dispersal. However, we see no obvious connection between surface conditions and deep flow.
Calculation of particle starting positions and dispersal probabilities
At the East Pacific Rise, larvae can be entrained in rising hydrothermal plumes, which results in an inverse
relationship between larval abundance and distance from the seafloor [S50]. The observations for the bivalve
Bathymodiolus thermophilus (Figure 3 in [S50]; on-vent pattern represented by black bars) were best described by a
power function [y = 0.1619x0.377
(R² = 0.79), where y denotes the number of larvae per m³ at height x in m above
bottom]. In our study, release positions of larvae were within 400 m above the seafloorthe maximum height of
hydrothermal plumes in the Atlantic [S51]. After release, larvae were allowed to drift in any depth with the three-
dimensional ocean currents, while no assumptions about mortality were made at any time of the simulations. Larval
dispersal probabilities were computed with the FORTRAN script "tcdfprob.f90" [S52] using a horizontal resolution
of 0.1° and a vertical resolution of 10 m.
Supplemental References
S1. Sayavedra, L., Kleiner, M., Ponnudurai, R., Wetzel, S., Pelletier, E., Barbe, V., Satoh, N., Shoguchi, E., Fink,
D., Breusing, C., et al. (2015). Abundant toxin-related genes in the genomes of beneficial symbionts from
deep-sea hydrothermal vent mussels. eLife 4, e07966.
S2. Francis, W.R., Christianson, L.M., Kiko, R., Powers, M.L., Shaner, N.C., and Haddock, S.H.D. (2013). A
comparison across non-model animals suggests an optimal sequencing depth for de novo transcriptome
assembly. BMC Genomics 14, 167.
S3. Del Fabbro, C., Scalabrin, S., Morgante, M., and Giorgi, F.M. (2013). An extensive evaluation of read
trimming effects on Illumina NGS data analysis. PLOS ONE 8, e85024.
S4. MacManes, M.D. (2014). On the optimal trimming of high-throughput mRNA sequence data. Front. Genet. 5,
13.
S5. Dodt, M., Roehr, J.T., Ahmed, R., and Dieterich, C. (2012). FLEXBAR‒Flexible barcode and adapter
processing for next-generation sequencing platforms. Biology 1, 895–905.
S6. Van Gurp, T.P., McIntyre, L.M., and Verhoeven, K.J.F. (2013). Consistent errors in first strand cDNA due to
random hexamer mispriming. PLOS ONE 8, e85583.
S7. Schmieder, R., Lim, Y.W., and Edwards, R. (2012). Identification and removal of ribosomal RNA sequences
from metatranscriptomes. Bioinformatics 28, 433–435.
S8. Langmead, B., and Salzberg, S.L. (2012). Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–
359.
S9. Grabherr, M.G., Haas, B.J., Yassour, M., Levin, J.Z., Thompson, D.A., Amit, I., Adiconis, X., Fan, L.,
Raychowdhury, R., Zeng, Q., et al. (2011). Full-length transcriptome assembly from RNA-Seq data without a
reference genome. Nat. Biotechnol. 29, 644–652.
S10. Li, B., and Dewey, C.N. (2011). RSEM: accurate transcript quantification from RNA-Seq data with or
without a reference genome. BMC Bioinformatics 12, 323.
S11. Li, W., and Godzik, A. (2006). Cd-hit: a fast program for clustering and comparing large sets of protein or
nucleotide sequences. Bioinformatics 22, 1658–1659.
S12. Parra, G., Bradnam, K., and Korf, I. (2007). CEGMA: a pipeline to accurately annotate core genes in
eukaryotic genomes. Bioinformatics 23, 1061–1067.
S13. DePristo, M.A., Banks, E., Poplin, R., Garimella, K.V., Maguire, J.R., Hartl, C., Philippakis, A.A., del Angel,
G., Rivas, M.A., Hanna, M., et al. (2011). A framework for variation discovery and genotyping using next-
generation DNA sequencing data. Nat. Genet. 43, 491–498.
S14. Koboldt, D.C., Chen, K., Wylie, T., Larson, D.E., McLellan, M.D., Mardis, E.R., Weinstock, G.M., Wilson,
R.K., and Ding, L. (2009). VarScan: variant detection in massively parallel sequencing of individual and
pooled samples. Bioinformatics 25, 2283–2285.
S15. Koboldt, D.C., Zhang, Q., Larson, D.E., Shen, D., McLellan, M.D., Lin, L., Miller, C.A., Mardis, E.R., Ding,
L., and Wilson, R.K. (2012). VarScan 2: somatic mutation and copy number alteration discovery in cancer by
exome sequencing. Genome Res. 22, 568–576.
S16. Li, H., Handsaker, B., Wysoker, A., Fennell, T., Ruan, J., Homer, N., Marth, G., Abecasis, G., Durbin, R.,
and 1000 Genome Project Data Processing Subgroup (2009). The Sequence Alignment/Map format and
SAMtools. Bioinformatics 25, 2078–2079.
S17. Tsagkogeorga, G., Cahais, V., and Galtier, N. (2012). The population genomics of a fast evolver: high levels
of diversity, functional constraint, and molecular adaptation in the tunicate Ciona intestinalis. Genome Biol.
Evol. 4, 740–749.
S18. Gayral, P., Melo-Ferreira, J., Glémin, S., Bierne, N., Carneiro, M., Nabholz, B., Lourenco, J.M., Alves, P.C.,
Ballenghien, M., Faivre, N., et al. (2013). Reference-free population genomics from next-generation
transcriptome data and the vertebrate-invertebrate gap. PLOS Genet. 9, e1003457.
S19. Danecek, P., Auton, A., Abecasis, G., Albers, C.A., Banks, E., DePristo, M.A., Handsaker, R.E., Lunter, G.,
Marth, G.T., Sherry, S.T., et al. (2011). The variant call format and VCFtools. Bioinformatics 27, 2156–
2158.
S20. Robinson, J.T., Thorvaldsdóttir, H., Winckler, W., Guttman, M., Lander, E.S., Getz, G., and Mesirov, J.P.
(2011). Integrative genomics viewer. Nat. Biotechnol. 29, 24–26.
S21. Benson, G. (1999). Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 27,
573–580.
S22. Altschul, S.F., Gish, W., Miller, W., Myers, E.W., and Lipman, D.J. (1990). Basic local alignment search
tool. J. Mol. Biol. 215, 403–410.
S23. Kearse, M., Moir, R., Wilson, A., Stones-Havas, S., Cheung, M., Sturrock, S., Buxton, S., Cooper, A.,
Markowitz, S., Duran, C., et al. (2012). Geneious Basic: an integrated and extendable desktop software
platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649.
S24. Untergasser, A., Cutcutache, I., Koressaar, T., Ye, J., Faircloth, B.C., Remm, M., and Rozen, S.G. (2012).
Primer3‒new capabilities and interfaces. Nucleic Acids Res. 40, e115.
S25. Hulce, D., Li, X., Snyder-Leiby, T., and Liu, C.S.J. (2011). GeneMarker® Genotyping Software: tools to
increase the statistical power of DNA fragment analysis. J. Biomol. Tech. 22 (Suppl), S35–S36.
S26. Arévalo, E., Davis, S.K., and Sites, J.W. (1994). Mitochondrial DNA sequence divergence and phylogenetic
relationships among eight chromosome races of the Sceloporus grammicus complex (Phrynosomatidae) in
central Mexico. Syst. Biol. 43, 387–418.
S27. Bielawski, J.P., and Gold, J.R. (1996). Unequal synonymous substitution rates within and between two
protein-coding mitochondrial genes. Mol. Biol. Evol. 13, 889–892.
S28. Breusing, C., Johnson, S.B., Tunnicliffe, V., and Vrijenhoek, R.C. (2015). Population structure and
connectivity in Indo-Pacific deep-sea mussels of the Bathymodiolus septemdierum complex. Conserv. Genet.
16, 1415–1430.
S29. Pritchard, J.K., Stephens, M., and Donnelly, P. (2000). Inference of population structure using multilocus
genotype data. Genetics 155, 945–959.
S30. Falush, D., Stephens, M., and Pritchard, J.K. (2003). Inference of population structure using multilocus
genotype data: linked loci and correlated allele frequencies. Genetics 164, 1567–1587.
S31. Evanno, G., Regnaut, S., and Goudet, J. (2005). Detecting the number of clusters of individuals using the
software STRUCTURE: a simulation study. Mol. Ecol. 14, 2611–2620.
S32. Meirmans, P.G. (2015). Seven common mistakes in population genetics and how to avoid them. Mol. Ecol.
24, 3223–3231.
S33. Excoffier, L., and Lischer, H.E.L. (2010). Arlequin suite ver 3.5: a new series of programs to perform
population genetics analyses under Linux and Windows. Mol. Ecol. Resour. 10, 564–567.
S34. Benjamini, Y., and Yekutieli, D. (2001). The control of the false discovery rate in multiple testing under
dependency. Ann. Stat. 29, 1165–1188.
S35. Foll, M., and Gaggiotti, O. (2008). A genome-scan method to identify selected loci appropriate for both
dominant and codominant markers: a Bayesian perspective. Genetics 180, 977–993.
S36. Blanke, B., Arhan, M., Madec, G., and Roche, S. (1999). Warm water paths in the equatorial Atlantic as
diagnosed with a general circulation model. J. Phys. Oceanogr. 29, 2753–2768.
S37. Madec, G. (2008). NEMO ocean engine. Technical Report 27. Note du Pôle de modélisation, (France: Institut
Pierre-Simon Laplace), ISSN No 1288–1619.
S38. Behrens, E. (2013). The oceanic response to Greenland melting: the effect of increasing model resolution.
Dissertation, Christian-Albrechts-Universität zu Kiel, 166 pp.
S39. Böning, C.W., Behrens, E., Biastoch, A., and Bamber, J.L. (2016). Emerging impact of Greenland meltwater
on deepwater formation in the North Atlantic Ocean. Nat. Geosci. doi: 10.1038/NGEO2740.
S40. Barnier, B., Madec, G., Penduff, T., Molines, J.-M., Treguier, A.-M., Le Sommer, J., Beckmann, A.,
Biastoch, A., Böning, C., Dengg, J., et al. (2006). Impact of partial steps and momentum advection schemes
in a global ocean circulation model at eddy-permitting resolution. Ocean Dyn. 56, 543–567.
S41. Debreu, L., Vouland, C., and Blayo, E. (2008). AGRIF. Adaptive grid refinement in Fortran. Comput. Geosci.
34, 8–13.
S42. National Geophysical Data Center (2006). 2-minute Gridded Global Relief Data (ETOPO2) v2. National
Geophysical Data Center, NOAA. doi:10.7289/V5J1012Q.
S43. Blanke, B., and Delecluse, P. (1993). Variability of the tropical Atlantic Ocean simulated by a general
circulation model with two different mixed-layer physics. J. Phys. Oceanogr. 23, 1363–1388.
S44. Levitus, S., Boyer, T.P., Conkright, M.E., Johnson, D., O' Brien, T., Antonov, J., Stephens, C., and Gelfeld,
R. (1998) NOAA Atlas NESDIS 18, World Ocean Database 1998. Volume 1: Introduction. U.S. Gov.
Printing Office, Washington D.C., 346 pp.
S45. Large, W.G., and Yeager, S. (2009). The global climatology of an interannually varying air-sea flux data set.
Climate Dyn. 33, 341–364.
S46. Griffies, S.M., Biastoch, A., Böning, C., Bryan, F., Danabasoglu, G., Chassignet, E.P., England, M.H, Gerdes,
R., Haak, H., Hallberg, R.W., et al. (2009). Coordinated Ocean-ice Reference Experiments (COREs). Ocean
Model. 26, 1–46.
S47. Mertens, C., Rhein, M., Walter, M., Böning, C.W., Behrens, E., Kieke, D., Steinfeldt, R., and Stöber, U.
(2014). Circulation and transports in the Newfoundland Basin, western subpolar North Atlantic. J. Geophys.
Res. 119, 7772–7793.
S48. Baltazar-Soares, M., Biastoch, A., Harrod, C., Hanel, R., Marohn, L., Prigge, E., Evans, D., Bodles, K.,
Behrens, E., Böning, C.W., and Eizaguirre, C. (2014). Recruitment collapse and population structure of the
European eel shaped by local ocean current dynamics. Curr. Biol. 24, 104–108.
S49. Khripounoff, A., Comtet, T., Vangriesheim, A., and Crassous, P. (2000). Near-bottom biological and mineral
particle flux in the Lucky Strike hydrothermal vent area (Mid-Atlantic Ridge). J. Mar. Sys. 25, 101–118.
S50. Mullineaux, L.S., Mills, S.W., Sweetman, A.K., Beaudreau, A.H., Metaxas, A., and Hunt, H.L. (2005).
Vertical, lateral and temporal structure in larval distributions at hydrothermal vents. Mar. Ecol. Prog. Ser.
293, 1–16.
S51. Speer, K.G., and Rona, P.A. (1989). A model of an Atlantic and Pacific hydrothermal plume. J. Geophys.
Res. 94, 6213–6220.
S52. Gary, S.F., Lozier, M.S., Biastoch, A., and Böning, C.W. (2012). Reconciling tracer and float observations of
the export pathways of Labrador Sea Water. Geophys. Res. Lett. 39, L24606.