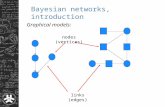
Bayesian Networks. Graphical Models Bayesian networks Conditional random fields etc.
Bayesian Networks
description
Transcript of Bayesian Networks

Bayesian Networks• A causal probabilistic network, or Bayesian network, is an directed acyclic graph (DAG) where nodes represent variables and links represent dependency relations, e.g. of the type cause-effect, between variables and quantified by (conditional) probabilities
• Qualitative component + quantitative component
A
B
C
D
E
F
G
H

Bayesian Networks
• Qualitative component : relations of conditional dependence / independence
I(A, B | C): A and B are independent given CI(A, B) = I(A, B | Ø): A and B are a priori independent
• Formal study of the properties of the ternary relation I
• A Bayesian network may encode three fundamental types of relations among neighbour variables.

Qualitative Relations : type I
FGH
Ex: F: smoke, G: bronchitis, H: respiratory problems (dyspnea)
Relations:¬ I(F, H)
I(F, H | G)

Qualitative Relations : type II
EFG
Ex: F: smoke, G: bronchitis,
E: lung cancer
Relations:¬ I(E, G)
I(E, G | F)

Qualitative Relations : type III
B C E
Ex: C: alarm, B: movement detection,
E: rain
Relations: I(B, E)
¬ I(B, E | C)

Probabilistic component
• Qualitative knowledge: a directed acyclic graph G (DAG)Nodes(G) = V = {X1, …, Xn} -- discrete variables --Edges(G) VxVParents(Xi) = {Xi : (Xj, Xi) Edges(G)}
• Probabilistic knowledge: P(Xi | parents(Xi))
These probabilities determine a joint probability distribution P over V = {X1, …, Xn}:
P(X1, …, Xn) = P(X1 | parents(X1)) · · · P(Xn | parents(Xn))
Bayesian Network = (G, P)

Joint Distribution
• P(X1,X2,...Xn) = P(Xn|Xn-1,...X1) ... P(X3|X2,X1) P(X2|X1) P(X1).
• Independence relations of each variable Xi with the set of predecessor variables of the parents of Xi:
P(Xi | parents(Xi), Y1,.., Yk) = P(Xi | parents(Xi))
P(X1, X2, ..., Xn) = i=1,n P(Xi | parents(Xi))
• to have in each node Xi the conditional probability distribution P(Xi | parents(Xi)) is enough to determine the full joint probability distribution P(X1,X2,...,Xn)

ExampleA
B
C
D
E
F
G
H
P(A): P(a) = 0.01P(B | A): P(b | a) = 0.05, P(b | ¬a) = 0.01P(C | B,E): P(c | b, e) = 1, P(c | b, ¬e) = 1, P(c | ¬b, e) = 1, P(c | ¬b, ¬e) = 0P(F): P(f) = 0.5P(D | C): P(d | c) = .98, P(d | ¬c) = 0.05P(E | F): P(e | f) = 0.1, P(e | ¬f) = 0.01P(G | F): P(g | f) = 0.6, P(g | ¬f) = 0.3P(H | C, G): P(h | c,g) =0.9 , P(h | c,¬g) = 0.7, P(h | ¬c,g) = 0.8, P(h | ¬c,¬g) = 0.1,
P(A,B,C,D,E,F,G,H) = P(D | C) P(H | C, G) P(C | B, E) P(G | F) P(E | F) P(F) P(B | A) P(A)
P(a,¬b,c,¬d,e,f,g,¬h) = P(¬d |c) P(¬h |c,g) P(c | ¬b,e) P(g | f) P(e | f) P(f) P(¬b | a) P(a) = (1- 0.98) (1-0.9) 1 0.6 0.1 0.5 (1-0.05) 0.01 = 5,7 10-7.
A: visit to Asia B: tuberculosisF: smoke E: lung cancerG: bronchitis C: B or ED: X-ray H: dyspnea

D-separation relations and probabilistic independence
Goal: precesely determine which independence relations (graphically) are defined by one DAG.
Previous definitions:
• A path is a sequence of connected nodes in the graph. • A non directed path is a path without taking into account the directions of the arrows. • A “head-to-head” link in a node is a (non directed) path of the form xyw, the node y is clalled a “head-to-head” node.

D-separation• A path c is called to be activated by a set of nodes Z if the following two conditions are satisfied:
1) Every node in c with links head-to-head is in Z or it has a descendent in Z.
2) Any other node in c does not belong to Z.Otherwise, the path c is called to be blocked by Z.
Definition. If X, Y and Z are three disjoint subsets of nodes disjunts in a DAG G, then Z d-separates X from Y, or equivalently X and Y are graphically independent given Z, when all the paths between any node from X and any node from Y are blocked by Z

D-separationA
B C
G
E
D
Theorem. Let G be a DAG and let X,Y and Z be subsets of nodes such that X and Y are d-separated by Z. Then, X and Y are conditionally independent from Zfor any probability P such that (G, P) is a causal network over G, that is, s.t. P(X | Y,Z) = P(X | Z) and P(Y | X,Z) = P(Y | Z).
{B} and {C} are d-separated by {A}:
Path B-E-C: E,G {A} - {A} blocks the path B-E-C
Path B-A-C: - {A} blocks the path B-A-C

Inference in Bayesian NetworksKnowledge about a domain encoded by a Bayesian network XB = (G, P).
Inference = updating probabilities: evidence E on values taken by some variables modify the probabilities of the rest of variables
P(X) --- > P’(X) = P(X | E)
Direct Method:
XB = < G = {A,B,C,D,E}, P(A,B,C,D,E) >
Evidence: A = ai, B = bjP ( a i , b j , c k , d m , e p )
m , p
∑
P ( a i , b j , c k , d m , e p )
k , m , p
∑
P(C = ck | A = ai, B = bj) =

Inference in Bayesian Networks• Bayesian networks allow local computations, which exploit the indepence relations among variables explictly induced by the corresponding DAG of the networks.
• They allow updating the probability of a variable using only the probabilities of the immediat predecessor nodes (parents), and in this way, step by step to update the probabilities of all non-instantiated variables in the network ---> propagation methods
• Two main propagation methods:
• Pearl method: message passing over the DAG
• Lauritzen & Spiegelhalter method: previous transformation of the DAG in a tree of cliques

Propagation method in trees of cliques
1) transformation of initial network in another graphical structure, a tree of cliques (subsets de nodes)
equivalent probabilistic information
BN = (G, P) ----> [Tree, P]
2) propagation algorithm over the new structure

Graphical TransformationDefinition: a “clique” in a non-directed graph is a complete
and maximal subgraph
To transform a DAG G in a tree of cliques:
1) Delete directions in edges of G: G’
2) Moralization of G’: add edges between nodes with common sons in the original DAG G: G’’
3) Triangularization of G’’ : G*
4) Identification of the cliques in G*
5) Suitable enumeration of the cliques (Running Inters. Prop.)
6) Construction of the tree according to the enumeration

Example (1)
A
B
C
D
E
F
G
H A
B
C
D
E
F
G
H
A
B
C
D
E
F
G
H
1)
2)

Example (2): triangularizationA
B
C
D
E
F
G
H
A
B
C
D
E
F
G
H
A
B
C
D
E
F
G
H
3) 3)

Example (3): cliques
A
B
C
D
E
F
G
H
A
B
C
D
E
F
G
H
Cliques:{A,B}, {B,C,E}, {E,F,G}, {C,E,G}, {C,G,H}, {C,D}
Cliques:4)

Ordering of cliques
Enumeration of cliques Clq1, Clq2, …, Clqn such that the following property holds:
Running Intersection Property: for all i=1,…, n there exists j < i such that Si Clqj , where Si = Clqi(Clq1Clq2...Clqi-1).
This property is guaranteed if: (i) nodes of the graph are enumerated following the criterion of “maximum cardinality search”(ii) cliques are ordered according to the node of the clique with a highest ranking in the former enumaration.

Example (4): ordering cliques
A
B
C
D
E
F
G
H
1
2
4
8 7
3
6
5
Clq1 = {A,B}, Clq2 = {B,E,C}, Clq3 = {E,C,G}, Clq4 = {E,G,F}, Clq5 = {C,G,H}, Clq6 = {C,D}

Tree Construction
Let [Clq1, Clq2, …, Clqn ] be an ordering satisfying R.I.P.
For each clique Clqi, define
Si = Clqi(Clq1Clq2...Clqi-1)Ri = Clqi-Si.
Tree of cliques:- (hyper) nodes: cliques- root: Clq1
- for each clique Clqi, its “father” candidates are
cliques Clqk with k < i and s.t. Si Clqk
(if more than one candidate, random selection)

Example (5): trees S2 = Clq2 Clq1{Clq1
S3 = Clq3(Clq1Clq2){E,CClq2
S4 = Clq4(Clq1Clq2 Clq3){GClq3
S5 = Clq5(Clq1Clq2 Clq3.Clq4){C,GClq3
S6 = Clq6( Clq1Clq2 Clq3.Clq4Clq5){CClq2, Clq3, Clq5
Clq1
Clq2
Clq3
Clq4 Clq5Clq6
Clq1
Clq2
Clq3
Clq4 Clq5
Clq6
Clq1
Clq2
Clq3
Clq4 Clq5
Clq6

Propagation Algorithm
• Potential Representation of the distribution P(X1, …, Xn):
([W1...Wp], ) is a potential representation of P, where the Wi
are subsets of V = {X1, …, Xn}, if P(V) =
• In a Bayesian network (G, P): P(X1, ..., Xn) = P(Xn| parents(Xn))·...· P(X1| parents(X1))
admits a potential representationP(X1, ..., Xn) = (Clq1) ·(Clq2) · ...·(Clqm)
with (Clqi)= ∏{P(Xj | parents(Xj)) | XjClqi, parents(Xj) Clqi ,
K ( W i )
i = 1
p
∏

Propagation Algorithm (2)
Fundamental property of the potential representations:
• Let ([W1, ..., Wm], ) be a potential representation for P. Evidence: X3 = a and X5 = b.
• Problem: update the probabilitaty P’(X1, ..., Xn) = P(X1, ..., Xn| X3=a,X5 = b) ??
Define: W^i = Wi - {X3, X5} ^(W^i) = (Wi (X3=a, X5=b))
([W^1, ..., W^m], ^) is a potential representation for P'.

Example (6): potentialsClq1
Clq2
Clq3
Clq4 Clq5
Clq6
Clq1 = {A,B}, Clq2 = {B,E,C}, Clq3 = {E,C,G}, Clq4 = {E,G,F}, Clq5 = {C,G,H}, Clq6 = {C,D}
A
B
C
D
E
F
G
H
(Clq1) = P(A)· P(B | A) (Clq2) = P(C | B,E), (Clq3) = 1 (Clq4) = P(F).P(E | F).P(G | F), (Clq5) = P(H | C, G)(Clq6) = P(D | C)
P(A,B,C,D,E,F,G,H) = P(D | C) P(H | C, G) P(C | B, E) P(G | F) P(E | F) P(F) P(B | A) P(A)
P(A,B,C,D,E,F,G,H) = (Clq1) • …. • (Clq6)

Example(6): potentials
(Clq1) = P(A)· P(B | A)(a,b) = P(a) · P(b | a) = 0.005(¬a,b) = P(¬a) · P(b | ¬a) = 0.0099(a,¬b) = P(a) · P(¬b | a) = 0.0095(¬a,¬b) = P(¬a) · P(¬b | ¬a) = 0.9801
(Clq5) = P(H | C, G)(c,g,h) = P(h | c,g) = 0.9 (c,g,¬h) = P(¬h | c,g) = 0.1(c,¬g,h) = P(h | c,¬g) = 0.7 (c,¬g,¬h) = P(¬h | c,¬g) = 0.3(¬c,g,h) = P(h | ¬c,g) = 0.8 (¬c,g,¬h) = P(¬h | ¬c,g) = 0.2(¬c,¬g,h) = P(h | ¬c,¬g) = 0.1 (¬c,¬g,¬h) = P(¬h | ¬c,¬g) = 0.9
…


Propagation algorithm: theoretical resultats
Causal network (G, P)([Clq1, ..., Clqp], ) is a potential representation for P
1) P(Clqi) = P(Ri|Si).P(Si)
2) P(Rp|Sp) = , where is the marginal
of the function with respect to the variables of Rp.
3) If father(Clqp) = Clqj, then ([Clq1,...Clqp-1], ') is a potential representation for the marginal distribution of P(V-Rp) where:
'(Clqi)=Clqi) for all i≠j, i < p'(Clqj)=Clqj)
( Clq p )
ψ ( Clq p )
R p
∑
( Clq p )
R p
∑
( Clq p )
R p
∑

Propagation algorithm: step by step (2)
Goal: to compute P(Clqi) for all cliques.
Two graph traversals: one bottom-up and one top-down
BU) start with clique Clqp . Combining properties 2 i 3 we have, an iterative form of computing the conditional distributions P(Ri|Si) in each clique until reaching the root clique Clq1.
Root: P(Clq1)=P(R1|S1).
TD) P(S2)= , and from there P(Si)=
--we can always compute in a clique Clqi the distribution P(Si) whenever we have already computed the distribution of its father clique Clqj --
P ( Clq 1 )
Clq 1 − S 2
∑P ( Clq j )
Clq j − S i
∑

Clq1
Clq2
Clq3
Clq4 Clq5
Clq6
Clq1
Clq2
Clq3
Clq4 Clq5
Clq6
P(Ri | Si)
P(Si)
P(Clqi) = P(Ri,Si) = P(Ri | Si) P(Si)

Clqi P(Ri|Si) = =
(Clqi)Ri(Clqi)
(Clqi)
(Clqi)’(Clqi) =
(Clqi) j(Sj) k(Sk) Clqi
Clqj Clqk
Clqi
Clqj Clqk
(Clqi) i(Si)
Case 1)
Case 2)

Clq1
Clq2
Clq3
Clq4 Clq5
Clq6
6(S6)
5(S5) 4(S4)
3(S3)
2(S2)

Example (7)
A) Bottom-up traversal: passing k(Sk) = Rk(Clqk),
Clique Clq6 = {C,D} (R6= {D}, S6 = {C}).
P(R6|S6) = P(D | C) =
6(c) = (c, d) + (c, ¬d) = 0.98 + 0.02 = 16(¬c) = (¬c, d) + (¬c, ¬d) = 0.05 + 0.95 = 1,
P(d | c) = P(¬d | c) = 0.02
P(d | ¬c) = P(¬d | ¬c) = 0.95
( R6
, S6
)
λ6
( S6
)
( c , d )
λ6
( c )
=
0 . 98
1
= 0 . 98
( ¬ c , d )
λ ( ¬ c )
=
0 . 05
1
= 0 . 05

Example (7)
Clique Clq5 = {C, G, H} (R5 = {H}, S5 = {C, G}).
This node is clique Clq6’s father. According to point [3], we modify the potential function of the clique Clq5:
'(Clq5)=Clq5)
P(R5 | S5) = P(H | C,G) =
where 5(C,G) =
5(c,g) = '(c, g, h) + '(e, g, ¬h) = 0.9 + 0.1 = 15(c,¬g) = '(c, ¬g, f) + '(c, ¬g, ¬h) = 0.7 + 0.3 = 15(¬c,g) = … = 5(c,¬g) = ...= 1
( Clq6
)
R6
∑ = ψ ( Clq 5 ) ⋅ λ6
( S6
)
' ( Clq5
)
ψ ' ( Clq 5 )
R 5
∑
=
ψ ' ( R5
, S5
)
λ5
( S5
)
' ( C , G , H )
H
∑

Exemple (7)
Clique Clq3 = {E,C,G} (R3 = {G}, S3 = {E,C})
Clq1
Clq2
Clq3
Clq4 Clq5
Clq6
Clq3 is father of two cliques: Clq4 and Clq5, both already processed
'(Clq3) = Clq3) R(Clq4) · R(Clq5)
= (Clq5) · 4(S4) · 5(S5)
'(E,C,G) = E,C,G) · 4(E,G) · 5(C,G)
P(R3 | S3) = P(G | E, C) =
where 3(E,C) =
' ( Clq3
)
ψ ' ( Clq3
)
R 3
∑
=
ψ ' ( R3
, S3
)
λ3
( S3
)
' ( E , C , G )
G
∑

Example (7)Root: Clique Clq1 = {A, B} (R1 = {A, B}, S1 = ).
'(A,B)=A,B) · 2(B)
P(R1) = P(R1 | S1) =
where 1 = '(a,b) + '(a,¬b)+'(¬a,b)+'(¬a,¬b).
P(A,B) = A,B) : P(a,b) = 0.005, P(a, ¬b) = 0.0095, P(¬a, b) = 0.099, P(¬a, ¬b) =
0.9801
' ( Clq1
)
ψ ' ( Clq 1 )
R 1
∑
=
ψ ' ( R1
)
λ1
( ∅ )
=
ψ ' ( A , B )
λ1

Clqi
Clqj Clqk
P(Clqi) = P(Ri|Si).P(Si)
P(Sk) = Clqi -Sk P(Clqi) = i(Sk) P(Sj) = Clqi -Sj P(Clqi) = i(Sj)

Clq1
Clq2
Clq3
Clq4 Clq5
Clq6
5(S6)
3(S5) 3(S4)
2(S3)
1(S2)

Example (7)
A) Top-down traversal:
Clique Clq2 = {B,E,C} (R2 = {E,C}, S2 = {B}).
P(B) = P(S2) =
P(b) = P(a, b) + P(¬a, b) = 0.005 + 0.099 = 0.104 , P(¬b) = P(a, ¬b) + P(¬a, ¬b) = 1- 0.104 = 0.896
*** P(Clq2) = P(R2 | S2) · P(S2)
P ( Clq 1 )
Clq 1 − S 2
∑

Example (7)
Clique Clq3 = {E,C,G} (R3 = G, S3 = {E,C}).
we have to compute P(S3) i P(Clq3)
Clique Clq4 ={E, G, F} (R4 = {F}, S4 = {E,G}).
we have to compute P(S4) i P(Clq4)
Clique Clq5 = {C, G, H} (R5 = {H}, S5 = {C, G}).
we have to compute P(S5) i P(Clq5)
Clique Clq6 = {C,D} (R6= {D}, S6 = {C}).
we have to compute P(S6) i P(Clq6)


Summary
Given a Bayesian network BN = (G, P), we have seen how
1) To transform G into a tree of cliques and factorize P as
P(X1, ..., Xn) = (Clq1) ·(Clq2) ·...·(Clqm)
where (Clqi)= ∏{P(Xj | parents(Xj)) | XjClqi, parents(Xj) Clqi,
2) To compute the probabilty distributions P(Clqi) with a propagation algorithm, and from there, to compute the probabilities P(Xj) for Xj Clqi, by marginalization.

Probability updating
It remains to see how to perform inference,
i.e. how to update probabilities P(Xj) when some information (evidence E) is available about some variables:
P(Xj) --- > P*(Xj) = P(Xj | E)
The updating mechanism is based in a fundamental property of the potential representations when applied to P(X1, ..., Xn) and its potential representation in terms of cliques:
P(X1, ..., Xn) = (Clq1) ·(Clq2) ·...·(Clqm)

Updating mechanismRecall:
• Let ([Clq1, ..., Clqm], ) be a potential representation for P(X1, …, Xn).
• We observe: X3 = a and X5 = b.
• Actualització de la probabilitat: P*(X1,X2,X4,X6,..., Xn) = P(X1, ...,Xn| X3=a,X5 = b)
Define: Clq^i = Clqi - {X3, X5} ^(Clq^i) = (Clqi (X3=a, X5=b))
([Clq^1, ..., Clq^m], ^) is a potential representation for P*.

Updating mechanism
Based on three steps:
A) build the new tree of cliques obtained by deleting from the original tree the instantiated variables,
B) re-compute the new potential functions ^ corresponding to the new cliques and, finally,
C) apply the propagation algorithm over the new tree of cliques and potential functions.

A,B
B,E,C
E,C,G
E,G,F C,G,H
C,D
Clq1
Clq2
Clq3
Clq4
Clq5
Clq6
B
B,E,C
E,C,G
E,G,F C,G
C,D
Clq’1
Clq’2
Clq’3
Clq’4
Clq’5
Clq’6
A = a, H = bP(Xj) P*(Xj) = P(Xj | X=a,H=h)

A = a, H = b

A = a, H = b


P(D = d | A = a, H = h) ?










![Learning Bayesian Networks in R · 2013-07-10 · Bayesian Networks Essentials Bayesian Networks Bayesian networks [21, 27] are de ned by: anetwork structure, adirected acyclic graph](https://static.fdocuments.in/doc/165x107/5f3267ce969e2b02050fd06c/learning-bayesian-networks-in-r-2013-07-10-bayesian-networks-essentials-bayesian.jpg)