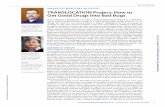
Balanced Translocation detected by FISH
description
Transcript of Balanced Translocation detected by FISH

1
Balanced Translocation detected by FISH

2
Red-Chrom. 5probe
Green-Chrom. 8probe

3
2D Protein Gels

4

5
MS-peptide size signature: match to all predicted proteins

6
1. Follow the mutation
2. Follow which regions of DNA are co-inherited (linked)
Positional Cloning by Recombination Mapping

7
1. Follow the mutation
To determine disease genepresence or absence (genotype) from phenotype you must first establish
Dominant / recessiveAurosomal / sex-linked
Positional Cloning by Recombination Mapping

8
SINGLE GENE DEFECTS
Modes of InheritanceTo deduce who (likely) has one or two copies of mutant gene
Affected FemaleUnaffected Male

9
AUTOSOMAL DOMINANT
+/+ D/+
D/++/+

10
RECESSIVEAUTOSOMAL X-LINKED
RECESSIVE
a/+
a/a
a/+ x/++/Y
x/+
x/Y
+/Y

11

12
2. Follow which DNAs are co-inherited (linked)
Use DNA sequences that differamong individuals within a family-Polymorphisms.
Positional Cloning by Recombination Mapping
T G
C A

13
VNTR / STRP DETECTION

14
A1A1A3
A1
A3
A4
A2
A1
A2A2 A4A4 A3
A3

15
X
X
X
2 3
Parent
Gamete
Child
A1
A2
B1
B2
C1
C2
B1
B1
C1
C1
A2
A2

16
Recombination Mapping
Measures distance between 2 sites on a chromosome according to frequency of recombination
Distance between 2 DNA markersor
Distance between a “disease gene”and a DNA marker

17
No fixed proportionalConversion between
Genetic distance (cM)
and
Physical distance (kb, Mb)

18
FAMILY A
A1 D
A2 +
NR NR NR NR NR RD D D D+ +

19
FAMILY B
A1 D
A2 +
NR NR NR NR NR R
A1 +
A2 D
R R R R R NR

20
INFORMATIVE MEIOSIS
Ideally:- unambiguous inheritance of mutation and markers
(requires heterozygosity for each in parent)
knowledge of which alleles linked in parent (phase)

21
Assign numbers to results of linkage analysis
to deal with non-ideal meioses
to sum data from many meioses in a family
to sum data from several families

22
Likelihood of R
Likelihood of NR
If linked and RF = If unlinked:-
1 -
1/2
1/2
Family A has 1 recombinant and 5 Non-RecombinantsLikelihood, given linkage of
L ( )
= . (1- ) 5
L (1/2) = (1/2) 6
Z = Lod = log { L ( ) / L (1/2)}
Or given unlinked:-
Z 0.58 0.62 0.51 0.3 0
0.1 0.2 0.3 0.4 0.5

23
Z = 3
Lod

24
FAMILY B
A1 D
A2 +
NR NR NR NR NR R
A1 +
A2 D
R R R R R NR

25
Family B:- Disease gene may be linked to A1 or A2
Consider equally likely50% chance Family B has 1 R and 5 NR50% chance Family B has 5 R and 1 NR
L ( )L (1/2) = (1/2) 6
Z = Lod = log { L ( ) / L (1/2)}Z 0.28 0.32 0.22 0.08 0
0.1 0.2 0.3 0.4 0.5
= . (1- ) 51/2 { } + . (1- ) 5
1/2 { }

26
Z 0.28 0.32 0.22 0.08 0
0.1 0.2 0.3 0.4 0.5
Phase unknown
Z 0.58 0.62 0.51 0.3 0
0.1 0.2 0.3 0.4 0.5
Phase known

27
Z = Z1 + Z2 + Z3 + Z4 +…..
Z = Z(A) + Z(B) + Z(C) + Z(D) + ….
For family “A” with meioses 1, 2, 3, 4 …..
For multiple families, “A”, “B”, “C”, “D”…..
Assumption: same gene responsible for disease in all families
Problem: locus heterogeneity

28
Z = 3
Lod

29

30
LINKAGE DISEQUILIBRIUM
Manygenerations

31
PCR test DNA segments

32

33
Testing for specific mutations

34
ARMS 3’ mis-match of primer

35

36
OLA

37

38

39

40

Aa BB CC DD Ee FF Gg HH II JJ AA BB CC Dd Ee FF GG HH II Jj
AA BB CC Dd Ee FF Gg HH II JJ
a B C D e/E F G H I J
A B C d E/e F G H I JA B C D E/e F g H I J
A B C D e/E F G H I j
Mother Father
Son/Daughter
Family Trio SNP genotypes reveal haplotypes
Deduced haplotypes- ignoring recombination

Creation of variant sequencesRearrangement of sequence variants by recombination
First, consider just the creation of variant sequences within a short stretch of DNA where there is no significant rearrangementdue to recombination (an assumption that turns out to be valid)

ABCDEFGHIJKLMNOPQRSTAbCDEFGHIJKLMNOPQRST
ABCDEFgHIJKLMNOPQRSTAbCDEFGHIJKLMNOPqRST
aBCDEFgHIJKLMNOPQRSTAbCDEFGHIJkLMNOPqRST
AbCDEFGhIJkLMNOPqRSTABCDEfGHIJKLMNOPQRST
aBCDEFgHIJKLMNOPQrSTaBCDEFgHIJKLMnOPQrST
bbqbqkbqkh
ggagargarn
f
History

ABCDEFGHIJKLMNOPQRSTAbCDEFGHIJKLMNOPQRST
ABCDEFgHIJKLMNOPQRSTAbCDEFGHIJKLMNOPqRST
aBCDEFgHIJKLMNOPQRSTAbCDEFGHIJkLMNOPqRST
AbCDEFGhIJkLMNOPqRSTABCDEfGHIJKLMNOPQRST
aBCDEFgHIJKLMNOPQrSTaBCDEFgHIJKLMnOPQrST
bbqbqkbqkh
ggagargarn
f
Retention & amplification of only a few haplotypes


For any short region of DNA typically only 4-6 haplotypesare found in a sampling of present day humans (of the manymillions that must have existed in at least one copy en route).These local haplotypes provide some information about ancestry.
Now consider how the major haplotypes of each short region ofDNA are associated with neighboring haplotypes to see whererecombination events took place.

aBCDEFgHIJKLMnOPQrSTUVwXyZ
aBCDEFgHIJKLMnOPQrSTUVwXyZ
aBCDEFgHIJKLMnOPQrSTUVwXyZ
aBCDEFgHIJKLMnOPQrSTUVwXyZ
aBCDEFgHIJKLMnOPQrSTUVwXyZ
aBCDEFgHIJKLMnOPQrSTUVwXyZ
aBCDEFgHIJKLMnOPQrSTUVwXyZ
High LD regions?

aBCDEFgHIJKLMnOPQrSTUVwXyZ
aBCDEFgHIJKLMnOPQrSTUVwXyZ
aBCDEFgHIJKLMnOPQrSTUVwXyZ
aBCDEFgHIJKLMnOPQrSTUVwXyZ
aBCDEFgHIJKLMnOPQrSTUVwXyZ
aBCDEFgHIJKLMnOPQrSTUVwXyZ
aBCDEFgHIJKLMnOPQrSTUVwXyZ
High LD segment High LD segment
Recombination hot-spot


85% of genome made up of 5-20kb high LD blocksOnly 4-5 different major haplotypes per block in the world!

Haplotype blocks

100 kb1 2 3 4 5 6 7 8

Disease No disease/2,000 /3,000Minor allele frequency
SNP-2a 93 130SNP-2b 21 27SNP-3a 140 62SNP-3b 24 35SNP-3c 140 260SNP-3d 87 120……. …. ….……. …. ….……. …. ..