ABCC transporters mediate the vacuolar accumulation of ... · 9/23/2019 · 148 transporters were...
Transcript of ABCC transporters mediate the vacuolar accumulation of ... · 9/23/2019 · 148 transporters were...

1
RESEARCH ARTICLE 1 2
ABCC Transporters Mediate the Vacuolar Accumulation of Crocins in Saffron 3
Stigmas 4
5 Olivia Costantina Demurtas
1, Rita de Brito Francisco
2, Gianfranco Diretto
1, Paola Ferrante
1, Sarah 6
Frusciante1, Marco Pietrella
1,3, Giuseppe Aprea
1, Lorenzo Borghi
2, Mistianne Feeney
4, Lorenzo Frigerio
4, 7
Adriana Coricello5, Giosuè Costa
5, Stefano Alcaro
5, Enrico Martinoia
2*, Giovanni Giuliano
1* 8
9 1ENEA, Italian National Agency for New Technologies, Energy and Sustainable Economic Development, C.R. 10 Casaccia, 00123, Rome, Italy 11 2Department of Plant and Microbial Biology, University of Zurich, 8008 Zurich, Switzerland 12 3Council for Agricultural Research and Economics (CREA), Research Center for Olive, Citrus and Tree Fruit, 13 47121 Forlì, Italy 14 4School of Life Sciences, University of Warwick, Coventry CV4 7AL, United Kingdom 15 5Department of Health Sciences, Magna Græcia University of Catanzaro, 88100 Catanzaro, Italy 16
17 *Corresponding authors: 18 [email protected] 19 [email protected] 20
21 Short title: Saffron vacuolar crocin transporters 22 One-sentence summary: A ‘transportomics’ approach allows identification of crocin vacuolar transporters 23 from saffron stigmas. 24
25
The author(s) responsible for distribution of materials integral to the findings presented in this article in 26 accordance with the policy described in the Instructions for Authors (www.plantcell.org) is Olivia Costantina 27 Demurtas ([email protected]) 28
29 ABSTRACT 30
Compartmentation is a key strategy enacted by plants for the storage of specialized metabolites. The saffron 31 spice owes its red color to crocins, a complex mixture of apocarotenoid glycosides that accumulate in 32 intracellular vacuoles and reach up to 10% of the spice dry weight. We developed a general approach, based on 33 co-expression analysis, heterologous expression in yeast, and in vitro “transportomic” assays using yeast 34 microsomes and total plant metabolite extracts, for the identification of putative vacuolar metabolite transporters, 35 and used it to identify Crocus sativus transporters mediating vacuolar crocin accumulation in stigmas. Three 36 transporters, belonging to both the Multidrug And Toxic Compound Extrusion (MATE) and ATP Binding 37 Cassette C (ABCC) families, were co-expressed with crocins and/or with the gene encoding the first dedicated 38 enzyme in the crocin biosynthetic pathway, CsCCD2. Two of these, belonging to the ABCC family, were able to 39 mediate transport of several crocins when expressed in yeast microsomes. CsABCC4a was selectively expressed 40 in C. sativus stigmas, was predominantly tonoplast localized, transported crocins in vitro in a stereospecific and 41 cooperative way, and was able to enhance crocin accumulation when expressed in Nicotiana benthamiana 42 leaves. 43
44 Keywords: Crocus sativus, stigma, ATP Binding Cassette (ABC) transporters, Multidrug And Toxic Compound 45 Extrusion (MATE) transporters, crocins, in vitro transport assay, microsomes, confocal microscopy. 46
47
48
Plant Cell Advance Publication. Published on September 23, 2019, doi:10.1105/tpc.19.00193
©2019 American Society of Plant Biologists. All Rights Reserved

2
49

3
INTRODUCTION 50
A fascinating feature of vascular plants is their capacity to produce extremely diverse specialized 51
metabolites (Pichersky and Lewinsohn, 2011) and to accumulate some of them to extremely high 52
concentrations: steviol glycosides in Stevia rebaudiana leaves (Brandle and Telmer, 2007) and crocins 53
in Crocus sativus stigmas (Bouvier et al., 2003), for instance, can make up to 10% of the tissue’s dry 54
weight. Plants have evolved a variety of strategies for storing specialized metabolites, such as the 55
accumulation in specialized cell types (McConkey et al., 2000), or their sequestration in the central 56
cellular vacuole (Martinoia et al., 2007). Well-known examples of the latter strategy are the vacuolar 57
sequestration of anthocyanins in Vitis vinifera berries (Francisco et al., 2013), of nicotine in Nicotiana 58
tabacum leaves (Morita et al., 2009), of steviol glycosides in Stevia rebaudiana leaves (Brandle and 59
Telmer, 2007) and of crocins in Crocus sativus stigmas (Bouvier et al., 2003). Vacuolar sequestration 60
has been proposed to prevent the feedback inhibition of the biosynthetic enzymes, and to reduce 61
toxicity effects induced by high cytosolic concentrations of the final products (Goodman et al., 2004). 62
63
Saffron - the most expensive spice on Earth - is composed of the dried stigmas of Crocus sativus 64
(Figure 1A-B), which accumulate large amounts of crocins, apocarotenoid glycosides that confer the 65
red color to the saffron spice (Tarantilis et al., 1995). Up to 15 different crocins have been identified in 66
mature C. sativus stigmas, consisting of both all-trans and 13-cis crocetin esterified with 1 to 5 glucose 67
moieties (Figure 1). The proposed pathway for crocin biosynthesis in C. sativus stigmas starts with the 68
cleavage, in the plastid, of zeaxanthin by Carotenoid Cleavage Dioxygenase 2 (CCD2) (Frusciante et 69
al., 2014). The cleavage product, crocetin dialdehyde, migrates to the endoplasmic reticulum (ER), 70
where it is dehydrogenated to crocetin by a membrane-associated CsALDH3I1, and then glycosylated 71
to crocins 1 and 2ʹ by CsUGT74AD1, localized in the cytosol (Figure 1C-D) (Demurtas et al., 2018). 72
More highly glycosylated crocins are synthesized by an unidentified UDP-glycosyl transferase (UGT), 73
probably localized in the cytosol (Figure 1D). Given their polar nature, crocins synthesized in the 74
cytosol must be transported to the vacuole by one or more tonoplast transporters (Figure 1D). This 75
paper describes their identification. 76
77
Vacuolar transport of glycosylated metabolites, such as glycosylated flavonoids or hormones (ABA 78
glucosyl esters), has been well documented (Gomez et al., 2009; Zhao and Dixon, 2009) and is effected 79
by both Multidrug And Toxic Compound Extrusion (MATE) and ATP Binding Cassette C (ABCC) 80

4
transporters (Martinoia et al., 2007; Francisco and Martinoia, 2018). ABC and MATE transporters have 81
been characterized through expression in yeast, animal or plant cells, isolation of transporter-loaded 82
microsomes, and transport assays using radiolabeled, fluorescent or light-absorbing compounds 83
transported into the microsomes (Marinova et al., 2007; Zhao and Dixon, 2009; Nour-Eldin et al., 84
2012; Burla et al., 2013; Francisco et al., 2013). This limits the number of compounds whose transport 85
can be studied, as well as the capacity to mimic in vivo conditions, where a transporter is exposed to 86
thousands of different compounds. In this work, we study the in vitro transport of multiple metabolites 87
from C. sativus stigma extracts, through a transportomic assay based on liquid chromatography-88
photodiode array-high resolution mass spectrometry (LC-PDA-HRMS), and its use to identify and 89
characterize C. sativus tonoplast transporters involved in vacuolar crocin accumulation. 90
91
RESULTS 92
Identification and characterization of candidate crocin transporters 93
94
A C. sativus stigma transcriptome (Supplemental Data set 1), was searched for expressed genes 95
belonging to the ABCC and MATE classes of tonoplast transporters. Nine ABCC and 11 MATE 96
transporters were expressed in stigmas (Figure 2A, Supplemental Table 1). The highest expressed 97
transcript in stigmas was CsMATE4, followed by CsABCC4a, CsABCC2, CsMATE1a and CsMATE1b. 98
Only CsABCC4a was specifically expressed in stigmas, while CsMATE4 was highly expressed also in 99
tepals and CsABCC2, CsMATE1a and CsMATE1b were mainly expressed in other tissues. To further 100
investigate their possible role in crocin transport, we performed a co-expression analysis (see Methods 101
for more details) of all 20 transcripts with total crocins and with the transcript encoding the first 102
dedicated enzyme in the crocin pathway, CsCCD2 (Frusciante et al., 2014). CsABCC4a expression 103
correlated with both CsCCD2 and crocins (Pearson correlation coefficient (ρ) = 0.99 and 0.95, 104
respectively), followed by CsMATE4 (ρ = 0.84 and 0.75) (Figure 2B, Supplemental Table 2). 105
CsABCC2 displayed a good correlation (ρ= 0.90) with CsCCD2, but not with crocins, while none of the 106
remaining 17 transporters showed significant positive correlations. 107
108
We decided to functionally characterize the five transporters that were most highly expressed in 109
stigmas (underlined in Figure 2A). The corresponding full-length transcripts were isolated from stigma 110
RNA and sequenced (see Methods). Phylogenetic analysis (Figure 2C) revealed that CsABCC4a is 111

5
closely related to Arabidopsis thaliana (At) ABCC4, involved in the vacuolar transport of folates 112
(Klein et al., 2004), while CsABCC2 shows high similarity to AtABCC2, which transports ABA 113
glucosyl esters (ABA-GEs) (Burla et al., 2013), phytochelatins (Song et al., 2010), glutathione 114
conjugates and chlorophyll catabolites (Lu et al., 1998), and to AtABCC1, involved in the transport of 115
phytochelatins (Song et al., 2010) and folates (Raichaudhuri et al., 2009). CsMATE4 is closely related 116
to the MATE1 transporter of Coptis japonica, proposed to mediate the transport of the alkaloid 117
berberine in rhizomes (Takanashi et al., 2017) and CsMATE1a and CsMATE1b are related to the 118
MATE2 protein of Sorghum bicolor that mediates the vacuolar accumulation of the cyanogenic 119
glucoside dhurrin (Darbani et al., 2016), and to Nicotiana tabacum MATE1 and MATE2, which are 120
responsible for the vacuolar accumulation of the alkaloid nicotine (Shoji et al., 2009). The typical ABC 121
signatures (Walker A and Walker B) in the two nucleotide binding domains (NBD1 and NBD2) are 122
well conserved in CsABCC transporters, but many differences were found in the transmembrane 123
domains known to be responsible for substrate recognition/specificity (Theodoulou, 2000; Jasinski et 124
al., 2003; Wilkens, 2015) (Supplemental Figure 1). 125
126
Crocin transporters must be localized in the plant tonoplast, since crocins are synthesized in the cytosol 127
and accumulate in the vacuole (Demurtas et al., 2018). We fused the five transporter genes most 128
expressed in stigmas to the enhanced Green Fluorescent Protein (eGFP) gene (Cinelli et al., 2000) and 129
cloned them in the pBI121 Agrobacterium transformation vector. Nicotiana benthamiana leaves were 130
co-infiltrated with Agrobacterium tumefaciens (strain C58C1) harboring constructs for expression of 131
different eGFP fusions, and the γTIP tonoplast marker fused to the red fluorescent protein (RFP) 132
(Nelson et al., 2007) (Figure 3). At this resolution, the GFP signal showed co-localization with the RFP 133
signal, suggesting that all tested transporters were predominantly tonoplast localized. 134
135
Establishment of a transportomic assay to identify crocin transporters 136
137
Transport of intracellular metabolites by vacuolar transporters is often stereospecific (Bhatia et al., 138
2008; Zhou et al., 2014; Schneider, 2015). Commercial crocin preparations are not representative of the 139
crocin content of C. sativus stigmas, being composed almost exclusively of all-trans crocin 4 140
(Supplemental Figure 2), while in C. sativus stigma six major and several minor crocins, including 13-141
cis isomers, are present (Supplemental Figure 2 and Figure 4B). Therefore, to study the transport of the 142

6
natural crocin substrates, as well as that of other glycosylated compounds accumulated in stigmas 143
(picrocrocin and flavonoid glycosides), we used a C. sativus stigma hydroalcoholic extract as a 144
substrate for the transport reaction. Simultaneous detection of multiple metabolites relied on their 145
detection by LC-PDA-HRMS (Figure 4). We expressed the five C. sativus transporters in yeast cells, 146
isolated microsomes from those cells, and used them to perform transportomic assays. Briefly, the 147
transporters were cloned in the pNEV plasmid and expressed in a yeast strain defective in the ABCC 148
Yeast Bile Transporter1 (YBT1) (ybt1 strain) (Sauer and Stolz, 1994; Paumi et al., 2009). Total 149
microsomes were isolated from the yeast transformants and their intactness was assessed by measuring 150
the transport of leukotriene C4 (LTC4) by LC-HRMS (see Methods) (Supplemental Figure 3). The 151
sensitivity of this method was comparable to the standard radiochemical assay (Leier et al., 1994), 152
indicating that the transportomic assay was sensitive enough to detect transport of metabolites present 153
at low concentrations (fmol to pmol). Microsomes from yeast cells transformed with the pNEV empty 154
vector were used as controls (Figure 4). 155
156
Microsomes expressing CsABCC4a and CsABCC2 were able to mediate the transport of different 157
crocins with different efficiency and in an ATP-dependent manner (Figures 4 and 5). CsABCC4a 158
transported with approximately equal efficiency trans crocin 1, cis crocin 3 and trans crocin 2ʹ, while 159
CsABCC2 showed lower transport efficiency on the latter crocin. In general, crocins carrying smaller 160
glucose groups were efficiently transported in the all-trans form, while crocins with larger gentiobiose 161
groups displayed preferential transport in the cis form. Low, but significant, levels of transport by 162
CsABCC4a and CsABCC2 were also observed for some flavonoids (Figure 5), in agreement with the 163
broad substrate range exhibited by ABCCs (Hwang et al., 2016). Crocin transport by CsABCC4a and 164
CsABCC2 was inhibited by probenecid, a known inhibitor of ABC-type transporters and by incubation 165
on ice, excluding non-specific binding to the yeast microsomes (Table 1). MATE transporters were 166
able to transport only flavonoid glycosides (Figure 5). We did not observe transport of picrocrocin, the 167
most abundant glycosylated metabolite in C. sativus stigma, by any of the transporters tested. 168
169
Given the similarity of CsABCC2 to the ABA-GE transporter AtABCC2 (Burla et al., 2013), we 170
investigated whether CsABCC2 and CsABCC4a could transport ABA-GE. At concentrations up to 7.5 171
µM, i.e. much higher than its physiological intracellular concentration (Burla et al., 2013), neither 172
transporter was able to transport ABA-GE, while they both efficiently transported cis crocin 3 (Table 173
2). 174

7
175
Detailed characterization of the stigma-specific CsABCC4a crocin transporter 176
Although Figure 3 shows that the majority of CsABCC4a localizes to the tonoplast of N. benthamiana 177
mesophyll cells, it does not completely rule out that a small fraction may also be localized to other 178
cellular membranes, such as the ER or the plasma membrane, that in mesophyll cells are not well 179
separated from the tonoplast. To further study the subcellular localization of CsABCC4a:eGFP in N. 180
benthamiana leaves, we conducted co-localization experiments with tonoplast, ER or plasma 181
membrane markers and analyzed the Spearman correlation coefficient (Rs) of the two signals (French 182
et al., 2008). The CsABCC4a:GFP signal strongly co-localized with the γTIP:RFP tonoplast marker 183
(Figure 6A), but did not show significant co-localization with the REFP:HDEL ER marker (Figure 6B) 184
or with the styryl dye FM4-64 labelling the plasma membrane (Figure 6C). Furthermore, GFP 185
fluorescence clearly localized to the tonoplast released after isolation of protoplasts from agroinfiltrated 186
leaves and their partial lysis (Figure 6D). These data confirm that the CsABCC4a transporter is 187
predominantly tonoplast localized when expressed in N. benthamiana leaves. 188
189
Transient expression in N. benthamiana leaves of CsCCD2, encoding a zeaxanthin cleavage 190
dioxygenase leading to the production of crocetin dialdehyde (Frusciante et al., 2014) resulted in the 191
production of low levels of both trans and cis crocetin and trans and cis crocins 1, 2ʹ, 2 and 3 (Figure 192
7). This suggests that N. benthamiana leaves contain endogenous aldehyde dehydrogenase (ALDH) 193
and UGT activities able to transform the CCD2 product (crocetin dialdehyde) into crocetin and crocins 194
(Demurtas et al., 2018). This finding allowed us to test the function of CsABCC4a in N. benthamiana 195
leaves. When CsABCC4a was expressed in combination with CsCCD2, we observed a significant 196
increase in the accumulation of crocins 1, 2 and 2ʹ, and a decrease of crocetin levels (Figure 7). 197
Transportomic assays present advantages (assaying hundreds of metabolites at a time) but also 198
complications (different metabolites can influence each other’s transport). To better investigate this 199
aspect, we studied the transport by CsABCC4a of cis and trans isomers of crocin 3, which are very 200
abundant in mature C. sativus stigmas and are readily separated by preparative HPLC. The two isomers 201
were purified from C. sativus stigmas and their purity was confirmed by LC-PDA-HRMS 202
(Supplemental Figure 4). At a concentration comparable to that of the whole extract (7.5 µM), the 203
purified cis isomer was efficiently transported in a time-dependent manner by CsABCC4a, while no 204
transport was observed for its trans isomer (Figure 8A). Since the trans isomer is transported in crude 205

8
extracts (Figure 5) we concluded that its transport requires a component present in the whole extract 206
that is lost during isomer purification. We tested the hypothesis that this component is the cis isomer by 207
mixing the two isomers at different concentrations and performing a transport assay. Transport of cis 208
crocin 3 was concentration-dependent and was also enhanced by the presence of its trans isomer 209
(Figure 8B, left panel). Transport of the trans isomer occurred only at the highest concentration, and 210
only in the presence of the cis one (Figure 8B, right panel). We conclude that crocin 3 transport by 211
CsABCC4a is cooperative and that this cooperativity allows transport of trans crocin 3 in whole 212
extracts (Figure 5). 213
214
215
Discussion 216
The capacity of specialized plant tissues to accumulate high levels of specialized metabolites relies on 217
their compartmentation capacity. Plant cells contain at least two types of vacuoles, storage and lytic. 218
While storage vacuoles of seeds are small, the lytic vacuoles, which are the main vacuoles in the 219
vegetative tissues, are generally very large and allow accumulation of large amounts of nutrients and 220
specialized metabolites. Lytic vacuoles also play an important role in the detoxification of toxic 221
compounds such as heavy metals and sodium. Additional important vacuolar functions include 222
metabolite storage as well as pH homeostasis, calcium signaling, and the participation in guard cell 223
movements (Martinoia et al., 2007; Martinoia et al., 2012). Plant specialized metabolites stored in 224
vacuoles are usually transported into the vacuolar lumen by different classes of tonoplast-localized 225
transporters. A single class of specialized metabolites can be a substrate for different types of 226
transporters, as shown for anthocyanins, which are transported by MATE-type antiporters in grape and 227
Medicago (Gomez et al., 2009; Zhao and Dixon, 2009), and also by ABC-type transporters in maize 228
and grape (Goodman et al., 2004; Francisco et al., 2013). Due to the fact that specialized compounds 229
exhibit an enormous variety of structures, it is likely that such structural modifications play an 230
important role in the recognition of a given substrate. 231
A single plant species can synthesize thousands of different molecules that may be transported across 232
one or more organellar membranes, which likely explains the large sizes of both ABC-type (>130) and 233
MATE-type (>110) transporter gene families (Hwang et al., 2016; Liu et al., 2016). Identifying the 234
primary transporter for a given molecule by brute force approaches is, therefore, extremely difficult and 235

9
time consuming. The situation is further complicated by the fact that <0.01% of plant specialized 236
metabolites are available in purified form, and even fewer as radiolabeled compounds for use in 237
radiochemical transport experiments. The transportomic approach described here is able to assess the 238
transport of any plant natural product that can be extracted by simple methods and detected by mass 239
spectrometry, i.e. >95% of plant metabolites. Its sensitivity is comparable to radiochemical transport 240
assays and it allows testing of a large number of substrates – potentially hundreds - in a single assay 241
without prior purification. This approach was first used for the study of a human transporter (Uchida et 242
al., 2007) and the term transportomics was coined by (Krumpochova et al., 2012) in characterization of 243
a murine transporter using mouse urine as substrate. Up to now, the maximum number of plant 244
substrates whose transport has been studied by LC-MS is two (Schaedler et al., 2014). This paper 245
demonstrates the possibility of studying transport in complex plant metabolite mixtures, similar to what 246
is already possible in other organisms (Krumpochova et al., 2012). 247
Given the well-known cooperative nature of the transport of several molecules (Liu et al., 2001), the 248
transportomic approach allows the study of the transport of substrates (e.g. trans crocin 3) that might 249
not be transported in purified form. With these advantages, it is surprising that LC-MS based 250
transportomic assays have been, to date, limited to the study of animal transporters (Uchida et al., 2007; 251
Krumpochova et al., 2012). 252
We have previously demonstrated that the second and third steps in crocin biosynthesis are localized in 253
the ER and cytosol, respectively (Demurtas et al., 2018) (Figure 1). Since crocins cannot passively 254
diffuse through membranes (Lautenschlager et al., 2015), an active transport system is required to 255
allow them to cross the tonoplast membrane and be accumulated within the vacuole. Combining 256
transcriptomic data from stigma with the transportomic approach, we identified the ABCC transporters 257
responsible for this transport step. Both CsABCC4a and CsABCC2 are highly expressed in stigmas, 258
show high co-expression with total crocin levels and/or CsCCD2 in C. sativus, predominantly localized 259
to the tonoplast in N. benthamiana leaves, and able to transport crocins in yeast microsomes. While 260
CsABCC4a is almost exclusively expressed in stigmas, CsABCC2 shows high levels of expression in 261
corms, suggesting that it may be involved in crocin transport in this tissue as well (Rubio-Moraga et al., 262
2010). 263
CsABCC4a and CsABCC2 are related to Arabidopsis transporters involved in the vacuolar transport of 264
folates and ABA-GE, respectively (Raichaudhuri et al., 2009; Burla et al., 2013). In spite of its 265

10
similarity to AtABCC2, CsABCC2 is unable to transport ABA-GE. Typical ABC motifs are well 266
conserved in both transporters, while several differences are found in the transmembrane domains, 267
known to be involved in substrate binding (Theodoulou, 2000; Jasinski et al., 2003; Wilkens, 2015) 268
(Supplemental Figure 1). Since crocin biosynthesis, in the Iridaceae family, is confined to the Crocus 269
genus, the most likely hypothesis is that crocin transport in C. sativus evolved from ancestral 270
transporters whose original substrate was different from crocins. As is frequently observed for ABCC 271
transporters (Hwang et al., 2016), CsABCC4a and CsABCC2 are, to a certain extent, promiscuous, in 272
that they transport crocins and, with lower efficiency, flavonoid glycosides. 273
Expression in N. benthamiana leaves of the first dedicated enzyme in the crocin pathway, CsCCD2 274
(Frusciante et al., 2014), was sufficient to cause crocetin and crocin accumulation, probably due to the 275
presence of ALDH and UGT enzymes able to convert the CsCCD2 product, crocetin dialdehyde, into 276
downstream compounds (Demurtas et al., 2018). CsABCC4a localizes to the tonoplast of N. 277
benthamiana mesophyll cells, and co-expression of it with CsCCD2 caused enhanced accumulation of 278
crocins and a decrease of their precursor, crocetin. This is likely due to increased vacuolar 279
sequestration of crocins and an increase of their stability and/or decrease of feedback inhibition of 280
cytosolic crocin synthesis. Unfortunately, crocin levels obtained in N. benthamiana leaves are too low 281
to assess their subcellular localization microscopically. 282
Crocin transport by CsABCC4a is stereospecific and cooperative, in that it shows a preference for cis 283
crocin 3 compared to its trans counterpart, and that one isomer enhances the transport of the other. 284
Cooperative transport has been previously observed with whole vacuoles or AtABCC2, and glutathione 285
and glucuronide conjugates (Lu et al., 1998; Klein et al., 2000). This cooperativity could be due to 286
structural changes in the large vestibule exhibited by ABCC transporters after binding a first substrate 287
(Johnson and Chen, 2017), which could allow binding of a second substrate molecule in the cavity. 288
Homology modeling and docking analysis is unlikely to provide useful insight into the cooperative 289
transport of trans and cis crocin as CsABCC4a has low similarity to ABCC transporters whose 290
structures have been solved (Supplemental Table 3). 291
Transportomic assays demonstrated that CsABCC4a and CsABCC2 transport cis crocins 3 and 4 more 292
efficiently, while the major crocins accumulated in C. sativus stigmas are the corresponding all-trans 293
counterparts. Acidic pH conditions like those found in the vacuole are known to cause polyene 294
isomerization (Re et al., 2001), and indeed cis crocin 3 is converted to its all-trans isomer at pH 5.2 295

11
(Supplemental Figure 5). We therefore hypothesize that crocins 3 and 4 are transported predominantly 296
in the cis form, and then converted to the all-trans one once exposed to the acidic vacuole sap. Such an 297
“isomerization trapping” mechanism has been demonstrated for instance for apigenin 7 (6-O-malonyl) 298
glucoside, a vacuolar pigment from Petroselinum hortense (parsley) that is subjected to conformational 299
changes under acidic pH (Matern et al., 1983) and suggested for the conversion of FCCs (fluorescent 300
Chl catabolites) to NCCs (non-fluorescent chlorophyll catabolites) during chlorophyll breakdown 301
(Hortensteiner and Krautler, 2011). 302
In this work, we have identified transporters for crocin in C. sativus stigmas using a transportomic 303
approach, which has been accomplished only for a few plant specialized metabolites. Using the method 304
developed here, identification of transporters for other important plant specialized metabolites can be 305
envisaged, which could in turn prove useful tools for increasing the accumulation of these metabolites 306
in planta. An example may be steviosides produced in Stevia rebaudiana leaves. Similar to crocins, 307
steviosides contain an isoprenoid moiety esterified with glucosyl groups, and in both cases the 308
synthesis starts in the plastid and produces a glycosylated product in the cytosol, which is finally 309
transported in the vacuole (Brandle and Telmer, 2007; Demurtas et al., 2018). 310

12
METHODS 311
Plant materials and growth conditions 312
Bioinformatic analyses 313
Phylogenetic analyses were conducted with MEGA version 7 (Kumar et al., 2016), using the neighbor-314
joining method (Saitou and Nei, 1987); the percentage of replicate trees in which the associated taxa 315
clustered together in the bootstrap test (500 replicates) are shown next to the branches (Felsenstein, 316
1985). The evolutionary distances were computed using the p-distance method (Nei and Kumar, 2000) 317
represent number of amino acid differences per site. All ambiguous positions were removed for each 318
sequence pair. Protein alignment was performed with Clustal Omega tool 319
(https://www.ebi.ac.uk/Tools/msa/clustalo/). Co-expression analysis was performed as previously 320
described (Coman et al., 2014; Ahrazem et al., 2018). Briefly, the pairwise Pearson correlation between 321
each transporter and CCD2/crocins was computed, and Fisher’s Z–transformation was used to test the 322
statistical significance of the pairwise correlations. Transmembrane domains were deduced using the 323
TMHMM 2.0 software (http://www.cbs.dtu.dk/services/TMHMM/). 324
325
Cloning 326
CsABCC and CsMATE CDSs were isolated from cDNA obtained by RNA extracted from C. sativus 327
stigma collected the day of anthesis using the SMART PCR cDNA synthesis kit (Clontech; Cat. No. 328
634902) and the Super-ScriptII reverse transcriptase (Life Technologies, Cat No. 18064-014). All the 329
oligonucleotides used to isolate and clone the genes are described in Supplemental Table 4. The 330
CsABCC4a partial sequence was reconstructed from contigs obtained by 454 Titanium sequencing 331
(Frusciante et al., 2014) and amplified from cDNA with the oligonucleotides 1 and 2; the amplicon 332
(amplicon 1) was then cloned in pBlueScript SK (+) vector (Stratagene) digested with EcoRV 333
restriction enzyme. 5ʹ RACE PCR was performed to obtain the 5ʹ sequence of the cDNA using a 334
commercial kit (Life Technologies, Cat No. 18374-058) according to the manufacturer’s instructions. 335
C. sativus cDNA was amplified with the oligonucleotide 3. Tailed cDNA was amplified using 5′-336
RACE abridged anchor primer supplied in the kit and oligonucleotide 4. The PCR product (amplicon 2) 337
was purified using a purification kit (Qiagen, Cat No. 28104) and cloned in pBlueScript SK (+) vector 338
digested with EcoRV. All PCR reactions were performed using Phusion High Fidelity DNA 339
polymerase (New England Biolabs, Cat No. M0530L). Amplicon 1 and 2 were then re-amplified, 340
assembled and cloned in the yeast expression vector pNEV-Ura (Sauer and Stolz, 1994) using Gibson 341

13
assembly method (Gibson et al., 2009). Amplicon 1 was amplified with the oligonucleotides 5 and 6, 342
producing the fragment 1; amplicon 2 with the oligonucleotides 7 and 8, producing the fragment 2; 343
pNEV-Ura plasmid was digested with NotI restriction enzymes (fragment 3). The three fragments were 344
assembled using the Gibson Assembly® Master Mix (New England Biolabs, Cat No. E2611S) 345
according to the manufacturer’s instructions thereby producing the pNEV:CsABCC4a construct. 346
Validation that amplicon 1 and 2 belonged to the same coding sequence was performed by PCR on C. 347
sativus cDNA using the oligonucleotides 9 and 10, that produced the expected full-length amplicon of 348
4491 bp (Supplemental Figure 6). 349
CsABCC2 and CsMATE full-length sequences were amplified from cDNA (Supplemental Figure 6) 350
with the following oligonucleotides: 11 and 12 for CsABCC2; 13 and 14 for CsMATE1a; 15 and 16 for 351
CsMATE1b (these primers anneal on the 5ʹ and 3ʹ UTR, respectively); 17 and 18 for CsMATE4. The 352
amplicons were then re-amplified with oligonucleotides that insert the NotI site and subcloned in the 353
pNEV-Ura plasmid NotI-digested, producing the pNEV:CsABCC4a, pNEV:CsABCC2, 354
pNEV:CsMATE1a, pNEV:CsMATE1b and pNEV:CsMATE4 constructs. All the amplicons and 355
constructs were verified by sequencing. The pNEV-based constructs were expressed in Saccharomyces 356
cerevisae cells as described in the “Transport assay” section. 357
358
For protein localization studies, CsABCC and CsMATE cDNAs were 3ʹ-fused to the coding sequence 359
for the enhanced Green Fluorescent Protein (eGFP) using Gibson assembly method. The pBI:eGFP 360
vector (Frusciante et al., 2014) was digested with XbaI restriction enzyme while cDNAs were 361
amplified from pNEV-based constructs, inserting the sequence encoding for Pro-Gly-Pro tripeptide 362
before the eGFP CDS, using the following oligonucleotides: 19 and 20 for CsABCC4a; 21 and 22 for 363
CsABCC2; 23 and 24 for CsMATE1a; 25 and 26 for CsMATE1b; 27 and 28 for CsMATE4. The purified 364
PCR fragments were then assembled with the pBI:eGFP vector generating the pBI:CsABCC4a:eGFP, 365
pBI:CsABCC2:eGFP, pBI:CsMATE1a:eGFP, pBI:CsMATE1b:eGFP and pBI:CsMATE4:eGFP 366
constructs. For protein overexpression in N. benthamiana leaves, the CsCCD2 cDNA including the 367
sequence coding for its chloroplast transit peptide (Demurtas et al., 2018) and the CsABCC4a cDNA 368
were cloned in the pBI121 vector by Gibson assembly method. The vector was digested with PacI 369
restriction enzyme while the cDNAs were obtained by PCR with the following oligonucleotides: 29 370
and 30 for CsCCD2; 31 and 32 for CsABCC4a. All the plasmids were checked by sequencing before 371
transformation in Agrobacterium tumefaciens strain C58C1. 372
373

14
Expression in Nicotiana benthamiana leaves 374
Agroinfiltration of N. benthamiana leaves was performed as described (Hamilton and Baulcombe, 375
1999). Leaves were co-infiltrated with C58C1 cells containing pBI121:CsABCC4a:eGFP or 376
pBI121:CsABCC2:eGFP or pBI121:CsMATE4:eGFP or pBI121:CsMATE1a:eGFP or 377
pBI121:CsMATE1b:eGFP and pBI:γ-TIP-RFP (tonoplast marker, (Nelson et al., 2007)). To reduce the 378
silencing of the transgenes, leaves were also infiltrated with the silencing suppressor RK19 (Francisco 379
et al., 2013). Leaves were also co-infiltrated with C58C1 cells containing pBI121:CsABCC4a:eGFP, 380
pBI-RFP-HDEL (ER marker (Lee et al., 2013)) and RK19. After 3-6 days, leaves were analyzed using 381
a confocal laser scanning microscope (Olympus FV1000 and Zeiss LSM880). Lasers at 488 nm and 382
543 nm were used to detect green and red fluorescence of GFP and γTIP-RFP or RFP-HDEL (emission 383
571-640 nm), respectively. FM4-64 was excited at 514 nm and detected in the range 616 nm – 645 nm.384
Images were acquired using a 40x oil immersion objective (N.A. 1,30) with optical zooming 1x or 3x. 385
All images were analyzed and processed with ImageJ. 386
For preparation of mesophyll protoplasts, cell walls of N. benthamiana leaves were digested for 2 h at 387
30°C in MCP (500 mM Sorbitol, 1 mM CaCl2, 10 mM MES-KOH, pH 5.6) supplemented with 1% 388
(w/v) Cellulase Y-C, 0.1% (w/v) Pectolyase Y-23. Protoplasts were collected by centrifugation at 1,200 389
x g for 5 min on a cushion constituted of 100% Percoll (v/v) in 500 mM Sorbitol, 20 mM MES. The 390
supernatant was removed and the protoplasts were mixed with the Percoll cushion, overlaid with 25% 391
Percoll and MCP. After centrifugation at 1,200 x g for 5 min the purified protoplasts were collected 392
between 25% Percoll and MCP fractions. Vacuoles were released by mixing (1:1) purified protoplasts 393
with lysis buffer (200 mM Sorbitol, 10% Ficoll, 10 mM HEPES-KOH, pH 8.0). Microscopy was 394
performed using a Leica TCS SP5 confocal microscope. For each experiment, at least three different 395
plants were infiltrated and observed. Each experiment was repeated at least twice (with different 396
batches of plants). At least 5 randomly chosen regions were observed, and representative images were 397
selected for each experiment. 398
For simultaneous expression of CsCCD2 and CsABCC4a, leaves were co-infiltrated with C58C1 cells 399
containing pBI121:CsCCD2, pBI121:CsABCC4a and the silencing suppressor RK19. Three 400
independent infiltration experiments (with different batches of plants) were performed. For each 401
experiment, one apical leaf of four different plants was infiltrated with the CsCCD2 construct, and the 402
other apical leaf with the CsCCD2 + CsABCC4a construct. At four d.p.i., the four leaves infiltrated 403

15
with the same construct were collected as a pool and stored at -80°C. The pools from the three 404
independent experiments were then analyzed by LC-PDA-HRMS. Semi-polar extracts for LC-PDA-405
HRMS analysis were prepared as described in “LC-PDA-HRMS analyses”. Data are presented as 406
average (avg) area of crocins ± standard deviation (std) normalized on the area of the internal standard 407
formononetin (fold IS). The results obtained were evaluated using Student's t-test for the calculation of 408
the significance of the difference between the means of the two groups (CsCCD2 and CsCCD2+ 409
CsABCC4a) (Supplemental Table 5). 410
411
Transport assay 412
The ybt1 yeast mutant (MATa; ura3D::HIS3; leu2-3, 112; his3-D200; bat1D1::URA3) was transformed 413
by electroporation as described (Becker and Guarente, 1991) with the pNEV-based constructs. As 414
control, cells were transformed with the empty vector pNEV-Ura, named pNEV for simplicity. 415
Transformants were selected on minimal synthetic dropout medium lacking uracil. For in vitro 416
transport studies, yeast microsomes were isolated as described (Tommasini et al., 1996). Uptake 417
experiments were performed using the rapid filtration technique (Tommasini et al., 1996). Briefly, 100 418
µL of vesicles (OD600= 4; total proteins content about 400 µg) were mixed with ice-cold transport 419
buffer (0.4 M glycerol, 0.1 M KCl, and 20 mM Tris-MES, pH 7.4) and freshly added with 1 mM DTT, 420
6 mM or 1 mM MgSO4 (in the presence or absence of MgATP, respectively), 100 μg/mL creatine 421
kinase, and 10 mM creatine phosphate. Substrate transport was assayed either in the presence or 422
absence of 4 mM MgATP in a total reaction volume of 650 µL. The transport assay was performed at 423
room temperature. 424
Intactness of microsomes was evaluated by the ability to transport the substrate leukotriene C4 (LTC4) 425
(Leier et al., 1994). The LTC4 substrate (Cayman Chemicals, Cat N. 20210) was used at concentration 426
of 100 µM. At two time points (30 sec and 8 min), 100 μl of the reaction mixture were immediately 427
loaded on a pre-wetted nitrocellulose filter (0.45µm pore size; Millipore) and rapidly washed with 3 x 428
2.5 mL of ice-cold transport buffer. Three technical replicates (100 μl each) were performed for each 429
condition and repeated for each vesicle preparation. The filter-bound vesicles were dissolved by adding 430
1 ml of 75% (v/v) spiked cold methanol (0.5 µg/ml formononetin; Sigma-Aldrich, Cat. No. 47752-431
25MG-F), and metabolites were then extracted at room temperature through continuous agitation for 30 432
min. To remove vesicle lipids, 800 µl of eluted samples was transferred in a microcentrifuge tube and 433
added to 400 µl of chloroform, vortexed and shaken in a Mixer Mill 300 (MM, Retsch) for 5 min at 20 434

16
Hz frequency. Ultrapure water (200 µl) was added to separate the phases, followed by vortexing and 435
centrifugation at 20,000 g for 20 min. Finally, 800 µl of the upper phase was collected, dried with a 436
vacuum concentrator and resuspended in 80 µl of cold 50% (v/v) methanol, and centrifuged at 20,000 g 437
for 10 min to remove precipitates. A 2 µl sample of the supernatant was subjected to LC-MS analysis. 438
All solvents used for the extraction were LC-MS grade (Merck Millipore). 439
To study the transport of C. sativus stigma metabolites, a C. sativus stigma hydroalcoholic extract and 440
purified crocins were used to perform uptake experiments. The C. sativus stigma hydroalcoholic extract 441
was prepared as following: 3 mg of pulverized C. sativus stigmas (Castilla-La Mancha, dried 2013) 442
was resuspended in 300 µl of cold 50% (v/v) methanol, homogenized for 40 min in a mixer mill MM 443
300 (Retsch GmbH, Haan, Germany) at 20 Hz frequency, and centrifuged 20 min at 20,000 g. The 444
supernatant was recovered, total crocin content quantified by LC-DAD, and a standardized amount 445
used to perform the uptake experiments: in a final reaction of 650 µl, we adjusted the C. sativus stigma 446
extract volume in order to have a final concentration of 320 µM of total crocins. Crocin 4 analytical 447
standard was purchased from PhytoLab (Cat. No. 80391). 448
Crocin 3 isomers (trans and cis) were purified from C. sativus stigma hydroalcoholic extract by 449
preparative HPLC. Briefly, 100 mg of pulverized stigmas were resuspended in 1 ml of cold 50% (v/v) 450
methanol and processed as described above. Crocin separation was performed on a LC system (Series 451
200, Perkin Elmer) injecting 100 µl of extract on a C18 reverse-phase column (250 x 10 mm, 5 µm; 452
Ascentis, Supelco) heated at 35 °C. The mobile phases used were water + 0.1% formic acid (A) and 453
acetonitrile + 0.1% formic acid (B) at a total flow rate of 4 ml/min. The separation was developed 454
using 10% B for 1 min, a 10 min linear gradient to 20% B followed by a 20 min linear gradient to 30% 455
B. LC conditions were kept for 10 more minutes, before going back to the initial conditions. Detection456
was performed at 440 nm with an online UV/VIS Detector (Series 200, Perkin Elmer). Two ml 457
fractions were collected, dried in a lyophilizer and resuspended in 50 µl of 50% (v/v) methanol. 458
Competitive assays between trans and cis crocin 3 were performed using concentrations ranging from 459
0.75 µM to 75 µM. 460
All uptake experiments with C. sativus stigma extract or purified crocins were performed using the 461
rapid filtration technique with acetate cellulose filters (0.45 µm pore size; Sartorius), instead of 462
nitrocellulose. Reactions were performed at room temperature in reaction mix as described for LTC4 463
and stopped at different time points. Inhibition assays were performed in the presence of the inhibitor 464
probenecid (SIGMA, Cat. No. P8761) (Francisco et al., 2013). Briefly, yeast microsomes were 465
incubated at room temperature in the reaction mix containing the inhibitor (1 mM); after 10 min, the C. 466

17
sativus stigma extract was added and the uptake experiment performed as described above. For 467
experiments on ice, the tubes containing the reaction mix and C. sativus stigma extract were placed on 468
ice immediately after the addition of yeast microsomes. All experiments were repeated at least three 469
times with independent vesicle preparations, and each experiment was performed with three technical 470
replicates (3 filters, each spotted with 100 µl of reaction mix). 471
For ABA-GE transport studies, in vitro transport assays were performed with the analytical standard of 472
ABA-GE (ChemIm Ltd, Cat. No. 013 2781), using concentrations ranging from 0.075 µM to 7.5 µM. 473
The transport of each metabolite presented was evaluated by LC-PDA-HRMS. 474
475
LC-PDA-HRMS analyses 476
C. sativus stigma polar extracts, standards, eluted fractions from in vitro transport studies and semi-477
polar extracts from N. benthamiana agroinfiltrated leaves were analyzed with a Q-Exactive quadrupole 478
Orbitrap mass spectrometry system (ThermoFisher Scientific), coupled to an LC system equipped with 479
a photodiode array detector (Dionex), as previously described (Demurtas et al., 2018). HPLC 480
separation was performed by injecting 2–5 µl of samples on a C18 Luna reverse-phase column (100 x 481
2.1 mm, 2.5µm; Phenomenex). The mobile phases used were water + 0.1% formic acid (A) and 482
acetonitrile + 0.1% formic acid (B) at a total flow rate of 250 µl/min. The separation was developed 483
using 5% B for 0.5 min, followed by a 24 min linear gradient to 75% B. The ionization was performed 484
in a heated electrospray ionization (HESI) source with nitrogen used as sheath and auxiliary gas, set to 485
45 and 30 units, respectively. The vaporizer temperature was 270 °C, the capillary temperature was 30 486
°C, the discharge current was set to 5 μA and S-lens RF level set at 50. The acquisition was performed 487
in the mass range 110/1600 m/z both in positive and in negative ion mode with the following 488
parameters: resolution 70000, microscan 1, AGC target 1e6, maximum injection time 50. UV-VIS489
detection was continuous from 220 to 700 nm. All solvents used were LC-MS grade (Merck Millipore). 490
Metabolite identification was achieved on the basis of accurate masses and by comparison with 491
authentic standards or with literature data. The ion peak areas were normalized to the ion peak area of 492
the internal standard (formononetin) (fold internal standard). To calculate the transport of each 493
metabolite, fold internal standard values of the reactions performed in the absence of ATP were 494
subtracted to those of the reactions performed in the presence of ATP (net uptake). Data are presented 495
as percentage of net uptake of three biological replicates (average values ± standard deviations are 496
shown in figures and tables). 497
498

18
Accession numbers 499
The sequences of genes reported in this paper have been deposited in the GenBank database: 500
CsABCC4a, accession no. MF966954; CsABCC2, accession no. MF966955; CsMATE4, accession no. 501
MF966956; CsMATE1a, accession no. MH475368; CsMATE1b, accession no. MH475369. The 502
sequences are also shown in the Supplemental Data set 2. The sequences of the transporter contigs 503
shown in Fig. 2C have likewise been deposited with the following accessions: CsABCC4a, accession 504
no. MN401321; CsABCC2, accession no. MN401322; CsMATE4, accession no. MN401323; 505
CsMATE1a, accession no. MN401324; CsMATE1b, accession no. MN401325; CsABCC8, accession 506
no. MN380444; CsABCC3a, accession no. MN380445; CsABCC3b, accession no. MN380446; 507
CsABCC4b, accession no. MN380447; CsABCC5, accession no. MN380448; CsABCC15, accession 508
no. MN380449; CsABCC4c, accession no. MN380450; CsMATE5a, accession no. MN380451; 509
CsMATE2e, accession no. MN380452; CsMATE5b, accession no. MN380453; CsMATE2b, accession 510
no. MN380454; CsMATE2d, accession no. MN380455; CsMATE2c, accession no. MN380456; 511
CsMATE2a, accession no. MN380457; CsMATE2f, accession no. MN380458. 512
Sequence data of ABCC and MATE transporters from Arabidopsis thaliana (At), Zea mays (Zm), Vitis 513
vinifera (Vv), Sorghum bicolor (Sb), Nicotiana tabacum (Nt), Medicago truncatula (Mt) and Coptis 514
japonica (Cj) shown in Figure 2 can be found under the following accession numbers: AtABCC1, 515
NP_181013; AtABCC2, NP_001031116; AtABCC3, NP_187915; AtABCC4, NP_182301; AtABCC5, 516
NP_171908; AtABCC6, NP_187916.3; AtABCC7, NP_187917; AtABCC8, Q8LGU1; AtABCC9, 517
Q9M1C7; AtABCC10, NP_191473; AtABCC11, NP_174331; AtABCC12, Q9C8H0; AtABCC13, 518
NP_001323940; AtABCC14, NP_191829; ZmABCC3, AAT37905; ZmABCC4, A7KVC2; 519
VvABCC1, AGC23330; AtDTX35, NP_194294; AtDTX41/TT12, NP_191462; AtDTX16, 520
NP_200058; AtDTX17, NP_177511; AtDX19, NP_566730; AtDTX29,NP_189291; AtDTX30, 521
NP_198619; AtDTX33, NP_175184; AtDTX40, NP_188806; CjMATE1, BAX73926; MtMATE1, 522
ACX37118; MtMATE2, ADV04045; NtJAT1, CAQ51477; NtJAT2, BAP40098; NtMATE1, 523
BAF47751; NtMATE2, BAF47752; SbMATE2, XP_021303040; VvAM1, ACN88706; VvAM3, 524
ACN91542. 525
526
Supplemental Data 527
Supplemental Figure 1. Conserved domains and transmembrane helices of CsABCC4a and 528
CsABCC2 and of the closely related AtABCC1, AtABCC2 and AtABCC4 transporters. 529

19
Supplemental Figure 2. Crocin composition of a commercial crocin standard and of a C. sativus 530
stigma hydroalcoholic extract. 531
Supplemental Figure 3. Assessment of physiological intactness of yeast microsomes. 532
Supplemental Figure 4. Purity of trans and cis crocin 3 purified from C. sativus stigma extract 533
through preparative HPLC. 534
Supplemental Figure 5. Isomerization of cis crocin 3 into trans crocin 3 at pH 5.2. 535
Supplemental Figure 6. MATE and ABCC full-length amplicons obtained from C. sativus cDNA with 536
the oligonucleotides described in Methods. 537
Supplemental Table 1. Transcript levels of CsMATE and CsABCC transporters in C. sativus tissues. 538
Supplemental Table 2. Co-expression analysis of CsABCC and CsMATE transcripts with total 539
crocins and CsCCD2 transcript. 540
Supplemental Table 3. Homology of CsABCC4a with ABCC transporters with known structures. 541
Supplemental Table 4. Oligonucleotides used in this study. 542
Supplemental Table 5. T-test result of data presented in Figure 7. 543
Supplemental Data set 1. Saffron transcriptome data 544
Supplemental Data set 2. Sequences of C. sativus transporters characterized in this paper. 545
Supplemental Data set 3. Text file of the ABC alignment used for the phylogenetic analysis in Figure 546
2C. 547
Supplemental Data set 4. Text file of the MATE alignment used for the phylogenetic analysis in 548
Figure 2C. 549
550
Author contributions 551
OCD, RBF, AC, GD, PF, SF, MP, GA, LB, MF and LF produced data. OCD, RBF, MP, LB, LF, MF, 552
EM, AC, GC, SA, and GG analyzed data. EM and GG developed the experimental strategy. GG 553
coordinated the study. OCD, RBF, AC, EM and GG wrote the manuscript. All authors reviewed the 554
results and approved the final version of the manuscript. The authors declare no conflict of interest. 555
556
Acknowledgments 557
We thank Barbara Bassin for the preparation of yeast microsomes, Elena Romano at the Centre of 558
Advanced Microscopy “Patrizia Albertano” of University of Rome Tor Vergata for help with confocal 559

20
experiments, Gaetano Perrotta, Paolo Facella, Fabrizio Carbone at the ENEA Trisaia Research Center 560
and IGA Technology Services for RNA-Seq data, and Andrea Aliboni for help with preparative HPLC. 561
This work was supported by grants from the European Union to GG [“From discovery to products: A 562
next generation pipeline for the sustainable generation of high-value plant products”, Contract 613153 563
and “Developing Multipurpose Nicotiana Crops for Molecular Farming using New Plant Breeding 564
Techniques”, Contract 760331] and by COST (European Cooperation in Science and Technology) 565
Action CA15136 “EUROCAROTEN”. 566
Figure legends 567
Figure 1. Saffron crocin biosynthesis and its compartmentation 568
(A) C. sativus flower at anthesis. (B) Scanning electron microscopy of mature stigma. (C) Proposed569
crocin biosynthetic pathway and (D) its compartmentation (Demurtas et al, 2018): the CCD2 enzyme 570
cleaves zeaxanthin in the plastid, producing crocetin dialdehyde, which then migrates to the 571
endoplasmic reticulum (ER) and is dehydrogenated to crocetin by an aldehyde dehydrogenase 572
(ALDH); the glycosylation steps are performed in the cytoplasm by UDP-glycosyltransferase (UGT) 573
enzymes; crocins are then transported into the vacuole by ABC tonoplast transporters functionally 574
characterized in this work. (E) Detailed crocin structures. 575
Figure 2. Expression and phylogenetic analysis of putative C. sativus tonoplast transporters. 576
(A) Heat map of transcript levels for ABCC and MATE transporter genes in different C. sativus577
tissues; data are expressed as log2 of reads per kilobase per million (RPKM) and sorted by decreasing 578
RPKM values in stigma. For further details, see Supplemental Data set 1. (B) Co-expression analysis of 579
genes for CsABCCs and CsMATEs (Bait) with CsCCD2 and with total crocins (Prey). Only ρ values 580
>0.50 are shown. For further details, see Supplemental Data set 1. (C) Phylogenetic relationships of581
ABCC (left) and MATE (right) transporters expressed in C. sativus (Cs) stigma (underlined) inferred 582
using the neighbor-joining method. Colored dots indicate the class of transported substrates for the 583
functionally characterized transporters. The CsABCCs and CsMATEs functionally characterized in this 584
work are indicated by arrows. The trees include transporters from Arabidopsis thaliana (At), Coptis 585
japonica (Cj), Medicago truncatula (Mt), Nicotiana tabacum (Nt), Sorghum bicolor (Sb), Vitis vinifera 586
(Vv) and Zea mays (Zm). The accession numbers are described in the Accession numbers section. The 587
percentage of replicate trees that clustered together in the bootstrap test (500 replicates) is indicated to 588
the left of the branches. The alignment files are shown in the Supplemental Data Sets 3 and 4. 589

21
Figure 3. Subcellular localization of C. sativus transporters in Nicotiana leaves. 590
Confocal images of GFP (green) and RFP (red) fluorescence in N. benthamiana leaves co-expressing 591
the indicated ABCC or MATE transporter fused to GFP and γTIP (a tonoplast marker) fused to RFP. 592
Scale bars, 10 µm. 593
Figure 4. Transportomic assay using C. sativus stigma extract and yeast microsomes. 594
(A) Schematic representation of the assay. Microsomes overexpressing a transporter are incubated with595
C. sativus stigma hydroalcoholic extract in the presence of ATP. Microsomes are separated by the rapid596
filtration technique and washed, and transported metabolites are eluted and quantified by LC-PDA-597
HRMS. 598
(B-E) Representative ESI+/MS chromatograms of the extracted accurate mass of crocetin (M+H+
599
329.1747) generated from crocin fragmentation. (B) C. sativus stigma hydroalcoholic extract; (C) 600
import into microsomes isolated from yeast cells transformed with the pNEV empty vector; (D) import 601
into microsomes expressing CsABCC4a incubated in the absence of ATP; (E) import into microsomes 602
expressing CsABCC4a incubated in the presence of ATP. Incubation was for 15 min. Different peaks 603
represent: trans crocin 4 (RT 10.58); trans crocin 3 (RT 11.52); trans crocin 2ʹ (RT 12.54); cis crocin 4 604
(RT 13.84); trans crocin 2 (RT 14.20); cis crocin 3 (RT 14.82); trans crocin 1 (RT 16.16); cis crocin 2ʹ 605
(RT 16.79); cis crocin 2 (RT 17.02); cis crocin 1 (RT 17.73). 606
Figure 5. CsABCC4a and CsABCC2 transport different crocins in vitro 607
(A) Major glycosylated metabolites present in the C. sativus stigma hydroalcoholic extract. 1:608
kaempferol 3-O-sophoroside-7-glucoside; 2: kaempferol 3,7,4ʹ-triglucoside; 3: kaempferol 7-609
sophoroside; 4: kaempferol 3-β-D-glucopyranoside; 5: kaempferol‐3‐O‐rutinoside; 6: 610
dihydrokaempferol 7-O-glucoside; 7: naringenin glucosides (3 isomers); 8: picrocrocin; 9: trans crocin 611
5 (t 5); 10: trans crocin 4 (t 4); 11: cis crocin 4 (c 4); 12: trans crocin 3 (t 3); 13: cis crocin 3 (c 3); 14: 612
trans crocin 2 (t 2); 15: cis crocin 2 (c 2); 16: trans crocin 2ʹ (t 2ʹ); 17: cis crocin 2ʹ (c 2ʹ); 18: trans 613
crocin 1 (t 1); 19: cis crocin 1 (c 1). Structures of the various crocins are shown in Figure 1E. Values 614
are presented as relative abundances compared to the internal standard used (formononetin) of mean 615
values ± stdev of three independent extract preparations. 616
(B) % net transport of glycosylated metabolites by yeast microsomes expressing candidate tonoplast617
transporters. CTRL: microsomes form yeast cells transformed with the empty vector. CsABCC4a, 618

22
CsABCC2, CsMATE4, CsMATE1a, CsMATE1b: microsomes expressing candidate tonoplast 619
transporters. % net transport was calculated by subtracting the peak areas in the absence of ATP from 620
those in the presence of ATP after 15 min of incubation, and normalizing to the peak areas in the initial 621
extract. Data are the avg ± stdev of three independent microsome preparations. 622
Figure 6. Tonoplast localization of CsABCC4a in Nicotiana leaves. 623
(A-C) N. benthamiana leaves were agroinfiltrated with the indicated constructs and imaged by confocal 624
microscopy. The Spearman correlation coefficient (Rs) between the two channels was calculated using 625
the ImageJ analysis program with the Pearson-Spearman correlation plug-in (French et al., 2008) to 626
calculate co-localization and to produce scatterplots using a threshold of 40. Scale bars, 5 µm. (D) 627
Confocal images of a protoplast from a N. benthamiana leaf expressing CsABCCa:GFP (green 628
channel). After induced lysis of the plasma membrane the vacuole (asterisk) is in the process of being 629
released. Scale bar, 12 µm. 630
Figure 7. Functional assay of the C. sativus ABCC4a transporter in N. benthamiana leaves. 631
Quantification of crocetin and crocins in leaves agroinfiltrated to express CsCCD2 or 632
CsCCD2+CsABCC4a. Data are the avg ± stdev of ion peak areas of crocetin and crocins, normalized to 633
the ion peak area of the internal standard formononetin (fold internal standard), in three independent 634
pools of agroinfiltrated leaves (see Methods for details). Asterisks indicate statistical significance of the 635
difference between the means of the two groups (CsCCD2 and CsCCD2+ CsABCC4a) (Student’s t-636
test; * p-value<0.05; ** p-value<0.01). 637
Figure 8. Kinetics and cooperativity of purified cis and trans crocin 3 transport by CsABCC4a 638
(A) Time-dependent transport of cis (left) or trans (right) crocin 3 at a concentration of 7.5 µM by639
yeast microsomes expressing CsABCC4a or control microsomes (pNEV). (B) Cooperative transport of 640
cis (left) and trans (right) crocin 3 in the presence of different concentrations of the other isomer. Left: 641
cis crocin 3 transport was evaluated at two different concentrations (0.75 µM and 7.5µM) in the 642
presence of increasing concentrations of trans crocin 3 (0, 0.75, 7.5 µM). Right: trans crocin 3 643
transport was evaluated in the same conditions and in the presence of increasing concentrations (0, 644
0.75, 7.5 µM) of cis crocin 3. Results are presented as net transport, calculated by subtracting the 645
values measured in the absence of ATP from the values measured in the presence of ATP after 15 min. 646
Data are the avg ± stdev of three independent microsome preparations. 647

23
Table 1. Inhibition of C. sativus stigma metabolite transport by CsABCC4a and CsABCC2 in the 648
presence of the probenecid inhibitor or by incubation on ice. 649
The table shows the percent residual transport after treatment with probenecid or on ice, compared to 650
control reactions performed at room temperature in absence of inhibitors. Data are the avg ± stdev of 651
three independent microsome preparations. n.d.= not detectable (detection limit of the mass 652
spectrometer in full MS mode: 500 fg). Since the CsABCC2 transporter doesn’t transport cis crocin 1 653
in normal conditions, the percent residual transport is indicated as “-”. 654
CsABCC4a CsABCC2
Compound name Probenecid 0°C Probenecid 0°C
kaempferol 3-O-
sophoroside-7-
glucoside
n.d. 19.3 ± 4.2 n.d. 53.5 ± 6.0
kaempferol 7-
sophoroside 23.0 ± 4.5 34.4 ± 6.9 10.4 ± 3.8 20.8 ± 4.6
Kaempferol 3‐O‐
rutinoside n.d. 35.4 ± 0.01 n.d. 2.3 ± 0.3
Naringenin 7-O-
glucoside 27.7 ± 1.2 11.1 ± 0.7 16.5 ± 0.6 4.4 ± 0.1
trans crocin 4 n.d. 20.0 ± 3.4 14.3 ± 2.7 n.d.
cis crocin 4 n.d. 19.6 ± 3.7 15.2 ± 2.7 n.d.
trans crocin 3 n.d. 48.4 ± 8.4 n.d. n.d.
cis crocin 3 48.5 ± 1.9 22.3 ± 2.2 0.7 ± 0.006 n.d.
trans crocin 2 n.d. 24.8 ± 5.0 18.2 ± 3.6 n.d.
cis crocin 2 26.8 ± 4.8 n.d. 33.6 ± 4.8 0.04 ± 0.005
trans crocin 2’ 28.4 ± 2.0 23.1 ± 1.2 n.d. n.d.
cis crocin 2’ 31.9 ± 2.9 n.d. 32.9 ± 5.2 n.d.
trans crocin 1 22.1 ± 1.5 42.0 ± 6.1 17.4 ± 1.3 n.d.
cis crocin 1 n.d. n.d. - -
655
Table 2. Transport of ABA-GE and cis crocin 3 by CsABCC4a and CsABCC2 transporters. 656
% transport at different substrate concentrations in standard assay conditions. Transport experiments 657
were performed with purified ABA-GE or cis crocin 3 (7.5 µM). Results are the avg ± stdev of three 658

24
independent microsome preparations. n.d.= not detectable (detection limit of the mass spectrometer in 659
full MS mode: 500 fg) 660
Substrate Transporter Substrate concentration (µM)
0.075 0.75 7.5
ABA-GE
CTRL n.d. n.d. n.d.
CsABCC4a n.d. n.d. n.d.
CsABCC2 n.d. n.d. n.d.
cis crocin 3
CTRL n.d. n.d. n.d.
CsABCC4a n.d. n.d. 8.8 ± 0.8
CsABCC2 n.d. n.d. 2.8 ± 0.5
661
References 662
Ahrazem, O., Argandona, J., Fiore, A., Aguado, C., Lujan, R., Rubio-Moraga, A., Marro, M., Araujo-Andrade, 663 C., Loza-Alvarez, P., Diretto, G., and Gomez-Gomez, L. (2018). Transcriptome analysis in tissue sectors 664 with contrasting crocins accumulation provides novel insights into apocarotenoid biosynthesis and 665 regulation during chromoplast biogenesis. Sci Rep 8, 2843. 666
Becker, D.M., and Guarente, L. (1991). [12] High-efficiency transformation of yeast by electroporation. 667 Methods in enzymology 194, 182-187. 668
Bhatia, P., Kolinski, M., Moaddel, R., Jozwiak, K., and Wainer, I.W. (2008). Determination and modelling of 669 stereoselective interactions of ligands with drug transporters: a key dimension in the understanding of 670 drug disposition. Xenobiotica 38, 656-675. 671
Bouvier, F., Suire, C., Mutterer, J., and Camara, B. (2003). Oxidative remodeling of chromoplast carotenoids: 672 identification of the carotenoid dioxygenase CsCCD and CsZCD genes involved in Crocus secondary 673 metabolite biogenesis. Plant Cell 15, 47-62. 674
Brandle, J., and Telmer, P. (2007). Steviol glycoside biosynthesis. Phytochemistry 68, 1855-1863. 675 Burla, B., Pfrunder, S., Nagy, R., Francisco, R.M., Lee, Y., and Martinoia, E. (2013). Vacuolar transport of 676
abscisic acid glucosyl ester is mediated by ATP-binding cassette and proton-antiport mechanisms in 677 Arabidopsis. Plant Physiol 163, 1446-1458. 678
Cinelli, R.A., Ferrari, A., Pellegrini, V., Tyagi, M., Giacca, M., and Beltram, F. (2000). The enhanced green 679 fluorescent protein as a tool for the analysis of protein dynamics and localization: local fluorescence 680 study at the single-molecule level. Photochem Photobiol 71, 771-776. 681
Coman, D., Rutimann, P., and Gruissem, W. (2014). A flexible protocol for targeted gene co-expression 682 network analysis. Methods Mol Biol 1153, 285-299. 683
Darbani, B., Motawia, M.S., Olsen, C.E., Nour-Eldin, H.H., Moller, B.L., and Rook, F. (2016). The biosynthetic 684 gene cluster for the cyanogenic glucoside dhurrin in Sorghum bicolor contains its co-expressed 685 vacuolar MATE transporter. Sci Rep 6, 37079. 686
Demurtas, O.C., Frusciante, S., Ferrante, P., Diretto, G., Azad, N.H., Pietrella, M., Aprea, G., Taddei, A.R., 687 Romano, E., Mi, J., Al-Babili, S., Frigerio, L., and Giuliano, G. (2018). Candidate Enzymes for Saffron 688 Crocin Biosynthesis Are Localized in Multiple Cellular Compartments. Plant Physiol 177, 990-1006. 689

25
Felsenstein, J. (1985). CONFIDENCE LIMITS ON PHYLOGENIES: AN APPROACH USING THE BOOTSTRAP. 690 Evolution; international journal of organic evolution 39, 783-791. 691
Francisco, R.B., and Martinoia, E. (2018). The Vacuolar Transportome of Plant Specialized Metabolites. Plant 692 Cell Physiol. 693
Francisco, R.M., Regalado, A., Ageorges, A., Burla, B.J., Bassin, B., Eisenach, C., Zarrouk, O., Vialet, S., Marlin, 694 T., Chaves, M.M., Martinoia, E., and Nagy, R. (2013). ABCC1, an ATP binding cassette protein from 695 grape berry, transports anthocyanidin 3-O-Glucosides. Plant Cell 25, 1840-1854. 696
French, A.P., Mills, S., Swarup, R., Bennett, M.J., and Pridmore, T.P. (2008). Colocalization of fluorescent 697 markers in confocal microscope images of plant cells. Nat Protoc 3, 619-628. 698
Frusciante, S., Diretto, G., Bruno, M., Ferrante, P., Pietrella, M., Prado-Cabrero, A., Rubio-Moraga, A., Beyer, 699 P., Gomez-Gomez, L., Al-Babili, S., and Giuliano, G. (2014). Novel carotenoid cleavage dioxygenase 700 catalyzes the first dedicated step in saffron crocin biosynthesis. Proc Natl Acad Sci U S A 111, 12246-701 12251. 702
Gibson, D.G., Young, L., Chuang, R.Y., Venter, J.C., Hutchison, C.A., 3rd, and Smith, H.O. (2009). Enzymatic 703 assembly of DNA molecules up to several hundred kilobases. Nat Methods 6, 343-345. 704
Gomez, C., Terrier, N., Torregrosa, L., Vialet, S., Fournier-Level, A., Verries, C., Souquet, J.M., Mazauric, J.P., 705 Klein, M., Cheynier, V., and Ageorges, A. (2009). Grapevine MATE-type proteins act as vacuolar H+-706 dependent acylated anthocyanin transporters. Plant Physiol 150, 402-415. 707
Goodman, C.D., Casati, P., and Walbot, V. (2004). A Multidrug Resistance–Associated Protein Involved in 708 Anthocyanin Transport in Zea mays. The Plant Cell 16, 1812-1826. 709
Hamilton, A.J., and Baulcombe, D.C. (1999). A species of small antisense RNA in posttranscriptional gene 710 silencing in plants. Science 286, 950-952. 711
Hortensteiner, S., and Krautler, B. (2011). Chlorophyll breakdown in higher plants. Biochim Biophys Acta 1807, 712 977-988. 713
Hwang, J.U., Song, W.Y., Hong, D., Ko, D., Yamaoka, Y., Jang, S., Yim, S., Lee, E., Khare, D., Kim, K., Palmgren, 714 M., Yoon, H.S., Martinoia, E., and Lee, Y. (2016). Plant ABC Transporters Enable Many Unique Aspects 715 of a Terrestrial Plant's Lifestyle. Mol Plant 9, 338-355. 716
Jasinski, M., Ducos, E., Martinoia, E., and Boutry, M. (2003). The ATP-binding cassette transporters: structure, 717 function, and gene family comparison between rice and Arabidopsis. Plant Physiol 131, 1169-1177. 718
Johnson, Z.L., and Chen, J. (2017). Structural Basis of Substrate Recognition by the Multidrug Resistance 719 Protein MRP1. Cell 168, 1075-1085.e1079. 720
Klein, M., Martinoia, E., Hoffmann-Thoma, G., and Weissenbock, G. (2000). A membrane-potential dependent 721 ABC-like transporter mediates the vacuolar uptake of rye flavone glucuronides: regulation of 722 glucuronide uptake by glutathione and its conjugates. Plant J 21, 289-304. 723
Klein, M., Geisler, M., Suh, S.J., Kolukisaoglu, H.U., Azevedo, L., Plaza, S., Curtis, M.D., Richter, A., Weder, B., 724 Schulz, B., and Martinoia, E. (2004). Disruption of AtMRP4, a guard cell plasma membrane ABCC-type 725 ABC transporter, leads to deregulation of stomatal opening and increased drought susceptibility. Plant 726 J 39, 219-236. 727
Krumpochova, P., Sapthu, S., Brouwers, J.F., de Haas, M., de Vos, R., Borst, P., and van de Wetering, K. 728 (2012). Transportomics: screening for substrates of ABC transporters in body fluids using vesicular 729 transport assays. Faseb j 26, 738-747. 730
Kumar, S., Stecher, G., and Tamura, K. (2016). MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 731 for Bigger Datasets. Molecular biology and evolution 33, 1870-1874. 732
Lautenschlager, M., Sendker, J., Huwel, S., Galla, H.J., Brandt, S., Dufer, M., Riehemann, K., and Hensel, A. 733 (2015). Intestinal formation of trans-crocetin from saffron extract (Crocus sativus L.) and in vitro 734 permeation through intestinal and blood brain barrier. Phytomedicine 22, 36-44. 735
Lee, H., Sparkes, I., Gattolin, S., Dzimitrowicz, N., Roberts, L.M., Hawes, C., and Frigerio, L. (2013). An 736 Arabidopsis reticulon and the atlastin homologue RHD3-like2 act together in shaping the tubular 737 endoplasmic reticulum. New Phytol 197, 481-489. 738

26
Leier, I., Jedlitschky, G., Buchholz, U., Cole, S.P., Deeley, R.G., and Keppler, D. (1994). The MRP gene encodes 739 an ATP-dependent export pump for leukotriene C4 and structurally related conjugates. J Biol Chem 740 269, 27807-27810. 741
Liu, G., Sanchez-Fernandez, R., Li, Z.S., and Rea, P.A. (2001). Enhanced multispecificity of arabidopsis vacuolar 742 multidrug resistance-associated protein-type ATP-binding cassette transporter, AtMRP2. J Biol Chem 743 276, 8648-8656. 744
Liu, J., Li, Y., Wang, W., Gai, J., and Li, Y. (2016). Genome-wide analysis of MATE transporters and expression 745 patterns of a subgroup of MATE genes in response to aluminum toxicity in soybean. BMC Genomics 17, 746 223. 747
Lu, Y.P., Li, Z.S., Drozdowicz, Y.M., Hortensteiner, S., Martinoia, E., and Rea, P.A. (1998). AtMRP2, an 748 Arabidopsis ATP binding cassette transporter able to transport glutathione S-conjugates and 749 chlorophyll catabolites: functional comparisons with Atmrp1. Plant Cell 10, 267-282. 750
Marinova, K., Pourcel, L., Weder, B., Schwarz, M., Barron, D., Routaboul, J.M., Debeaujon, I., and Klein, M. 751 (2007). The Arabidopsis MATE transporter TT12 acts as a vacuolar flavonoid/H+ -antiporter active in 752 proanthocyanidin-accumulating cells of the seed coat. Plant Cell 19, 2023-2038. 753
Martinoia, E., Maeshima, M., and Neuhaus, H.E. (2007). Vacuolar transporters and their essential role in plant 754 metabolism. J Exp Bot 58, 83-102. 755
Martinoia, E., Meyer, S., De Angeli, A., and Nagy, R. (2012). Vacuolar transporters in their physiological 756 context. Annu Rev Plant Biol 63, 183-213. 757
Matern, U., Heller, W., and Himmelspach, K. (1983). Conformational changes of apigenin 7-O-(6-O-758 malonylglucoside), a vacuolar pigment from parsley, with solvent composition and proton 759 concentration. Eur J Biochem 133, 439-448. 760
McConkey, M.E., Gershenzon, J., and Croteau, R.B. (2000). Developmental regulation of monoterpene 761 biosynthesis in the glandular trichomes of peppermint. Plant Physiol 122, 215-224. 762
Morita, M., Shitan, N., Sawada, K., Van Montagu, M.C., Inze, D., Rischer, H., Goossens, A., Oksman-763 Caldentey, K.M., Moriyama, Y., and Yazaki, K. (2009). Vacuolar transport of nicotine is mediated by a 764 multidrug and toxic compound extrusion (MATE) transporter in Nicotiana tabacum. Proc Natl Acad Sci 765 U S A 106, 2447-2452. 766
Nei, M., and Kumar, S. (2000). Molecular evolution and phylogenetics. (Oxford university press). 767 Nelson, B.K., Cai, X., and Nebenfuhr, A. (2007). A multicolored set of in vivo organelle markers for co-768
localization studies in Arabidopsis and other plants. Plant J 51, 1126-1136. 769 Nour-Eldin, H.H., Andersen, T.G., Burow, M., Madsen, S.R., Jorgensen, M.E., Olsen, C.E., Dreyer, I., Hedrich, 770
R., Geiger, D., and Halkier, B.A. (2012). NRT/PTR transporters are essential for translocation of 771 glucosinolate defence compounds to seeds. Nature 488, 531-534. 772
Paumi, C.M., Chuk, M., Snider, J., Stagljar, I., and Michaelis, S. (2009). ABC transporters in Saccharomyces 773 cerevisiae and their interactors: new technology advances the biology of the ABCC (MRP) subfamily. 774 Microbiol Mol Biol Rev 73, 577-593. 775
Pichersky, E., and Lewinsohn, E. (2011). Convergent evolution in plant specialized metabolism. Annu Rev Plant 776 Biol 62, 549-566. 777
Raichaudhuri, A., Peng, M., Naponelli, V., Chen, S., Sanchez-Fernandez, R., Gu, H., Gregory, J.F., 3rd, Hanson, 778 A.D., and Rea, P.A. (2009). Plant Vacuolar ATP-binding Cassette Transporters That Translocate Folates779 and Antifolates in Vitro and Contribute to Antifolate Tolerance in Vivo. J Biol Chem 284, 8449-8460.780
Re, R., Fraser, P.D., Long, M., Bramley, P.M., and Rice-Evans, C. (2001). Isomerization of lycopene in the 781 gastric milieu. Biochem Biophys Res Commun 281, 576-581. 782
Rubio-Moraga, A., Trapero, A., Ahrazem, O., and Gomez-Gomez, L. (2010). Crocin transport in Crocus sativus: 783 the long road from a senescent stigma to a newborn corm. Phytochemistry 71, 1506-1513. 784
Saitou, N., and Nei, M. (1987). The neighbor-joining method: a new method for reconstructing phylogenetic 785 trees. Molecular biology and evolution 4, 406-425. 786

27
Sauer, N., and Stolz, J. (1994). SUC1 and SUC2: two sucrose transporters from Arabidopsis thaliana; expression 787 and characterization in baker's yeast and identification of the histidine-tagged protein. Plant J 6, 67-77. 788
Schaedler, T.A., Thornton, J.D., Kruse, I., Schwarzlander, M., Meyer, A.J., van Veen, H.W., and Balk, J. (2014). 789 A conserved mitochondrial ATP-binding cassette transporter exports glutathione polysulfide for 790 cytosolic metal cofactor assembly. J Biol Chem 289, 23264-23274. 791
Schneider, S. (2015). Inositol transport proteins. FEBS Lett 589, 1049-1058. 792 Shoji, T., Inai, K., Yazaki, Y., Sato, Y., Takase, H., Shitan, N., Yazaki, K., Goto, Y., Toyooka, K., Matsuoka, K., 793
and Hashimoto, T. (2009). Multidrug and toxic compound extrusion-type transporters implicated in 794 vacuolar sequestration of nicotine in tobacco roots. Plant Physiol 149, 708-718. 795
Song, W.Y., Park, J., Mendoza-Cozatl, D.G., Suter-Grotemeyer, M., Shim, D., Hortensteiner, S., Geisler, M., 796 Weder, B., Rea, P.A., Rentsch, D., Schroeder, J.I., Lee, Y., and Martinoia, E. (2010). Arsenic tolerance in 797 Arabidopsis is mediated by two ABCC-type phytochelatin transporters. Proc Natl Acad Sci U S A 107, 798 21187-21192. 799
Takanashi, K., Yamada, Y., Sasaki, T., Yamamoto, Y., Sato, F., and Yazaki, K. (2017). A multidrug and toxic 800 compound extrusion transporter mediates berberine accumulation into vacuoles in Coptis japonica. 801 Phytochemistry 138, 76-82. 802
Tarantilis, P.A., Tsoupras, G., and Polissiou, M. (1995). Determination of saffron (Crocus sativus L.) 803 components in crude plant extract using high-performance liquid chromatography-UV-visible 804 photodiode-array detection-mass spectrometry. J Chromatogr A 699, 107-118. 805
Theodoulou, F.L. (2000). Plant ABC transporters. Biochim Biophys Acta 1465, 79-103. 806 Tommasini, R., Evers, R., Vogt, E., Mornet, C., Zaman, G.J., Schinkel, A.H., Borst, P., and Martinoia, E. (1996). 807
The human multidrug resistance-associated protein functionally complements the yeast cadmium 808 resistance factor 1. Proc Natl Acad Sci U S A 93, 6743-6748. 809
Uchida, Y., Kamiie, J., Ohtsuki, S., and Terasaki, T. (2007). Multichannel liquid chromatography-tandem mass 810 spectrometry cocktail method for comprehensive substrate characterization of multidrug resistance-811 associated protein 4 transporter. Pharm Res 24, 2281-2296. 812
Wilkens, S. (2015). Structure and mechanism of ABC transporters. F1000Prime Rep 7, 14. 813 Zhao, J., and Dixon, R.A. (2009). MATE transporters facilitate vacuolar uptake of epicatechin 3'-O-glucoside for 814
proanthocyanidin biosynthesis in Medicago truncatula and Arabidopsis. Plant Cell 21, 2323-2340. 815 Zhou, Q., Yu, L.S., and Zeng, S. (2014). Stereoselectivity of chiral drug transport: a focus on enantiomer-816
transporter interaction. Drug Metab Rev 46, 283-290. 817
818

Figure 1. Saffron crocin biosynthesis and its compartmentation
(A) Saffron flower at anthesis. (B) Scanning electron microscopy of mature stigma. (C) Proposed crocin biosynthetic
pathway and (D) its compartmentation (Demurtas et al, 2018): the CCD2 enzyme cleaves zeaxanthin in the plastid,
producing crocetin dialdehyde, which then migrates to the endoplasmic reticulum (ER) and is dehydrogenated to
crocetin by an aldehyde dehydrogenase (ALDH); the glycosylation steps are performed in the cytoplasm by UDP-
glycosyltransferase (UGT) enzymes; crocins are then transported into the vacuole by ABC tonoplast transporters
functionally characterized in this work. (E) Detailed crocin structures.
C
A B
500 µm
ChromoplastZeaxanthin V
ac
uo
le
Crocetin
dialdehyde
ER
Crocins
Cytoplasm
Tonoplast
transporters?
500 nm
UGTs
ALDH
CCD2
Crocetin
D
zeaxanthin OH
HO
picrocrocin
safranal
O
crocetin dialdehyde3-OH-β-cyclocitral
OHO
O
R1OO
O
UGT?
?
CCD2
ALDH3I1
OO
UGTsHO
OH
OO
crocetin
crocins
R1O
R2O
E

Figure 2. Expression and phylogenetic analysis of putative C. sativus tonoplast transporters.
(A) Heat map of transcript levels for ABCC and MATE transporter genes in different C. sativus tissues; data are
expressed as log2 of reads per kilobase per million (RPKM) and sorted by decreasing RPKM values in stigma. (B) Co-
expression analysis of genes for CsABCCs and CsMATEs (Bait) with CsCCD2 and with total crocins (Prey). Only ρ
values >0.50 are shown. (C) Phylogenetic relationships of ABCC (left) and MATE (right) transporters expressed in C.
sativus (Cs) stigma (underlined) inferred using the neighbor-joining method. Colored dots indicate the class of
transported substrates for the functionally characterized transporters. The CsABCCs and CsMATEs functionally
characterized in this work are indicated by arrows. The trees include transporters from Arabidopsis thaliana (At), Coptis
japonica (Cj), Medicago truncatula (Mt), Nicotiana tabacum (Nt), Sorghum bicolor (Sb), Vitis vinifera (Vv) and Zea mays
(Zm). The accession numbers are described in Methodsthe Accession numbers section. The percentage of replicate
trees that clustered together in the bootstrap test (500 replicates) is indicated to the left of the branches.
Acorm leaf stamen tepal stigma
CsABCC4aCsABCC2
CsABCC4bCsABCC8CsABCC4cCsABCC5CsABCC15
CsABCC3b
CsABCC3a
CsMATE4
CsMATE1aCsMATE1bCsMATE2aCsMATE2f
CsMATE2dCsMATE5bCsMATE2b
CsMATE5aCsMATE2e
CsMATE2c
B Bait Pearson’s
ρ
Prey
CsCCD2 0.96 Total crocins
CsABCC4a 0.99 Total crocins
CsMATE4 0.84 Total crocins
CsABCC4a 0.95 CsCCD2
CsABCC2 0.90 CsCCD2
CsMATE4 0.75 CsCCD2
AtABCC5
ZmABCC4
AtABCC2
AtABCC1AtABCC1
AtABCC2
CsABCC2
AtABCC11
AtABCC12
AtABCC13
AtABCC4
AtABCC14
CsABCC4a
CsABCC4b
CsABCC4c
CsABCC8
AtABCC8
ZmABCC3
AtABCC10
VvABCC1
AtABCC9
AtABCC15
CsABCC15
CsABCC3a
CsABCC3b
AtABCC3
AtABCC6
AtABCC7
ZmABCC4
CsABCC5
AtABCC5
CsABCC3a
CsABCC3b
AtABCC10
AtABCC3
ZmABCC3
CsABCC4b
CsABCC2
AtABCC9
AtABCC11
AtABCC12
AtABCC13
AtABCC4
AtABCC14
CsABCC4a
CsABCC4c
CsABCC8
AtABCC8
VvABCC1
AtABCC15
CsABCC15
AtABCC7
CsABCC5
AtABCC6
CsMATE2e
AtDTX35
NtJAT1
AtDTX33
CsMATE5b
SbMATE2
NtMATE1
CsMATE1b
CsMATE1a
VvAM3
VvAM1
NtJAT2
AtDTX40AtDTX40
SbMATE2
CsMATE1a
CsMATE1b
NtMATE1
NtMATE2
AtDTX41/TT12
MtMATE1
AtDX19
NtJAT1
CsMATE5a
CsMATE5b
AtDTX16
AtDTX17
CsMATE2c
CsMATE2b
CsMATE2a
MtMATE2
AtDTX35
NtJAT2
VvAM1
VvAM3
CsMATE2d
AtDTX33
CJMATE1
CsMATE4
AtDTX29
AtDTX30
CsMATE2e
CsMATE2f
NtMATE2
AtDTX41/TT12
MtMATE1
AtDTX19
CsMATE5a
AtDTX16
AtDTX17
CsMATE2c
CsMATE2b
CsMATE2a
MtMATE2
CsMATE2d
CjMATE1
CsMATE4
AtDTX29
AtDTX30
CsMATE2f
Substrates
flavonoids
chlorophyll catabolites
folates
phytate
ABA-GE
phytochelatins
glutathione conjugates
alkaloids
cyanogenic glycosides
C

Figure 3. Subcellular localization of C. sativus transporters in Nicotiana leaves.
Confocal images of GFP (green) and RFP (red) fluorescence in N. benthamiana leaves co-expressing the indicated
ABCC or MATE transporter fused to GFP and γTIP (a tonoplast marker) fused to RFP. Scale bars, 10 µm.
CsA
BC
C4
a:G
FP
CsA
BC
C2
:GF
P
γTIP:RFP mergeGFP
CsM
AT
E4
:GF
PC
sM
AT
E1
a:G
FP
CsM
AT
E1
b:G
FP

Figure 4. Transportomics assay using C. sativus stigma extract and yeast microsomes.
(A) Schematic representation of the assay. Microsomes overexpressing a transporter are incubated with
saffron hydroalcoholic extract in the presence of ATP. Microsomes are separated by the rapid filtration technique
and washed, and transported metabolites are eluted and quantified by LC-PDA-HRMS.
(B-E) Representative ESI+/MS chromatograms of the extracted accurate mass of crocetin (M+H+ 329.1747)
generated from crocin fragmentation. (B) C. sativus stigma hydroalcoholic extract; (C) import into microsomes
isolated from yeast cells transformed with the pNEV empty vector; (D) import into microsomes expressing
CsABCC4a incubated in the absence of ATP; (E) import into microsomes expressing CsABCC4a incubated in the
presence of ATP. Incubation was for 15 min. Different peak represent: trans crocin 4 (RT 10.58); trans crocin 3 (RT
11.52); trans crocin 2ʹ (RT 12.54); cis crocin 4 (RT 13.84); trans crocin 2 (RT 14.20); cis crocin 3 (RT 14.82); trans
crocin 1 (RT 16.16); cis crocin 2ʹ (RT 16.79); cis crocin 2 (RT 17.02); cis crocin 1 (RT 17.73).
ATP
ATP
ATP
Yeast cells
Saffron extract
In vitro transport assay
Microsome
recovery
Analysis of uptake
by LC-PDA-HRMS
A
0
0
0
trans
4
106
Ion
in
ten
sit
y
106
Ion
in
ten
sit
y
107
Saffron extract
CsABCC4a - ATP
CsABCC4a + ATP
Ion
in
ten
sit
y
B
0
106
Ion
in
ten
sit
y
205 10 15 25 30
CTRL + ATP
trans
3
trans2’
cis
4 trans
2cis
3tr
ans
1cis2’
cis
2
trans
4
trans
3tr
ans
3tr
ans
3tr
ans
4tr
ans 4
trans2’
trans2’
trans2’
cis
4cis
4tr
ans
2cis
3cis
3
trans
2tr
ans 2
cis
4
cis
3
trans
1cis2’
cis
1
D
C
E

Figure 5. CsABCC4a and CsABCC2 transport
different crocins in vitro
(A) Major glycosylated metabolites present in the C.
sativus stigma hydroalcoholic extract. 1: kaempferol 3-
O-sophoroside-7-glucoside; 2: kaempferol 3,7,4ʹ-
triglucoside; 3: kaempferol 7-sophoroside; 4:
kaempferol 3-β-D-glucopyranoside; 5:
kaempferol‐3‐O‐rutinoside; 6: dihydrokaempferol 7-O-
glucoside; 7: naringenin glucosides (3 isomers); 8:
picrocrocin; 9: trans crocin 5 (t 5); 10: trans crocin 4 (t
4); 11: cis crocin 4 (c 4); 12: trans crocin 3 (t 3); 13: cis
crocin 3 (c 3); 14: trans crocin 2 (t 2); 15: cis crocin 2
(c 2); 16: trans crocin 2ʹ (t 2’); 17: cis crocin 2ʹ (c 2’);
18: trans crocin 1 (t 1); 19: cis crocin 1 (c 1). Structures
of the various crocins are shown in Figure 1E. Values
are presented as relative abundances compared to the
internal standard used (formononetin) of mean values
± stdev of three independent extract preparations.
(B) % net transport of glycosylated metabolites by
yeast microsomes expressing candidate tonoplast
transporters. CTRL: microsomes form yeast cells
transformed with the empty vector. CsABCC4a,
CsABCC2, CsMATE4, CsMATE1a, CsMATE1b:
microsomes expressing candicate tonoplast
transposters. % net transport was calculated by
subtracting the peak areas in the absence of ATP from
those in the presence of ATP after 15 min of
incubation, and normalizing to the peak areas in the
initial extract. Data are the avg ± stdev of three
independent microsome preparations.
B
0
2
4
6
8
10
% t
ran
sp
ort
t4 t
3c4
c3
t2
c2
t2’
c2’
t1
c1
CsABCC4a
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19
0
0.2
0.4
0.6
0.8
1Saffron extract
t 4
t 5
t3
c4
c3 t2
c 2 t2’
c2’
t1
c1
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19
Flavonoids Crocins
Rela
tive a
bu
nd
an
ce
0
2
4
6
8
10CsMATE4
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19
0
2
4
6
8
10
t 4 t3
c4
c3
t2
c2
t2’ c2’
t1
% t
ran
sp
ort
CsABCC2
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19
0
2
4
6
8
10
% t
ran
sp
ort
CTRL
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19
0
2
4
6
8
10CsMATE1a
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19
0
2
4
6
8
10CsMATE1b
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19
A
% t
ran
sp
ort
% t
ran
sp
ort
% t
ran
sp
ort

Figure 6. Tonoplast localization of CsABCC4a in Nicotiana leaves.
(A-C) N. benthamiana leaves were agroinfiltrated with the indicated constructs and imaged by confocal microscopy.
The Spearman correlation coefficient (Rs) between the two channels was calculated using the ImageJ analysis
program with the Pearson-Spearman correlation plug-in (French et al., 2008), to calculate co-localization and to
produce scatterplots using a threshold of 40. Scale bars, 5 µm. (D) Confocal images of a protoplast from a N.
benthamiana leaf expressing CsABCCa:GFP (green channel). After induced lysis of the plasma membrane the
vacuole (asterisk) is in the process of being released. Scale bar, 12 µm.
* **
CsABCC4a:GFP
mergebright field CsABCC4a:GFP chlorophyll
D
RFP-HDEL
merge
CsABCC4a:GFP
20 µm12 µm
Rs = -0.05
FM4-64
γTIP-RFP merge
CsABCC4a:GFP merge
Rs = + 0.75
Rs = - 0.05
colocalization
A
B
C
colocalization
colocalization

Figure 7. Functional assay of the C. sativus ABCC4a transporter in N. benthamiana leaves.
Quantification of crocetin and crocins in leaves agroinfiltrated to express CsCCD2 or CsCCD2+CsABCC4a. Data are
the avg ± stdev of ion peak areas of crocetin and crocins, normalized to the ion peak area of the internal standard
formononetin (fold internal standard), in three independent pools of agroinfiltrated leaves (see Methods for details).
Asterisks indicate statistical significance of the difference between the means of the two groups (CsCCD2 and
CsCCD2+ CsABCC4a) (Student’s t-test; * p-value<0.05; ** p-value<0.01).
0
2
4
6
8
Fold
IS
CCD2 CCD2+ABCC4
**
** **
*
CsCCD2 + CsABCC4aCsCCD2**
**

Figure 8. Kinetics and cooperativity of purified cis and trans-crocin 3 transport by CsABCC4a
(A) Time-dependent transport of cis (left) or trans (right) crocin 3 at a concentration of 7.5 µM by yeast microsomes
expressing CsABCC4a or control microsomes (pNEV). (B) Cooperative transport of cis (left) and trans (right) crocin 3
in the presence of different concentrations of the other isomer. Left: cis crocin 3 transport was evaluated at two different
concentrations (0.75 µM and 7.5µM) in the presence of increasing concentrations of trans crocin 3 (0, 0.75, 7.5 µM).
Right: trans crocin 3 transport was evaluated in the same conditions and in the presence of increasing concentrations
(0, 0.75, 7.5 µM) of cis crocin 3. Results are presented as net transport, calculated by subtracting the values measured
in the absence of ATP from the values measured in the presence of ATP after 15 min. Data are the avg ± stdev of three
independent microsome preparations.
0
20
40
60
80
100
120
140
0 5 10 15 20 25
ng
cis
cro
cin
3/m
g p
rote
in
CsABCC4a
pNEV
cis 7.5 μM
A
B
0
50
100
150
200
250
300
trans 0 trans 0.75 trans 7.5
ng
cis
cro
cin
3/m
g p
rote
in
0
50
100
150
200
250
300
cis 0 cis 0.75 cis 7.5
ng
tra
ns
cro
cin
3/m
g p
rote
in
0
20
40
60
80
100
120
140
0 5 10 15 20 25
ng
tra
ns
cro
cin
3/m
g p
rote
in
CsABCC4a
pNEV
trans 7.5 μM
cis trans
0.75 μM
7.5 μM
0.75 μM
7.5 μM

DOI 10.1105/tpc.19.00193; originally published online September 23, 2019;Plant Cell
Giosuè Costa, Stefano Alcaro, Enrico Martinoia and Giovanni GiulianoPietrella, Giuseppe Aprea, Lorenzo Borghi, Mistianne Feeney, Lorenzo Frigerio, Adriana Coricello,
Olivia Costantina Demurtas, Rita Francisco, Gianfranco Diretto, Paola Ferrante, Sarah Frusciante, MarcoABCC transporters mediate the vacuolar accumulation of crocins in saffron stigmas
This information is current as of October 24, 2020
Supplemental Data /content/suppl/2019/09/23/tpc.19.00193.DC1.html
Permissions https://www.copyright.com/ccc/openurl.do?sid=pd_hw1532298X&issn=1532298X&WT.mc_id=pd_hw1532298X
eTOCs http://www.plantcell.org/cgi/alerts/ctmain
Sign up for eTOCs at:
CiteTrack Alerts http://www.plantcell.org/cgi/alerts/ctmain
Sign up for CiteTrack Alerts at:
Subscription Information http://www.aspb.org/publications/subscriptions.cfm
is available at:Plant Physiology and The Plant CellSubscription Information for
ADVANCING THE SCIENCE OF PLANT BIOLOGY © American Society of Plant Biologists















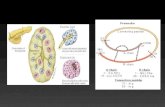

![Ions channels/transporters and chloroplast regulation · transporters/pumps and secondary transporters (according to the Transport Classification system [1]). Channels transport](https://static.fdocuments.in/doc/165x107/601623c1d6936b1074546c48/ions-channelstransporters-and-chloroplast-transporterspumps-and-secondary-transporters.jpg)

